Nassariidae comprised about 300 extant and almost 600 extinct nassariid species. Nassarius, a species-rich genus of Nassariinae, is distributed throughout worldwide oceans and are usually less than 50 mm in adult shell height (Cernohorsky, 1972; Zou et al., 2012). Most nassariids of Nassarius inhabited inter- to subtidal shallow marine environments. As scavengers, nassariids of Nassarius is important in maintaining the balance of ecological system, especially for the balance of benthic community. However, food safety problem exists in Nassarius. As economic species, most Nassarius species are consumed as food in China and other Asian countries. However, eating nassariids of Nassarius have caused hundreds of food poisoning incidents in the last several years in China. It is frequently reported that many people died of the poisoning incidents (Lin et al., 2015).
It has been indicated that the food poisoning incident is due to the different toxins concentrated in Nassarius sp's body. Some studies detected that the toxins were tetrodotoxin (TTX) and homologues (Zhang et al., 2012). Nevertheless, the origin of the toxins is still not clear. It was inferred that the TTX was from food chain or bacteria in their body (Wang, 2008). Recent studies found that the toxicity of Nassarius was related to species and populations (Xu et al., 2007; Zhang et al., 2009), which indicated that N. hepaticus was toxic gastropods and N. festiva was non-toxic gastropods while the toxicity of N. succinctus and N. variciferus probably changed with the different sea areas (Zhang et al., 2009). The reason why the toxicity of Nassarius samples changes with the species, populations and different sea areas can greatly contribute to control the poisoning incidents. The possible reason of TTX-resistance in Nassarius may be that the species and populations evolve differently in genetic level (Moczydlowski, 2013).
However, so far, the identification of Nassarius species is still confusing, especially for the toxic species. Since the intraspecific shell variation and the various shell forms often lead to the difficulty of Nassarius species discrimination it is arbitrary to identify Nassarius species just based on morphological characters (Haasl, 2000). Thus, it is significant to combine the toxicity detection and the species assignment to determine the toxicity of Nassarius in species and population level, especially based on the molecular tool. Although our previous study revealed some cryptic diversity in Nassarius based on some gene loci the toxicity of the nassariids was not tested. Thus, the species assignment of the various toxic and nontoxic Nassarius is still not clear at the molecular level.
In this study, we detected the toxicity and the toxin components of Nassarius samples, then used several gene loci (COI, 16S and ITS) to assign the Nassarius species and populations and analyzed their phylogenetic relationship. We aim to reveal the relationship of toxicity and species in Nassarius at the genetic level.
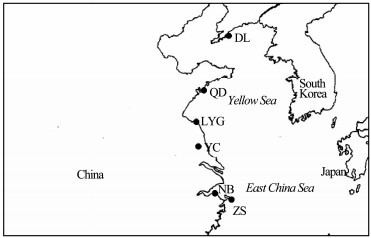
2 Materials and Methods 2.1 Sample Collection and Identification by Morphological CharactersNassarius samples were collected from the China coast, including Liaoning, Shandong, Jiangsu and Zhejiang provinces from 2015.6 to 2015.9 (Fig. 1, Table S1). Both toxic and nontoxic species were included, e.g. N. succinctus, N. semiplicatus, N. variciferus. In addition, the samples were collected from different seasons. Firstly, the samples were identified by morphological characters, and then the identification is further confirmed by molecular sequences.
|
Fig. 1 The locations along the North China coast where the Nassarius were collected. DL, Dalian; QD, Qingdao; LYG, Lianyungang; YC, Yancheng; NB, Ningbo; ZS, Zhoushan. |
Firstly, the mud was washed from the nassariid bodies and the whole tissue was taken out by smashing the shell. Then the tissue was blended. 1 g mixed tissue was added into 5 mL HCL solution (0.1 mol L-1). The PH of the mixed HCL solution was then adjusted to 2-4. After heating and cooling, the PH of the mixed HCL solution was adjusted to 2-4 again. Then, the mixed HCL solution was adjusted to 10 mL using 0.1 mol L-1 HCL. Finally, the intermixture was centrifuged for 15 min under 6000 r min-1, and the supernatant was got for toxicity test.
The toxicity test was conducted by intraperitoneal injection of mice, and the toxicity calculation followed the method in Hwang et al., (1991). The Kunming male mice weighting 18-21 g was used for toxicity test. 1 mL toxin supernatant was injected into the abdomen of the mice. The reaction of the mice was observed and the death time was recorded. The Mouse Unit was used for toxicity (1 mouse unit means that the TTX could kill the mouse within 30 minutes).
