文章信息
- 安东, 袁正伟
- AN Dong, YUAN Zhengwei
- 利用同位素标记相对和绝对定量技术筛查神经管畸形胚胎孕鼠血清差异蛋白质谱
- Identification of Differentially Expressed Proteins in Sera of Pregnant Rats with Fetuses Having Neural Tube Defects Using Isobaric Tag Technology for Relative and Absolute Quantitation
- 中国医科大学学报, 2019, 48(4): 315-319, 323
- Journal of China Medical University, 2019, 48(4): 315-319, 323
-
文章历史
- 收稿日期:2018-03-27
- 网络出版时间:2019-04-13 14:59
2. 中国医科大学附属盛京医院卫生部小儿先天畸形重点实验室, 沈阳 110004
2. Key Laboratory of Health Ministry for Congenital Malformation, Shengjing Hospital, China Medical University, Shenyang 110004, China
神经管畸形(neural tube defects,NTDs)是儿童常见的畸形,发病率为1‰~5‰[1]。70%~80%的NTDs可能通过产前筛查手段如超声及孕妇血清甲胎蛋白(alpha fetoprotein,AFP)筛查出来,但目前AFP等产前检查标记物的特异性并不高[2-3]。孕期胎儿和孕妇之间存在相互交换,其交换物质包括细胞内、细胞外因子[4-5]。血清蛋白谱变化能够反应孕妇和胎儿的生理和病理变化。因此,研究孕妇血清蛋白谱变化规律对畸形的早期发现和研究畸形的发生机制具有十分重要的意义。
近年来,蛋白质组学研究对疾病的早期诊断、发病机制探讨有重要意义[6]。本课题组已用蛋白质组学方法对先天性显性脊柱裂(spina bifida aperta,SBA)胎鼠的脊髓组织及羊水进行了蛋白质表达谱的分析,结果发现了新的特异性的差异表达蛋白[7-8]。本研究采用同位素标记相对和绝对定量(isobaric tag for relative and absolute quantitation,iTRAQ)技术[9],利用维甲酸致畸动物模型,研究比较SBA致畸早期孕11、13 d(E11、E13)与正常孕鼠血清蛋白表达谱的变化,以筛选出多种与NTDs相关的差异表达蛋白,为寻找NTDs早期血清蛋白标志物及探讨发病机制奠定了基础。
1 材料与方法 1.1 研究对象成熟未孕Wistar大鼠(体质量250~300 g)由中国医科大学附属盛京医院实验动物中心提供,雌雄比例5:1午夜合笼,晨8:00阴道涂片镜下见到精子记为孕0 d(E0)。将孕10 d(E10)孕鼠随机分为2组,致畸组以维甲酸混悬液(140 mg/kg)经胃管灌饲,对照组给予同体积的橄榄油。孕鼠分别于E11、E13行水合氯醛腹腔麻醉,心脏穿刺取血,剖宫取出胎鼠,解剖显微镜下辨认胎鼠是否为BA。血标本用无抗凝剂的5 mL普通采血试管,室温放置2 h,使血液凝固析出血清,3 000 g离心20 min,冰上分装于1.5 mL的EP管中,每管200 μL,-80 ℃保存。iTRAQ实验分组为维甲酸致畸组(E11、E13)和正常对照组(E11、E13),每组5例血清预混。本研究符合中国医科大学实验动物伦理委员会所制定的伦理学标准。
1.2 方法 1.2.1 蛋白质提取与浓度测定将致畸组和对照组各组内5个样本等体积混合为一个样本,用Proteo miner试剂盒去除血清高丰度蛋白,使用Brandford方法进行蛋白浓度测量。
1.2.2 iTRAQ标记、SCX分离与液相串联质谱分析每个样品取出蛋白100 μg,胰蛋白酶酶解消化,按照操作手册进行iTRAQ标记。E11正常对照组(E11N),标记114标签;E11维甲酸致畸组(E11SBA),标记117标签;E13正常对照组(E13N),标记118标签;E13维甲酸致畸组(E13SBA),标记121标签。室温培养2 h,标记后的各组肽段混合。采用岛津LC-20AB液相系统、分离柱为4.6 mm×250 mm型号的Ultremex SCX柱对样品进行液相分离,分离后组分经Triple TOF 5600的液相色谱电离串联质谱(LC-ESI-MSMS)分析采集数据。
1.3 质谱数据分析及生物信息学分析应用Mascot检索软件,选择数据库Uniprot_RAT(34908sequences)进行数据库搜索,对蛋白质进行鉴定。根据蛋白质丰度水平,当差异倍数 > 1.5,且经统计检验其P < 0.05时,视为致畸组与正常对照组之间的表达差异蛋白。对鉴定出的差异蛋白,通过检索Gene Ontology数据库和COG数据库,进行GO生物学功能富集分析和COG功能分类分析。
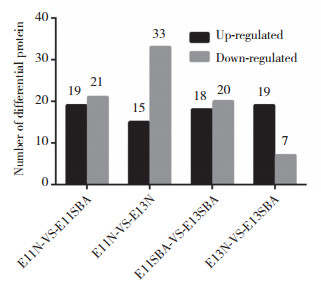
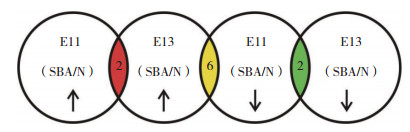
