2. 上海海洋大学 水产科学国家级实验教学示范中心, 上海 201306
脊椎动物的性别决定主要分为基因型性别决定(genetic sex determination,GSD)和环境依赖型性别决定(environmental sex determination,ESD)[1],其中,温度依赖型性别决定(temperature sex determination,TSD)属于ESD中最典型的一种模式。TSD动物的性别不由遗传因素决定,而是取决于特定胚胎时期——温度敏感期(temperature sensitive period,TSP)的环境温度,该时期性腺对热信号敏感并具有性别双向分化的潜能[2-3]。红耳龟(Trachemys scripta)被视为TSD研究的一种理想的动物模型,26 ℃产雄温度(male producing temperature, MPT)下产出100%的雄性后代,32 ℃产雌温度(female producing temperature, FPT)下产出100%的雌性后代[4]。与哺乳动物[5-6]和鸡[7]等GSD研究相比,目前TSD研究较为落后,主要局限在组织学观察、性别分化相关基因的两性差异表达等关联性分析,尚未涉及功能鉴定和机制研究。20世纪90年代开始,学者们通过PCR和原位杂交等技术对TSD动物相关基因进行了克隆和表达分析,锚定了一些潜在的性别发育基因,如Wt1[8-9]、Sf1[10-11]、Dax1[12-13]、Sox9[12, 14-15]、Dmrt1[12, 16-17]、Cyp19a1[18-19]等。参与红耳龟性别分化的雄性关联基因有Sox9[14-15]、Dmrt1[16-17]、Mis/Amh[18]等,雌性分化关联基因有Cyp19a1[19-20]、Rspo1[21]、Foxl2[18]等。这些基因是GSD动物性别分化的关键保守基因,提示TSD和GSD机制很可能共享一套性别分化的下游信号通路。而关于环境温度如何转换成生物信号调控性别决定基因表达(温度感受机制)的研究非常少,导致TSD机制长期未得到破解。采用基因组、转录组学等技术,筛选一些能直接感受温度的温度敏感因子并进行功能鉴定将是推进TSD研究的一项重要举措。
目前,TSD分子机制已有几种假说。首先,类固醇激素被认为在TSD非哺乳脊椎动物的性别分化中发挥关键作用[22]。糖皮质激素已被证明在包括爬行动物的多种脊椎动物中可诱导性逆转[23-24],而固醇类激素合成如何由温度来调节,仍不清楚。其次,表观遗传修饰能将环境因素和基因表达联系起来,也已被证明参与性别发育过程,如Kdm6B [25-26]。此外,其他与温度应激相关的因素,如热休克蛋白HSPs(heat shock proteins, HSPs)和冷诱导RNA结合蛋白(cold inducible RNA binding proteins, CIRBPs)也被视为在TSD中发挥作用[19, 27]。最近,CASTELLI等[28]还提出了关于环境信号与性别决定通路内在联系机制的模型,指出ESD物种中细胞通过高度保守的氧化还原态钙(CaRe)——主要由Ca2+和活性氧(reactive oxygen species,ROS)刺激后激活并感知环境信号,转导相应信号通路,或影响表观遗传过程来使性别有关基因差异表达。RADHAKRISHNAN等[29]为检测TSD和GSD物种在转录水平机制和作用环路是否一致,比较分析了TSP前和TSP阶段的锦龟(Chrysemy spicta)性腺以及同时期的角鳖(Apalone spinifera)性腺转录组数据,发现TSD物种对温度更为敏感。此外,Cirbp基因在两个物种较高温度下上调,表明热信号能转为生物信号来调控性别发育基因表达,期间可能由“信号和激素通路”“RNA绑定”“热休克通路”“Jak-Stat信号通路”“Nf-κB信号通路”“Wnt信号通路”等不同调控途径实现。但这仅是推测,如何将温度和性别决定基因耦联起来有待进一步研究。
本研究选取第16期(性别决定关键时期)红耳龟MPT和FPT胚胎性腺组织,制备测序文库,比较分析性腺转录本,筛选差异表达基因,对温度敏感因子和类固醇激素通路着重展开讨论,为研究红耳龟TSD的分子调控机制提供基础数据。
1 材料与方法 1.1 红耳龟受精卵孵化与采集本实验所用红耳龟龟卵购自湖南汉寿,均采集于产卵后12 h内。设置温度26 ℃(MPT)和32 ℃(FPT)于恒温恒湿孵化箱进行孵化,湿度保持在75%~85%。丢弃未受精的龟卵,待发育到15期(性腺开始分化)时,参考红耳龟发育图谱[30],收集16期(性腺初形成)、23和25期(性别决定后)的两性性腺,一部分置于Trizol试剂(Invitrogen,美国)以提取RNA,另一部分于4%多聚甲醛(上海,国药集团)中固定24 h,乙醇梯度脱水后,于70%乙醇4 ℃长期保存,以切片备用。
1.2 RNA提取、测序文库构建及测序分别采集F16和M16性腺,每20对为1个混样,各设3个生物学重复。根据Trizol试剂盒法(Invitrogen,美国),用Recombinant DNaseⅠ(TaKaRa,日本)除去基因组,检测RNA完整程度,采用微量核酸测定仪(Nannodrop)测定总RNA浓度、纯度,根据Fermentas反转录试剂盒(K1622,Thermo Scientific,美国)法实现文库构建。根据Qubit2.0、Agilent 2100和Q-PCR法确保文库浓度、质量后,送至IlluminaHiSeq 2500平台上机测序(委托北京百迈客完成)。
1.3 测序数据质量分析所得的原始数据(Raw Reads)过滤后得到高质量的测序数据(Clean Data)。计算Q10、Q20、Q30、Q40、GC含量和重复序列水平,经Trinity进行序列拼接、过滤和组装获得高质量的Unigene。
1.4 基因表达水平检测与差异表达筛选根据RSEM估计基因定量表达水平,结合FPKM (Fragment Per Kilo Bases per Million reads)值来呈现Unigene的表达量,参照Benjamini-Hochberg法,经显著性P域值校正,以最终的P——FDR (False Discovery Rate)为标准。筛选出FDR≤0.01且差异倍数FC(Fold Change)≥2为显著性差异表达基因。
1.5 差异表达基因GO注释和KEGG代谢途径分析使用KOBAS 2.0得到Unigene在KEGG中的注释结果,运用WEGO和KEGG进行功能统计和代谢通路分类,分析差异基因的富集情况,挖掘更多性别相关且具有性别二态性表达的基因。
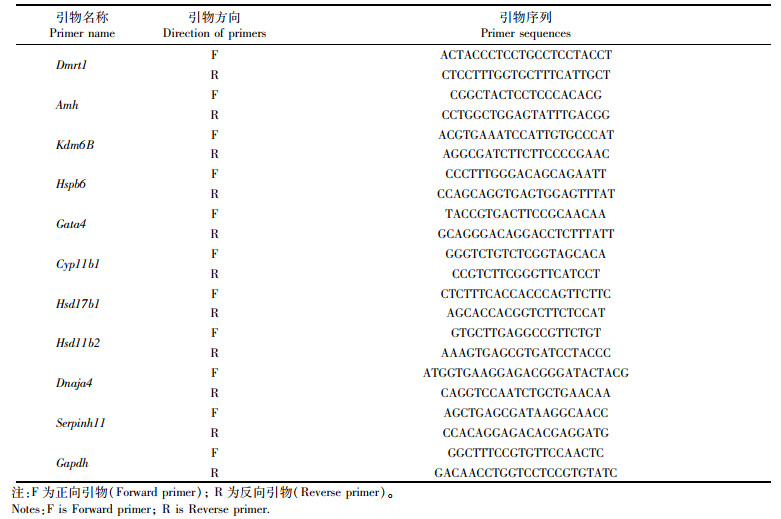
1.6 实时荧光定量PCR分析将合成的16期两性性腺cDNA模板,使用SYBR®Premix(TaKaRa)选择10个基因进行qRT-PCR扩增,配置12.5 L反应液体系如下:6.25 μL SYBR® Green Ⅰ,上下游引物各0.5 μL,cDNA模板1 μL,灭菌蒸馏水4.25 μL。Primer 5.0设计引物,以Gapdh作为内参,采用2-ΔΔCt计算分析转录过程的表达情况,所用引物见表 1。