2.3 Toxicity Type TestSome representative toxic Nassarius samples were collected for testing the toxin. The toxin content was tested by the method of TTX and homologues with High Performance Liquid Chromatography (HPLC).
The condition of Chromatographic column is: Agilent Zorbax SB C18 (250 mm × 4.6 mm × 5 μm), Agilent C18, Detection wave te: 196 nm, column temperature: room temperature, mobile phase: mixture of 8 mmol L-1 sodium heptanesulfonate and 0.005%TFA. The pH of mobile phase was adjusted to 5.0 by NaOH.
TTX standard (1 g) was dissolved by 0.1% HAc since TTX is easily dissolved in acid and slightly dissolved in water, which was diluted to 10, 20, 25, 50, 80, and 400 μg mL-1 respectively.
Tissue extraction of Nassarius samples was filtered with 0.22 μm filter membrane. Finally 500 μL tissue extraction was tested.
2.4 Molecular Identification and Phylogenetic AnalysisDNA was extracted from small pieces of foot tissue by the CTAB method as modified by Winnepenninckx et al. (1993). PCR reactions were carried out in HotstarTaq Master Mix Kit, Qiagen. PCR and sequencing primers for COI, 16S rRNA and ITS-1 genes followed Zou et al. (2012). The PCR products were confirmed by 1.5% agarose gel electrophoresis and purified using EZ Spin Column PCR Product Purification Kit, Sangon. Purified products were sequenced in both directions using the BigDye Terminator Cycle Sequencing Kit (Ver. 3.1, Applied Biosystems) and an AB PRISM 3730 (Applied Biosystems) automatic sequencer.
Forward and reverse sequences of COI, 16S and ITS genes were edited, assembled and merged into consensus sequences using the software program Sequencher 4.5 (Genecodes Corporation, Ann Arbor, MI). Sequences were aligned by MAFFT 6.717 (Katoh et al., 2009). Phylogenetic analysis of COI, 16S and ITS sequences were conducted based on Neighbour Joining (NJ), Maximum Likelihood (ML) and Bayesian methods. The species Fusinus longicaudus, Euplica scripta, Mitrella burchardi and Pseudamycla formosa were selected as the outgroups as our previous studies (Zou et al., 2012). NJ analyses were conducted using K2P distance model in MEGA 7.0 (Kumar et al., 2016). Bayesian analyses were carried out following the information in Zou et al. (2012). ML analyses were conducted using RAxML (v7.0.4, Stamatakis, 2006) executed on the CIPRES server (Miller et al., 2009). Rapid nonparametric bootstrapping with 1000 replicates was used to evaluate the branch support (Stamatakis et al., 2008). Nucleotide substitution models were selected separately using the Akaike Information Criterion (AIC) as implemented in the jModeltest (v.0.1.1, Posada, 2008).
Some published COI, 16S and ITS data of Nassarius in our previous study Zou et al. (2012) were also included in the assignment analysis.
3 Results 3.1 ToxicityThe toxicity of 64 samples were tested using Kunming male mice. It was indicated that the toxicity varied among different species and different populations (Table S1). The two species N. festivus and N. conoidalis were nontoxic species, all samples of which from different sea regions were nontoxic. But different populations within N. succinctus, N. semoplicatus and N. variciferus had different toxicity. In N. succinctus, the samples from Lianyungang and Ningbo had high toxicity while the samples from Lalian and Qingdao were nontoxic. In N. variciferus, while the samples from Lianyungang and Ningbo were toxic the samples from Qingdao, Dalian and Rizhao were nontoxic.
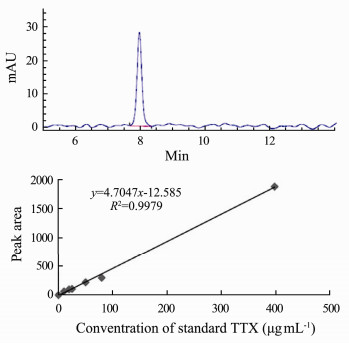
3.2 Toxin DetectionThe TTX standard chromatomap and canonical plotting detected by HPLC are shown in Fig. 2. It was indicated that the Agilent Zorbax had good retention effect for TTX. The retention time is about 8.05 min. The TTX concentration and signal had good linear relation while TTX standard toxin concentration was between 1-400 µg mL-1 (R2 = 0.9979).