2 结果 2.1 鉴定得到差异蛋白本研究共得到谱图 157 422张,通过Mascot软件进行分析后,匹配到的谱图数量7 303张,其中Unique谱图数量为6 013张,共鉴定到2 167个肽段,其中含1 880个Unique肽段和390个蛋白。对E11、E13维甲酸致畸组与正常对照组差异蛋白(表 1)、E11与E13正常对照组差异蛋白、E11与E13维甲酸致畸组差异蛋白4组资料进行两两比较,共发现显著差异表达的152个蛋白(图 1)。在这些有差异表达的蛋白中,维甲酸致畸组与正常对照组比较,蛋白APOM和PCSK9在E11和E13同时下调,蛋白FGG和Uncharacterized在E11和E13同时上调,蛋白PLA2g2a、FGl2、SERPINE2、TUBB5、FGl1和HP在E11下调而在E13上调,见图 2。
| Accession | Uniprot_swissprot description | Mascot score | % of coverage | Fold differences | |
| 11SBA/11N | 13SBA/13N | ||||
| IPI00205036 | Hemoglobin subunit alpha-1/2 | 335 | 46.5 | 11.717 | - |
| IPI00231192 | Hemoglobin subunit beta-2 | 547 | 62.6 | 6.090 | - |
| IPI00230897 | Hemoglobin subunit beta-1 | 770 | 57.8 | 4.095 | - |
| IPI00205389 | Fibrinogen beta chain | 279 | 21.7 | 3.970 | - |
| IPI00190759 | Fibrinogen gamma chain | 183 | 18.9 | 3.623 | 1.648 |
| IPI00190622 | Muellerian-inhibiting factor | 35 | 1.1 | 3.308 | - |
| IPI00202651 | Fibrinogen alpha chain | 428 | 16.1 | 2.793 | - |
| IPI00187799 | Kininogen-1 | 913 | 34.9 | 2.401 | - |
| IPI00199497 | Leukemia inhibitory factor receptor | 529 | 17.7 | 2.385 | - |
| IPI00569842 | Ig heavy chain V region J558 | 106 | 19.1 | 2.210 | - |
| IPI00564154 | Ig heavy chain V regions TEPC 15/S107/HPCM1/HPCM2/HPCM3 | 115 | 20.8 | 2.177 | - |
| IPI00782787 | Ig gamma-2C chain C region | 262 | 17.0 | 2.053 | - |
| IPI00205022 | Nucleobindin-1 | 103 | 10.2 | 1.960 | - |
| IPI00363901 | Immunoglobulin J chain | 372 | 41.5 | 1.959 | - |
| IPI00211401 | Matrix Gla protein | 86 | 23.3 | 1.872 | - |
| IPI00568091 | Ig kappa chain V19-17 | 194 | 30.3 | 1.794 | - |
| IPI00557643 | Ig heavy chain V region T601 | 207 | 19.8 | 1.683 | - |
| IPI00365922 | Ig heavy chain V region MOPC 47A | 358 | 51.3 | 1.543 | 2.077 |
| IPI00565300 | Ig heavy chain V-Ⅲ region VH26 | 441 | 47.9 | 1.515 | - |
| IPI00197780 | Growth-regulated protein homolog alpha | 126 | 15.3 | - | 3.744 |
| IPI00382202 | Haptoglobin | 265 | 21.5 | 0.557 | 2.670 |
| IPI00205248 | Phospholipase A2, membrane associated | 380 | 42.5 | 0.267 | 2.508 |
| IPI00393058 | Multimerin-1 | 310 | 11.1 | - | 2.459 |
| IPI00324102 | Fibroleukin | 253 | 14.9 | 0.278 | 2.141 |
| IPI00205465 | Phospholipase A1 member A | 129 | 18.2 | 2.131 | |
| IPI00876637 | Glia-derived nexin | 234 | 18.9 | 0.293 | 2.053 |
| IPI00361346 | Ig gamma-2A chain C region | 1 545 | 33.6 | - | 1.888 |
| IPI00959940 | Ig heavy chain V region PJ14 | 32 | 12.6 | - | 1.856 |
| IPI00198887 | Protein disulfide-isomerase | 118 | 6.7 | - | 1.837 |
| IPI00325135 | 14-3-3 protein epsilon | 153 | 14.5 | - | 1.836 |