|
表 1 所有引物序列 Tab.1 Primers sequences |
将70%乙醇中保存的23、25期两性性腺,经乙醇梯度脱水、二甲苯透明、浸蜡后,进行石蜡包埋,Leica石蜡切片机(RM2235)制片,厚度为6~8 μm。成片通过脱蜡、乙醇梯度复水后,抗原修复液95 ℃处理20~30 min,室温备用。(1)苏木素-伊红染色:苏木素核染、分化、伊红染色、脱水、中性树胶封片、晾干,正置显微镜(Nikon)下观察拍照;(2)免疫荧光染色:封闭液室温孵育1 h,吸去液体,分别孵育兔抗SOX9(1∶ 500,Millipore,USA)、兔抗AMH(1∶ 200,华安生物)、羊抗FOXL2(1∶ 200,华安生物)和鼠抗CTNNB1(1∶ 500,Sigma,USA),4 ℃过夜。用磷酸盐Triton X-100缓冲液(PBST)清洗15 min,重复3次。在室温避光环境下,滴加羊抗兔IgG-488(1∶ 500,Invitrogen)、驴抗羊IgG-488(1∶ 500,联科生物)、羊抗鼠IgG-594(1∶ 500,Invitrogen)和DAPI(1∶ 50,Sigma)稀释液,孵育2 h,用PBST和洗脱液清洗15 min,重复3次。采用抗荧光淬灭剂封片,于共聚焦荧光显微镜(Nikon,A1 Plus)下观察拍照。
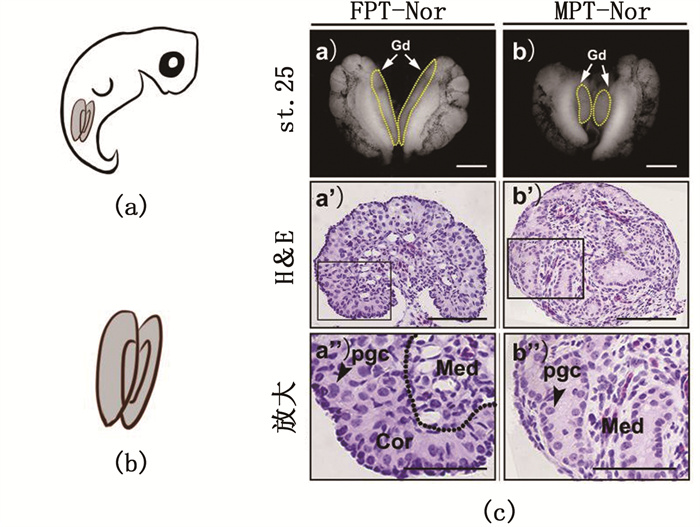
2 结果 2.1 红耳龟胚胎性腺采集及形态学观察16期红耳龟龟胚示意图 1a和1b可见,中肾复合物在靠近尾部位置。体视显微镜下观察到,25期FPT性腺发育呈扁平、细长状(图 1c-a),MPT性腺则相对短粗(图 1c-b),经苏木素-伊红染色后,结果显示:相比25期FPT性腺高度发育的皮质区,MPT性腺退化为单细胞层;MPT性腺髓质区中,原始性索上分布着许多生殖细胞,FPT性腺的髓质区已完全退化(图 1c-a’,1c-b’)。
|
Gd.性腺,标尺.1mm; pgc.原始生殖细胞; Cor.皮质区; Med.髓质区,标尺.50 μm。 Gd. gonad, Scale bar.1 mm; pgc. primordial germ cell; Cor. cortical area; Med. medulla area, Scale bar. 50 μm. 图 1 红耳龟胚胎性腺取样示意及组织形态学观察 Fig. 1 Gonad sampling scheme and histological observation of T.scripta gonads |
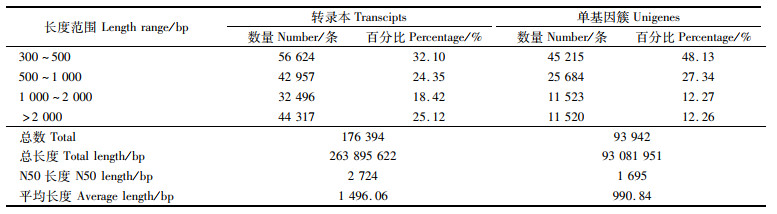
测序所得的性腺转录本样品Clean Data均达到2.38 Gb,Q30碱基百分比均大于89.54%,GC含量在50.32%~52.32%。经组装共得176 394条转录本和93 942条Unigene,Transcript、Unigene的N50与平均长度分别为2 724 bp、1 695 bp和1 496.06 bp、990.84 bp,见表 2。
|
表 2 红耳龟胚胎性腺转录组组装结果 Tab.2 Transcriptome assembly of T.scripta embryonic gonads |
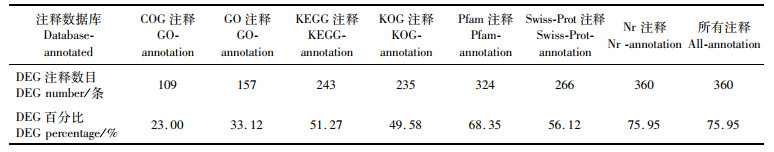
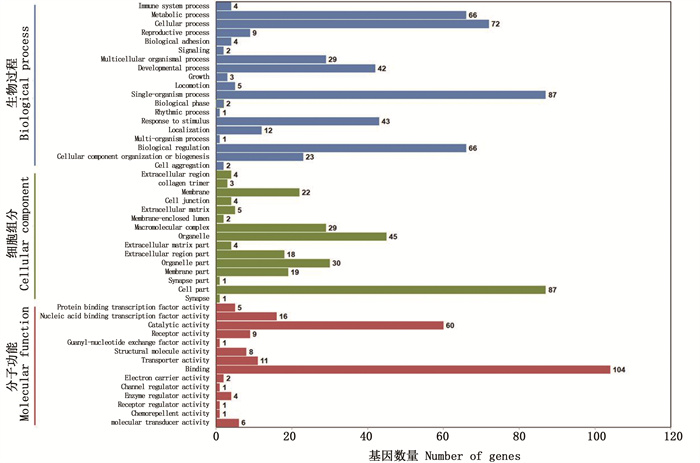
经COG、GO、KEGG、KOG、Pfam及Swiss-Prot数据库注释,共获得75.95%(474个中的360个)DEG,其中,NR数据库中注释到的基因数最多,360(75.95%)个,具体DEG注释统计信息见表 3。GO注释分为3个大类别,257个DEG被注释到60个GO term上,注释数量最多三大子类别有:生物学过程中的细胞过程,87个;细胞组成中的细胞部分,87个;分子功能中的绑定,104个,见图 2。
|
表 3 差异表达的编码转录本注释概述 Tab.3 Overview of differentially expressed coding transcripts annotation |
|
GO三大类别下的子分支,分为生物过程(蓝色柱)、细胞组分(绿色柱)和分子功能(红色柱)。 A sub-branch of the three GO categories, divided into biological processes (blue bar), cellular components (green bar), and molecular functions (red bar). 图 2 差异表达基因GO功能注释统计 Fig. 2 Statistics of GO function annotation of differentially expressed genes |
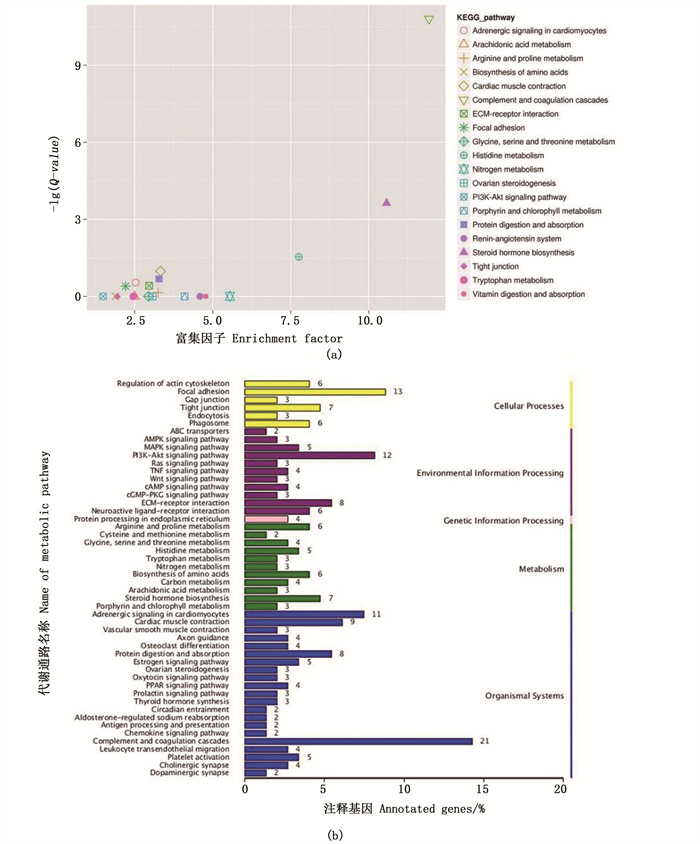
KEGG注释结果显示,474个显著性差异基因被映射到139条代谢通路,显著性富集最可靠的前20条通路[即log10(Q-value)越大,可靠性越高]中,排前三的有“补体和凝血级联”(21个DEG)、“类固醇激素代谢合成通路”(7个DEG)、“组氨酸代谢”(5个DEG),见图 3a。同样,在一级分类的“环境信息处理”中,筛选到6条涉及性别分化的途径:“PI3K-Akt信号通路”、“TGF-β信号通路”、“cAMP信号通路”、“雌激素信号通路”、“Wnt信号通路”、“炎症介质对TRP通道的调节)”(图 3b)。经统计,共有243个DEG涉及这前20条显著富集的KEGG通路中。
|
(a) X、Y轴分别代表KEGG富集因子、KEGG途径中差异表达的编码转录本的富集可靠性;(b) X、Y轴分别为所注释对应通路的基因个数及其占所有注释基因数的比率和代谢通路名称。 (a) X axis and Y axis respectively represent KEGG enrichment factors and the enrichment reliability of coding transcripts differentially expressed in the KEGG pathway; (b) X and Y axis are the number of genes in the corresponding pathway annotated and their ratio to all annotated genes and the name of metabolic pathway, respectively. 图 3 差异表达基因KEGG通路分类 Fig. 3 Annotation classification of differentially expressed genes KEGG pathway |
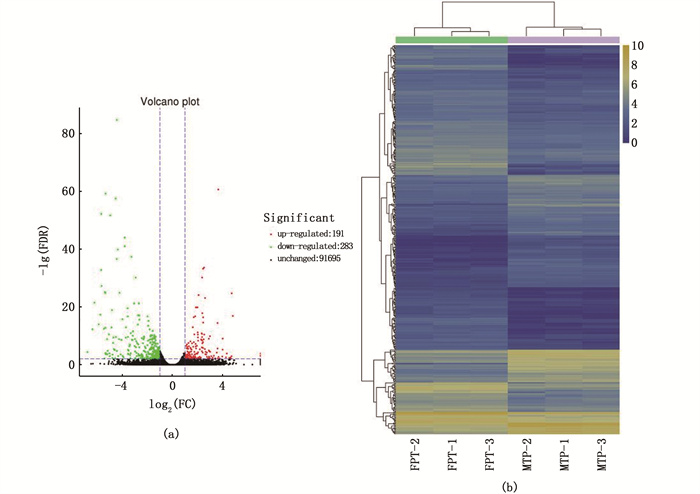
所得测序结果,按FDR≤0.01且差异倍数FC(Fold Change)≥2和聚类分析筛选差异表达基因,共获474个性别偏向表达基因,相对于FPT性腺DEGs,上调表达基因283个,下调表达基因191个(图 4a)。火山图展示了差异基因的表达情况,基因聚类分析呈现了基因整体的表达模式(图 4)。
|
(a)红色、绿色分别代表上、下调基因,黑色代表无显著性差异表达基因;(b)蓝色为弱表达基因,黄色为高表达基因。 (a)Red and green represent up-regulated and down-regulated genes, respectively, while black represent genes with no significant differential expression. (b) Blue is weakly expressed gene and yellow is highly expressed gene. 图 4 性腺差异表达基因火山图及聚类热图 Fig. 4 Volcano plot and heatmap of differentially expressed genes in turtle gonads |
为挖掘一些在TSD中起重要作用的基因,通过红耳龟早期性腺转录本,结合GO注释和KEGG代谢通路,筛选获得了参与性别决定、性腺发育和分化、信号转导等相关靶基因。转录组数据显示,以FPT性腺为对照组,红耳龟Dmrt1、Amh、Sf1、Gata4基因在MPT性腺中显著上调。其中,Kdm6B、Dmrt1和Amh的表达量分别约为FPT性腺中的134倍、16倍和30倍。Dmrt1参与雄性性别决定(GO: 0030238)和支持细胞分化(GO: 0060008)。Amh的KEGG编码通路为K04665,调控缪勒氏管的形成,指导雄性性别决定和性腺分化。Sox9是脊椎动物雄性决定和睾丸分化的一种重要转录因子,该基因在MPT和FPT性腺中均高表达,推测其参与红耳龟早期性腺发育过程。
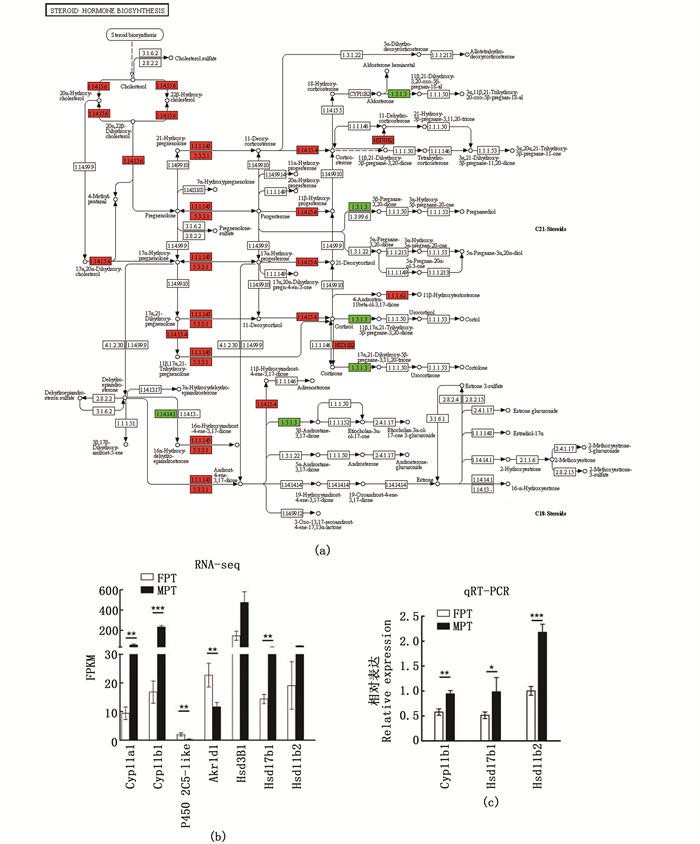
KEEG富集散点图显示,类固醇激素代谢合成通路中有7个显著性差异表达的基因,cytochrome P450 2C5-like、Akr1d1在FPT性腺中高表达,Cyp11a1、Cyp11b1、Hsd3b1、Hsd17b1、Hsd11b2在MPT性腺中高表达(图 5a)。cytochrome P450 2C5-like编码细胞色素P450 2C5酶,能催化孕酮的21-羟基化,从而形成脱氧皮质酮。Cyp11a1基因编码胆固醇侧链裂解酶是一种线粒体酶,催化胆固醇为孕烯醇酮,启动类固醇激素合成的第一步,是合成途径的关键调控位点。Hsd3b1编码一种双功能酶,将3beta-羟基-delta5甾体前体氧化和异构化为3-oxo-delta4甾体,这是甾体激素生物合成的必要步骤。Hsd17b1编码17Beta-羟基类固醇脱氢酶,催化雌激素活化的最后一步,利用NADPH将雌激素E1和E2以及4-雄烯二酮等雄激素转化为睾酮。同时,qRT-PCR结果也显示,Cyp11b1、Hsd17b1、Hsd11b2在MPT中高表达,与测序数据一致(图 5c)。以上结果强烈提示类固醇激素对红耳龟性腺分化起着重要的调控作用。
|
(a)红色、绿色分别代表上、下调基因;(b)和(c) n≥3,柱状图上方“*”代表差异显著(P<0.05)。 (a)Red and green represent up-regulated and down-regulated genes, respectively; (b) and (c) n≥3, "*"at the top of the bar chart represents significant difference (P<0.05). 图 5 类固醇激素代谢合成通路中的差异表达基因及部分验证结果 Fig. 5 Differentially expressed genes and several validation results in steroid hormone synthesis pathway |
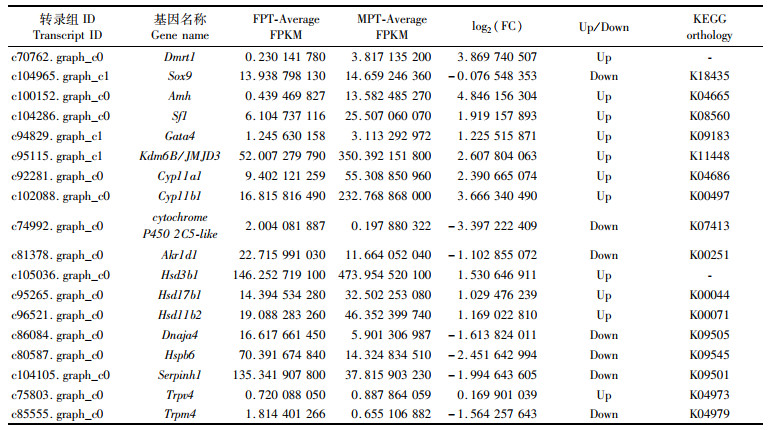
近年来,热休克蛋白(HSP)和瞬时受体电位(TRP)离子通道蛋白是TSD温度感受机制研究的关注热点。转录组数据检测到6个TRP家族的基因和7个HSP家族的基因,在16期MPT和FPT性腺转录水平上出现差异表达,显著性差异表达有6个(Trpv6、Trpm2、Trpm4、Dnaja4、Hspb6和Serpinh1),见表 4。TRP家族的差异表达基因中,与温度感受有关的Trpv2、Trpv4和Trpm4温度激活范围分别为>52 ℃、>27 ℃和15~35 ℃。其中Trpv4和Trpm4通道的温度激活范围与红耳龟孵化温度(26~32 ℃)相吻合,这提示Trpv4和Trpm4很可能介导红耳龟TSD的温度感受过程。
|
表 4 红耳龟雌雄性腺差异表达的部分基因 Tab.4 Several genes differentially expressed in male and female gonads of T. scripta |
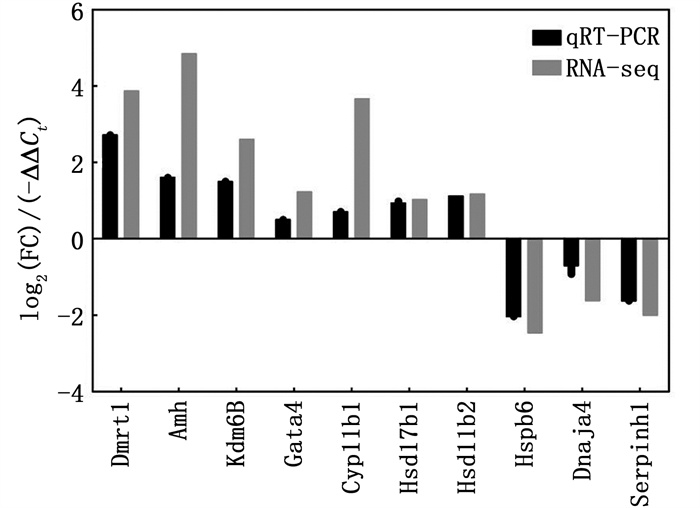
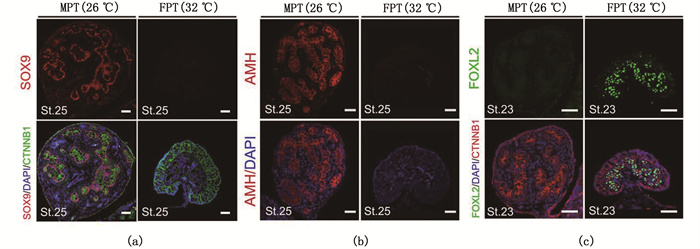
为确定转录组测序结果的可靠性,选取9个基因进行qPCR验证,结果表明:在16期两性性腺中,所选基因的mRNA表达水平与转录组测序结果相一致,且Dmrt1、Amh、Kdm6B、Gata4、Cyp11b1、Hsd17b1、Hsd11b2在MPT性腺中的表达水平较高,而Hspb6、Dnaja4、Serpinh1在FPT性腺中表达较高(图 6)。同样,SOX9、AMH和FOXL2蛋白的免疫荧光染色也表明:在25期MPT性腺中,SOX9和AMH蛋白大量表达,分别定位在髓质区支持细胞的细胞核和细胞质中,FPT性腺中则未检测到荧光信号(图 7a,7b)。相反,FOXL2在23期FPT性腺中大量表达,分布于髓质区体细胞细胞核,MPT性腺中未见表达,见图 7c。
|
图 6 部分差异表达基因的qRT-PCR与转录组的比较分析 Fig. 6 Comparative analysis of several selected differentially expressed genes by qRT-PCR and transcriptome |
|
标尺.50μm。 Scale bar. 50 μm. 图 7 3个差异表达基因的免疫组化验证 Fig. 7 Immunohistochemical verification of three differentially expressed genes |