| IPI00189795 | Tubulin alpha-1A chain | 503 | 38.1 | 0.314 | 1.671 |
| IPI00201347 | Histidine-rich glycoprotein | 351 | 12.0 | - | 1.606 |
| IPI00201262 | Alpha-1-inhibitor 3 | 1 475 | 28.1 | - | 1.587 |
| IPI00203721 | Fibrinogen-like protein 1 | 117 | 15.3 | 0.449 | 1.570 |
| IPI00197579 | Tubulin beta chain | 178 | 15.1 | 0.341 | 1.535 |
| IPI00203319 | Ficolin-1 | 304 | 22.4 | - | 1.521 |
| IPI00768167 | Tubulin beta-1 chain | 122 | 11.5 | 0.153 | - |
| IPI00896162 | Fermitin family homolog 3 | 165 | 6.6 | 0.236 | - |
| IPI00210071 | Coronin-1A | 64 | 4.1 | 0.264 | - |
| IPI00555171 | Transgelin-2 | 125 | 22.1 | 0.309 | - |
| IPI00214905 | Tropomyosin alpha-4 chain | 265 | 13.3 | 0.332 | - |
| IPI00362014 | Talin-1 | 474 | 7.6 | 0.362 | - |
| IPI00211075 | Serine protease inhibitor A3N | 1 123 | 49.3 | 0.395 | - |
| IPI00197684 | Xaa-Pro aminopeptidase 2 | 79 | 7.3 | 0.502 | - |
| IPI00396889 | Proprotein convertase subtilisin/kexin type 9 | 448 | 16.2 | 0.535 | 0.639 |
| IPI00207275 | Apolipoprotein M | 494 | 19.5 | 0.536 | 0.561 |
| IPI00421302 | Hyaluronan-binding protein 2 | 300 | 14.6 | 0.588 | - |
| IPI00194097 | Vitamin D-binding protein | 1 083 | 38.0 | 0.625 | - |
| IPI00210947 | Heparin cofactor 2 | 1 972 | 53.7 | 0.663 | - |
| IPI00199368 | C-type lectin domain family 11 member A | 147 | 18.6 | 0.665 | - |
| IPI00190501 | Carboxypeptidase B2 | 280 | 21.8 | - | 0.498 |
| IPI00190500 | Carboxypeptidase N catalytic chain | 165 | 16.2 | - | 0.608 |
| IPI00948319 | Complement component C8 alpha chain | 879 | 32.0 | - | 0.612 |
| IPI00778224 | Complement factor H-related protein 4 | 221 | 11.0 | - | 0.613 |
| IPI00192657 | Carboxypeptidase N subunit 2 | 510 | 30.4 | - | 0.665 |
|
| E, embryonic day; N, normal; SBA, spina bifida aperta. 图 1 各比较组间差异蛋白数量统计 Fig.1 Comparison between the differentially expressed serum proteins in pregnant rats with SBA fetuses and normal controls at E11 and E13 |
|
| The yellow region includes 6 proteins (sPLA2, fibroleukin, GDN, tubulin beta chain, fibrinogen-like protein 1, and haptoglobin) that were upregulated at E13 and downregulated at E11. The green region includes 2 proteins (APOM, PCSK9) that were downregulated at both E11 and E13. The red region includes 2 proteins (fibrinogen gamma chain and Ig heavy chain V region MOPC 47A) that were upregulated at both E11 and E13. 图 2 差异蛋白表达交集图 Fig.2 Differentially expressed serum proteins |
2.2 生物信息学分析 2.2.1 GO注释
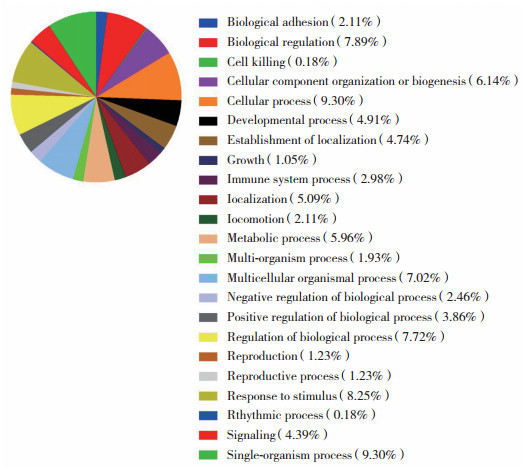
对差异蛋白质进行GO分析,发现其主要参与的生物学功能包括细胞进程(9.30%)、单有机体过程(9.30%)、应激反应(8.25%)、生物功能调节(7.89%)、发育过程(4.91%)和信号转导(4.39%),见图 3。
|
| 图 3 GO分类图(参与生物学功能) Fig.3 Gene ontology annotation based on biological functions |
2.2.2 COG注释
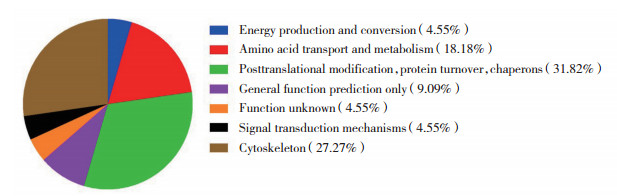
COG注释,差异蛋白主要参与蛋白质翻译后修饰、折叠、分子伴侣和细胞骨架功能,见图 4。
|
| 图 4 COG分类图 Fig.4 COG annotation |
3 讨论
目前,比较蛋白质组学应用于出生缺陷研究,在寻找产前诊断标记物、探索胚胎畸形发生机制和治疗靶点等方面已做了大量研究。iTRAQ技术是先进的定量比较蛋白质组学方法之一,优点有定量敏感、标记完全、标记效率高。在本研究中,采用iTRAQ相关技术对脊柱裂动物模型致畸早期的孕鼠血清总蛋白进行鉴定和比较蛋白组学相对定量分析,共鉴定了390个蛋白,其中152个蛋白在不同组间出现显著差异表达。结果表明,将iTRAQ标记串联质谱技术用于NTDs孕鼠血清的比较蛋白组学分析是一种行之有效的方法。
维甲酸致畸组与正常对照组相比,E11鉴定差异表达的蛋白为40个,远远多于E13鉴定差异表达的蛋白26个,说明致畸早期有更多的蛋白分子参与异常神经发育过程。在这些差异蛋白中,部分蛋白曾在NTDs、神经疾病、胚胎发育畸形的蛋白质组学研究中报道。如应用双向电泳和质谱分析技术,对正常对照组和维甲酸致畸组孕17 d胎鼠脊髓组织的蛋白质组进行分析,发现了差异蛋白14-3-3ε蛋白、蛋白二硫化物异构酶(protein disulphide-isomerase,PDI)家族[8];在唐氏综合征孕妇血清中筛查出标志物富组氨酸糖蛋白(histidine-richglycoprotein)和维生素D结合蛋白(vitamin D-binding protein)[10-11];脐血中发现新生儿特异蛋白Transgelin-2[12];阿尔兹海默病脑脊液筛查出基质Gla-protein前体和结合珠蛋白前体;帕金森病脑脊液筛查出载脂蛋白M和维生素D结合蛋白前体[13]。提示iTRAQ标记技术用于蛋白质相对定量的比较研究有可比性和延续性。用iTRAQ标记技术对致畸早期E11、E13维甲酸致畸孕鼠血清进行相对定量分析,共鉴定出38个在畸形组中表达显著上调的蛋白和28个表达显著下调的蛋白。这些致畸早期差异蛋白可能参与NTDs的发生过程,有望作为NTDs产前诊断的标志物。
在血清学中鉴定到的差异蛋白,从其GO分类可知这些蛋白质参与细胞进程、应激反应、生物功能调节、发育过程、信号转导等诸多方面的生物过程。所鉴定的差异蛋白中,部分是功能已知与畸形明确相关的分子,如14-3-3ε蛋白和PDI。14-3-3ε是哺乳动物神经系统正常发育所必需的一种蛋白质,该蛋白缺失或缺陷时,会发生脑死亡或严重大脑畸形[14]。PDI蛋白具有蛋白质转录后修饰、协助蛋白质折叠、装配、运输功能,进而抑制细胞凋亡、保护细胞、减少应激对器官造成的伤害等多种作用[15-16]。此外,还有很多新发现的蛋白,其中部分蛋白参与胚胎的正常发育。对这些蛋白进行进一步的验证分析和深入的体内外实验和功能验证研究,将有助于阐明这些蛋白参与和调控脊柱裂发生机制的方式,为NTDs产前诊断与新的靶向治疗打下基础。
| [1] |
ZAGANJOR I, SEKKARIE A, TSANG BL, et al. Describing the prevalence of neural tube defects worldwide:a systematic literature review[J]. PLoS One, 2016, 11(4): e0151586. DOI:10.1371/journal.pone.0151586 |
| [2] |
杨岚, 赵丽, 江静颖, 等. 孕中期血清学筛查在产前诊断及妊娠结局预测中的应用[J]. 南方医科大学学报, 2015, 35(7): 1059-1062, 1072. DOI:10.3969/j.issn.1673-4254.2015.07.25 |
| [3] |
KRANTZ DA, HALLAHAN TW, SHEWIN JE. Screening for open neural tube defects[J]. Clin Lab Med, 2010, 30(3): 721-725. DOI:10.1016/j.cll.2010.04.010 |
| [4] |
CHOOLANI M, NARASIMHAN K, KOLLA V, et al. Proteomic technologies for prenatal diagnostics:advances and challenges ahead[J]. Expert Rev Proteomics, 2009, 6(1): 87-101. DOI:10.1586/14789450.6.1.87 |
| [5] |
HERR F, BAAL N, ZYGMUNT M. Studies of placental vasculogenesis:a way to understand pregnancy pathology?[J]. Z Geburtshilfe Neonatol, 2009, 213(3): 96-100. DOI:10.1055/s-0029-1224141 |