动物性别决定和分化长期以来都是生物学的研究热点。自Sry雄性性别决定基因[31]被发现以来,GSD动物中其他性别决定基因如Dmy、Dm-w、Dmrt1、Amhy、Gsdfy、Amhr2陆续被研究报道[32-38],并获得许多阐述详尽的分子调控通路。在TSD动物中,GE等在红耳龟上发现组蛋白去甲基化酶Kdm6B通过去除Dmrt1启动子区的H3K27me3标记,直接激活Dmrt1转录,从而启动雄性性腺分化[39]。但TSD的机制研究仅仅局限在某几个物种上,研究深度远落后于GSD。本研究分析了红耳龟早期性腺转录组,筛选到474个性别相关DEGs,除报道较多的Dmrt1、Amh、Sox9和Sf1[37-38, 40-43]外,通过GO和KEGG分析发现Cyp11a1、Cyp11b1、Akr1d1等在“类固醇激素代谢合成”性别分化相关的通路中显著富集(图 5a),以及Trpv4、Trpm4、Dnaja4、Hspb6、Serpinh1等在热信号传导中呈差异表达(表 4)。因此,TSD机制很可能构成了一个极其复杂的网络,由多基因多途径共同调控性别决定和分化。本研究主要对类固醇激素代谢合成通路和温度敏感因子重点筛选分析。
类固醇激素代谢合成通路中,Cyp11a1作为控制类固醇生物合成的第一步和限速步骤,经垂体激素的控制,通过CAMP信号通路在肾上腺皮质和性腺中表达[44]。Sf1与Cyp11a1的启动子结合,在类固醇基因的组织特异性和激素调控表达中起重要的作用[45]。我们转录组数据显示Cyp11a1和Cyp11b1在MPT性腺中显著上调。MARTÍN等认为Cyp11a1在胡子鲇(Clarias batrachus)发育成熟的睾丸中高表达,可能参与调节硬骨鱼性腺外或循环类固醇水平[46]。Cyp11b1是编码由睾酮产生11β-羟基睾酮的关键酶,博纳里牙汉鱼(Odontes thesbonariensis)中已证实Cyp11b1表达与睾丸的形态分化呈正相关[47]。由此可推出,这两个基因具有促进睾丸分化的重要作用。转录组数据中Akr1d1表达下调(即FPT高表达),而CREWS等认为红耳龟、鸟类中Akr1d1在MPT表达是另一个潜在的雄性化机制[48],这与国外推测的研究结果相反。cytochrome P450 2C5-like涉及含血清素的神经突触、炎症介质对TRP通道的调节、类固醇类激素生物合成等过程,表明其参与着重要基础生命代谢合成过程。此外,转录组和qPCR虽检测到Hsd3b1、Hsb17b1、Hsd11b2的差异表达,但目前并没有这些基因参与性别调控过程的相关报道,有待进一步研究。
PIEAU[49]于1999年已提出热敏感基因能直接或间接激活芳香化酶,作用于GPCR型受体后,由蛋白激酶A使Trpv4敏感;另一种则由热休克蛋白参与雌激素受体的结合或解离发挥作用[50]。研究[50]表明TRP通道响应温度刺激形成钙离子流,降钙素能作为温度调节器调节哺乳动物的血清钙激素。因此,它作为介质使温度诱导性腺中的钙离子流,本研究结果显示,与钙离子有关的热敏感基因Trpv4、Trpm4呈差异表达,其温度激活范围分别为>27 ℃和15~35 ℃,与红耳龟孵化温度(26~32 ℃)相吻合[51-52],推测很可能通过温度激活下游靶基因参与调控决定性别和分化过程。本研究还发现热休克蛋白家族的Dnaja4、Hspb6、Serpinh1呈现差异表达。目前,大鳞大马哈鱼Oncorhynchus tshawytscha、虹鳟Oncorhynchus mykiss等鱼类中HSP70和HSP90的表达会随着水温变化而改变[53-54]。此外,HSP70已被证实能与Sox9a、Gata4和Lhx6一起参与罗非鱼的性腺发育过程,且HSP70在睾丸中呈高表达[55]。同样,猜测Dnaja4、Hspb6、Serpinh1很可能在红耳龟性别分化过程中也起到高温候选传感器的作用。综上,TRP蛋白编码基因和HSP热休克蛋白基因很可能响应温度通过钙调素或其他某种未知方式,直接调控下游靶基因或激素来影响性腺性别分化过程。
激素、热敏性基因与性别相关基因的网络尤为复杂,候选基因能否在性别分化过程中起到关键作用,还有待进一步证实。通过此次研究,我们筛选获得了与红耳龟性别有关的候选基因,结合相关通路和研究报道,分析并预测了性别分化的靶向基因,为进一步阐明红耳龟TSD机制提供基础数据。
4 总结本研究对红耳龟16期两性性腺进行了转录组测序,经GO注释和KEGG功能富集分析,发现雌雄差异表达转录本主要在“类固醇激素代谢合成通路”“PI3K-Akt信号通路”“cAMP信号通路”“TGF-β信号通路”“Wnt信号通路”“雌激素信号通路”“TRP通道”等与性别分化相关的通路中显著富集。所筛基因可能是爬行动物TSD机制研究的重要目标基因,包括Dmrt1、Sox9、Foxl2等高度保守的性别分化基因,Cyp11b1、Hsd11b2等类固醇激素代谢相关基因,以及Hspb6、Dnaja4、Serpinh1等热敏性蛋白编码基因,这为解析TSD分子机制提供了基础数据。
| [1] |
VALENZUELA N, ADAMS D C, JANZEN F J. Pattern does not equal process: exactly when is sex environmentally determined?[J]. The American Naturalist, 2003, 161(4): 676-683. DOI:10.1086/368292 |
| [2] |
SHOEMAKER C M, QUEEN J, CREWS D. Response of candidate sex-determining genes to changes in temperature reveals their involvement in the molecular network underlying temperature-dependent sex determination[J]. Molecular Endocrinology, 2007, 21(11): 2750-2763. DOI:10.1210/me.2007-0263 |
| [3] |
SHOEMAKER-DALY C M, JACKSON K, YATSU R, et al. Genetic network underlying temperature-dependent sex determination is endogenously regulated by temperature in isolated cultured Trachemys scripta gonads[J]. Developmental Dynamics, 2010, 239(4): 1061-1075. DOI:10.1002/dvdy.22266 |
| [4] |
MATSUMOTO Y, CREWS D. Molecular mechanisms of temperature-dependent sex determination in the context of ecological developmental biology[J]. Molecular and Cellular Endocrinology, 2012, 354(1/2): 103-110. |
| [5] |
LIU C F, LIU C, YAO H C. Building pathways for ovary organogenesis in the mouse embryo[J]. Current Topics in Developmental Biology, 2010, 90: 263-290. |
| [6] |
MUNGER S C, NATARAJAN A, LOOGER L L, et al. Fine time course expression analysis identifies cascades of activation and repression and maps a putative regulator of mammalian sex determination[J]. PLoS Genetics, 2013, 9(7): e1003630. DOI:10.1371/journal.pgen.1003630 |
| [7] |
AYERS K L, DAVIDSON N M, DEMIYAH D, et al. RNA sequencing reveals sexually dimorphic gene expression before gonadal differentiation in chicken and allows comprehensive annotation of the W-chromosome[J]. Genome Biology, 2013, 14(3): R26. DOI:10.1186/gb-2013-14-3-r26 |
| [8] |
SPOTILA L D, SPOTILA J R, HALL S E. Sequence and expression analysis of WT1 and Sox9 in the red-eared slider turtle, Trachemysscripta[J]. Journal of Experimental Zoology, 1998, 281(5): 417-427. DOI:10.1002/(SICI)1097-010X(19980801)281:5<417::AID-JEZ7>3.0.CO;2-R |
| [9] |
VALENZUELA N. Relic thermosensitive gene expression in a turtle with genotypic sex determination[J]. Evolution, 2007, 62(1): 234-240. DOI:10.1111/j.1558-5646.2007.00279.x |
| [10] |
VALENZUELA N, LECLERE A, SHIKANO T. Comparative gene expression of steroidogenic factor 1 in Chrysemys picta and Apalone mutica turtles with temperature-dependent and genotypic sex determination[J]. Evolution & Development, 2006, 8(5): 424-432. |
| [11] |
RAMSEY M, SHOEMAKER C, CREWS D. Gonadal expression of Sf1 and aromatase during sex determination in the red-eared slider turtle (Trachemys scripta), a reptile with temperature-dependent sex determination[J]. Differentiation, 2007, 75(10): 978-991. DOI:10.1111/j.1432-0436.2007.00182.x |
| [12] |
MALDONADO L C T, PIEDRA A L, MENDOZA N M, et al. Expression profiles of Dax1, Dmrt1, and Sox9 during temperature sex determination in gonads of the sea turtle Lepidochelys olivacea[J]. General and Comparative Endocrinology, 2002, 129(1): 20-26. DOI:10.1016/S0016-6480(02)00511-7 |
| [13] |
VALENZUELA N. Evolution of the gene network underlying gonadogenesis in turtles with temperature-dependent and genotypic sex determination[J]. Integrative and Comparative Biology, 2008, 48(4): 476-485. DOI:10.1093/icb/icn031 |
| [14] |
BARSKE L A, CAPEL B. Estrogen represses SOX9 during sex determination in the red-eared slider turtle Trachemys scripta[J]. Developmental Biology, 2010, 341(1): 305-314. DOI:10.1016/j.ydbio.2010.02.010 |
| [15] |
MATSUMOTO Y, YATSU R, TAYLOR C, et al. Changes in gonadal gene network by exogenous ligands in temperature-dependent sex determination[J]. Journal of Molecular Endocrinology, 2013, 50(3): 389-400. DOI:10.1530/JME-12-0260 |
| [16] |
VALENZUELA N. Multivariate expression analysis of the gene network underlying sexual development in turtle embryos with temperature-dependent and genotypic sex determination[J]. Sexual Development, 2010, 4(1/2): 39-49. |
| [17] |