| [6] |
ASLAM B, BASIT M, NISAR MA, et al. Proteomics:technologies and their applications[J]. J Chromatogr Sci, 2017, 55(2): 182-196. DOI:10.1093/chromsci/bmw167 |
| [7] |
SHAN L, FAN Y, LI H, et al. Proteomic analysis of amniotic fluid of pregnant rats with spinabifida aperta[J]. J Proteomics, 2012, 75(4): 1181-1189. DOI:10.1016/j.jprot.2011.10.033 |
| [8] |
FAN Y, WANG L, ZHOU F, et al. Comparative proteomics of spinal cords of rat fetuses with spina bifida aperta[J]. J Proteomics, 2011, 75(2): 668-676. DOI:10.1016/j.jprot.2011.09.006 |
| [9] |
WONG YK, ZHANG J, HUA ZC, et al. Recent advances in quantitative and chemical proteomics for autophagy studies[J]. Autophagy, 2017, 13(9): 1472-1486. DOI:10.1080/15548627.2017.1313944 |
| [10] |
NAGALLA SR, CANICK JA, JACOB T, et al. Proteomic analysis of maternal serum in down syndrome:identification of novel protein biomarkers[J]. J Proteome Res, 2007, 6(4): 1245-1257. DOI:10.1021/pr060539h |
| [11] |
GRAVETT MG, NOVY MJ, ROSENFELD RG, et al. Diagnosis of intra-amniotic infection by proteomic profiling and identification of novel biomarkers[J]. JAMA, 2004, 292(4): 462-469. DOI:10.1001/jama.292.4.462 |
| [12] |
SONG HJ, ZHANG P, GUO XJ, et al. The proteomic analysis of human neonatal umbilical cord serum by mass spectrometry[J]. Acta Pharmacol Sin, 2009, 30(11): 1550-1558. DOI:10.1038/aps.2009.140 |
| [13] |
ABDI F, QUINN JF, JANKOVIC J, et al. Detection of biomarkers with a multiplex quantitative proteomic platform in cerebrospinal fluid of patients with neurodegenerative disorders[J]. J Alzheimers Dis, 2006, 9(3): 293-348. DOI:10.3233/JAD-2006-9309 |
| [14] |
TOYO-OKA K, SHIONOYA A, GAMBELLO MJ, et al. 14-3-3epsilon is important forneuronal migration by binding to NUDEL:a molecular explanation for Miller-Dieker syndrome[J]. Nat Genet, 2003, 34(3): 274-285. DOI:10.1038/ng1169 |
| [15] |
PERRI E, PARAKH S, ATKIN J. Protein disulphide isomerases:emerging roles of PDI and ERp57 in the nervous system and as therapeutic targets for ALS[J]. Expert Opin Ther Targets, 2017, 21(1): 37-49. DOI:10.1080/14728222.2016.1254197 |
| [16] |
SOARES MORETTI AI, MARTINS LAURINDO FR. Protein disulfide isomerases:redox connections in and out of the endoplasmic reticulum[J]. Arch Biochem Biophys, 2017, 617: 106-119. DOI:10.1016/j.abb.2016.11.007 |
 2019, Vol. 48
2019, Vol. 48