KETTLEWELL J R, RAYMOND C S, ZARKOWER D. Temperature-dependent expression of turtle Dmrt1 prior to sexual differentiation[J]. Genesis, 2000, 26(3): 174-178. DOI:10.1002/(SICI)1526-968X(200003)26:3<174::AID-GENE2>3.0.CO;2-J |
| [18] |
SHOEMAKER C M, CREWS D. Analyzing the coordinated gene network underlying temperature-dependent sex determination in reptiles[J]. Seminars in Cell and Developmental Biology, 2009, 20(3): 290-303. |
| [19] |
MATSUMOTO Y, BUEMIO A, CHU R, et al. Epigenetic control of gonadal aromatase(cyp19a1) in temperature-dependent sex determination of red-eared slider turtles[J]. PLoS One, 2013, 8(6): e63599. DOI:10.1371/journal.pone.0063599 |
| [20] |
VALENZUELA N, SHIKANO T. Embryological ontogeny of aromatase gene expression in Chrysemy spicta and Apalone muticaturtles: comparative patterns within and across temperature-dependent and genotypic sex-determining mechanisms[J]. Development Genes and Evolution, 2007, 217(1): 55-62. DOI:10.1007/s00427-006-0106-3 |
| [21] |
SCHROEDER A L, METZGER K J, MILLER A, et al. A novel candidate gene for temperature-dependent sex determination in the common snapping turtle[J]. Genetics, 2016, 203(1): 557-571. DOI:10.1534/genetics.115.182840 |
| [22] |
LANCE V A. Is regulation of aromatase expression in reptiles the key to understanding temperature-dependent sex determination?[J]. Journal of Experimental Zoology Part A: Ecological Genetics and Physiology, 2009, 311A(5): 314-322. DOI:10.1002/jez.465 |
| [23] |
NAKAMURA M. The mechanism of sex determination in vertebrates-are sex steroids the key-factor?[J]. Journal of Experimental Zoology Part A: Ecological Genetics and Physiology, 2010, 313A(7): 381-398. DOI:10.1002/jez.616 |
| [24] |
WARNER D A, RADDER R S, SHINE R. Corticosterone exposure during embryonic development affects offspring growth and sex ratios in opposing directions in two lizard species with environmental sex determination[J]. Physiological and Biochemical Zoology, 2009, 82(4): 363-371. DOI:10.1086/588491 |
| [25] |
PIFERRER F. Epigenetics of sex determination and gonadogenesis[J]. Developmental Dynamics, 2013, 242(4): 360-370. DOI:10.1002/dvdy.23924 |
| [26] |
PARROTT B B, KOHNO S, CLOY-MCCOY J A, et al. Differential incubation temperatures result in dimorphic DNA methylation patterning of the SOX9 and aromatase promoters in gonads of alligator (Alligator mississippiensis) embryos[J]. Biology of Reproduction, 2014, 90(1): 2. |
| [27] |
NAVARRO-MARTIN L, VIÑAS J, RIBAS L, et al. DNA methylation of the gonadal aromatase (cyp19a) promoter is involved in temperature-dependent sex ratio shifts in theEuropean sea bass[J]. PLoS Genetics, 2011, 7(12): e1002447. DOI:10.1371/journal.pgen.1002447 |
| [28] |
CASTELLI M A, WHITELEY S L, GEORGES A, et al. Cellular calcium and redox regulation: the mediator of vertebrate environmental sex determination?[J]. Biological Reviews, 2020, 95(3): 680-695. DOI:10.1111/brv.12582 |
| [29] |
RADHAKRISHNAN S, LITERMAN R, NEUWALD J, et al. Transcriptomic responses to environmental temperature by turtles with temperature-dependent and genotypic sex determination assessed by RNAseq inform the genetic architecture of embryonic gonadal development[J]. PLoS One, 2017, 12(3): e0172044. DOI:10.1371/journal.pone.0172044 |
| [30] |
GREENBAUM E. A standardized series of embryonic stages for the emydid turtle Trachemys scripta[J]. Canadian Journal of Zoology, 2002, 80(8): 1350-1370. DOI:10.1139/z02-111 |
| [31] |
KOOPMAN P, GUBBAY J, VIVIAN N, et al. Male development of chromosomally female mice transgenic forSry[J]. Nature, 1991, 351(6322): 117-121. DOI:10.1038/351117a0 |
| [32] |
MATSUDA M, NAGAHAMA Y, SHINOMIYA A, et al. DMY is a Y-specific DM-domain gene required for male development in the medakafish[J]. Nature, 2002, 417(6888): 559-563. DOI:10.1038/nature751 |
| [33] |
MYOSHO T, OTAKE H, MASUYAMA H, et al. Tracing the emergence of a novel sex-determining gene in medaka, Oryzias luzonensis[J]. Genetics, 2012, 191(1): 163-170. DOI:10.1534/genetics.111.137497 |
| [34] |
YOSHIMOTO S, OKADA E, UMEMOTO H, et al. A W-linked DM-domain gene, DM-W, participates in primary ovary development in Xenopuslaevis[J]. Proceedings of the National Academy of Sciences of the United States of America, 2008, 105(7): 2469-2474. DOI:10.1073/pnas.0712244105 |
| [35] |
SMITH C A, ROESZLER K N, OHNESORG T, et al. The avian Z-linked gene Dmrt1 is required for male sex determination in the chicken[J]. Nature, 2009, 461(7261): 267-271. DOI:10.1038/nature08298 |
| [36] |
CAPEL B. Vertebrate sex determination: evolutionary plasticity of a fundamental switch[J]. Nature Reviews Genetics, 2017, 18(11): 675-689. DOI:10.1038/nrg.2017.60 |
| [37] |
BISHOP C E, WHITWORTH D J, QIN Y J, et al. A transgenic insertion upstream of Sox9 is associated with dominant XX sex reversal in the mouse[J]. Nature Genetics, 2000, 26(4): 490-494. DOI:10.1038/82652 |
| [38] |
VALENZUELA N, NEUWALD J L, LITERMAN R. Transcriptional evolution underlying vertebrate sexual development[J]. Developmental Dynamics, 2013, 242(4): 307-319. DOI:10.1002/dvdy.23897 |
| [39] |
GE C T, YE J, WEBER C, et al. The histone demethylase KDM6B regulates temperature-dependent sex determination in a turtle species[J]. Science, 2018, 360(6389): 645-648. DOI:10.1126/science.aap8328 |
| [40] |
EGGERS S, OHNESORG T, SINCLAIR A. Genetic regulation of mammalian gonad development[J]. Nature Reviews Endocrinology, 2014, 10(11): 673-683. DOI:10.1038/nrendo.2014.163 |
| [41] |
KUROKI S, MATOBA S, AKIYOSHI M, et al. Epigenetic regulation of mouse sex determination by the histone demethylase Jmjd1a[J]. Science, 2013, 341(6150): 1106-1109. DOI:10.1126/science.1239864 |
| [42] |
MORRISH B C, SINCLAIR A H. Vertebrate sex determination: many means to an end[J]. Reproduction, 2002, 124(4): 447-457. DOI:10.1530/rep.0.1240447 |
| [43] |
KOHNO S, PARROTT BB, YATSU R, et al. Gonadal differentiation in reptiles exhibiting environmental sex determination[J]. Sexual Development, 2014, 8(5): 208-226. DOI:10.1159/000358892 |
| [44] |
TOIT R L D, STORBECK K H, CARTWRIGHT M, et al. Progestins used in endocrine therapy and the implications for the biosynthesis and metabolism of endogenous steroid hormones[J]. Molecular and Cellular Endocrinology, 2017, 441: 31-45. DOI:10.1016/j.mce.2016.09.004 |
| [45] |
GUO I C, HU M C, CHUNG B C. Transcriptional regulation of CYP11A1[J]. Journal of Biomedical Science, 2003, 10(6): 593-598. |
| [46] |
RAJAKUMAR A, SENTHILKUMARAN B. Expression analysis of cyp11a1 during gonadal development, recrudescence and after hCG induction and sex steroid analog treatment in the catfish, Clarias batrachus[J]. Comparative Biochemistry and Physiology Part B: Biochemistry and Molecular Biology, 2014, 176: 42-47. DOI:10.1016/j.cbpb.2014.07.007 |
| [47] |
BLASCO M, FERNANDINO J I, GUILGUR L G, et al. Molecular characterization of cyp11a1 and cyp11b1 and their gene expression profile in pejerrey (Odontes thesbonariensis) during early gonadal development[J]. Comparative Biochemistry and Physiology Part A: Molecular & Integrative Physiology, 2010, 156(1): 110-118. |
| [48] |
CZERWINSKI M, NATARAJAN A, BARSKE L, et al. A timecourse analysis of systemic and gonadal effects of temperature on sexual development of the red-eared slider turtle Trachemys scripta elegans[J]. Developmental Biology, 2016, 420(1): 166-177. DOI:10.1016/j.ydbio.2016.09.018 |
| [49] |
PIEAU C, DORIZZI M, RICHARD-MERCIER N. Temperature-dependent sex determination and gonadal differentiation in reptiles[J]. Cellular and Molecular Life Sciences CMLS, 1999, 55(6): 887-900. |
| [50] |
DARBY W G, GRACE M S, BARATCHI S, et al. Modulation of TRPV4 by diverse mechanisms[J]. The International Journal of Biochemistry and Cell Biology, 2016, 78: 217-228. DOI:10.1016/j.biocel.2016.07.012 |
| [51] |
MARTíNEZ-JUÁREZ A, LÓPEZ-LUNA M A, PORRAS-GÓMEZ T J, et al. Expression of the Sox9, Foxl2, Vasa, and TRPV4 genes in the ovaries and testes of the Morelet's crocodile, Crocodylus moreletii[J]. Journal of Experimental Zoology Part B: Molecular and Developmental Evolution, 2018, 330(3): 148-164. DOI:10.1002/jez.b.22799 |
| [52] |
MONTELL C. The TRP superfamily of cationchannels[J]. Science's STKE, 2005(272): re3. |
| [53] |
PALMISANO A N, WINTON J R, DICKHOFF W W. Tissue-specific induction of Hsp90 mRNA and plasma cortisol response in Chinook salmon following heat shock, seawater challenge, and handling challenge[J]. Marine Biotechnology, 2000, 2(4): 329-338. DOI:10.1007/s101260000005 |
| [54] |
NAKANO K, IWAMA G K. The 70-kDa heat shock protein response in two intertidal sculpins, Oligocottus maculosus and O. snyderi: relationship of hsp70 and thermal tolerance[J]. Comparative Biochemistry and Physiology Part A: Molecular & Integrative Physiology, 2002, 133(1): 79-94. |
| [55] |
TAOW J, CHEN J L, TAND J, et al. Transcriptome display during tilapia sex determination and differentiation as revealed by RNA-Seqanalysis[J]. BMC Genomics, 2018, 19(1): 363. DOI:10.1186/s12864-018-4756-0 |
2. National Demonstration Center for Experimental Fisheries Science Education, Shanghai Ocean University, Shanghai 201306, China
 2022,
Vol. 31
2022,
Vol. 31