|
Fig. 2 The TTX standard chromatomap and canonical plotting detected by HPLC. |
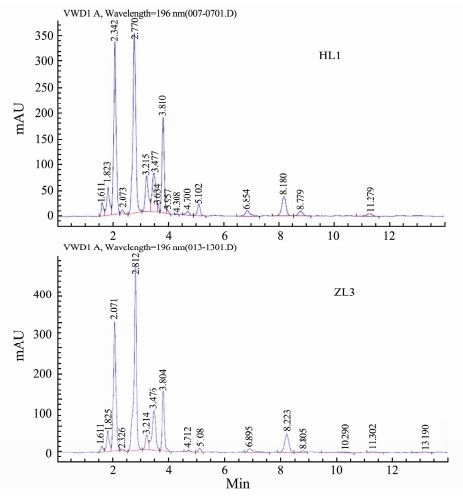
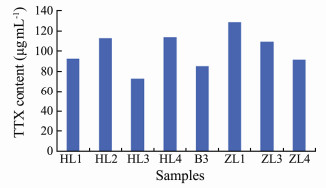
Some representative toxic N. succinctus, N. semoplicatus and N. variciferus samples were selected for TTX content detection. The test indicated that the toxin in Nassarius samples was TTX. The representative chromatomap of HL1 and ZL3 samples are shown in Fig. 3. The toxicity varied in different species or different populations. For example, the TTX content in N. succinctus (HL1) was 87 µg mL-1, the TTX content in N. succinctus (HL2) was 118 µg mL-1, and the TTX content in N. variciferus (ZL3 and ZL4) was 110 µg mL-1 and 90 µg mL-1 respectively (Fig. 4).
|
Fig. 3 The TTX chromatomap of HL1, HL2, ZL3 and ZL4 samples. |
|
Fig. 4 The TTX content detected in representative Nassarius samples. |
The COI, 16S and ITS sequences of all different toxic Nassarius samples were analyzed by NJ, ML and Bayesian analysis respectively. The Genbank number of COI, 16S and ITS were among MG679290-MG679358, and MH113174-MH113272. We also included the previous published COI, 16S and ITS data of Nassarius in the identification analysis, which could assign the various toxic samples more correctly. The Buccinum pemphigus was chosen as the outgroup in the COI and 16S analysis. Since there were no published Buccinum pemphigus ITS sequences the ITS data in this study was analyzed without outgroup. The most appropriate models for ML and Bayesian analyses were GTR + G for COI, TVMef+I+G for 16S, and GTR for ITS. And the molecular taxonomy assignment and the phylogenetic relationships of COI, 16S and ITS are all showed in Figs. 5, 6 and 7 respectively.
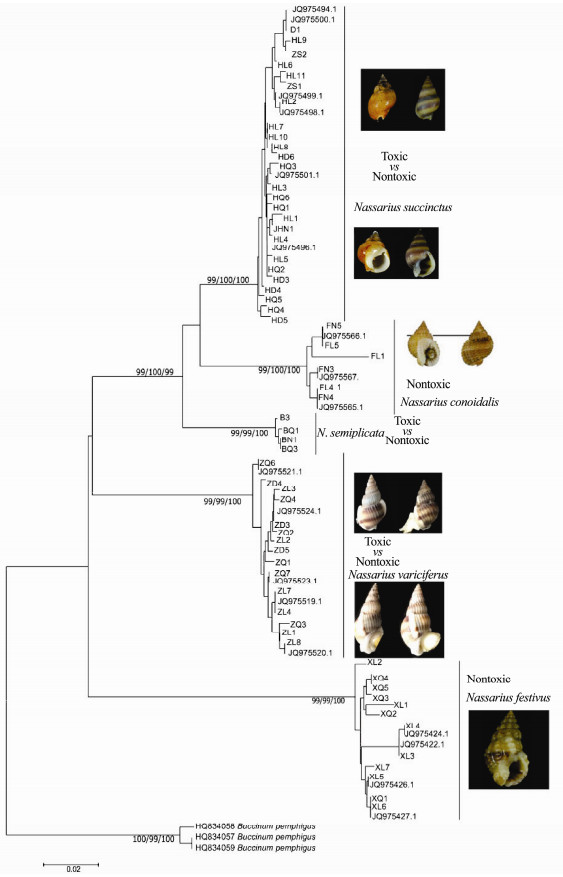
|
Fig. 5 Bayesian tree of the COI gene sequences. Left to right, maximum likelihood bootstraps/NJ bootstraps and posterior probabilities. |
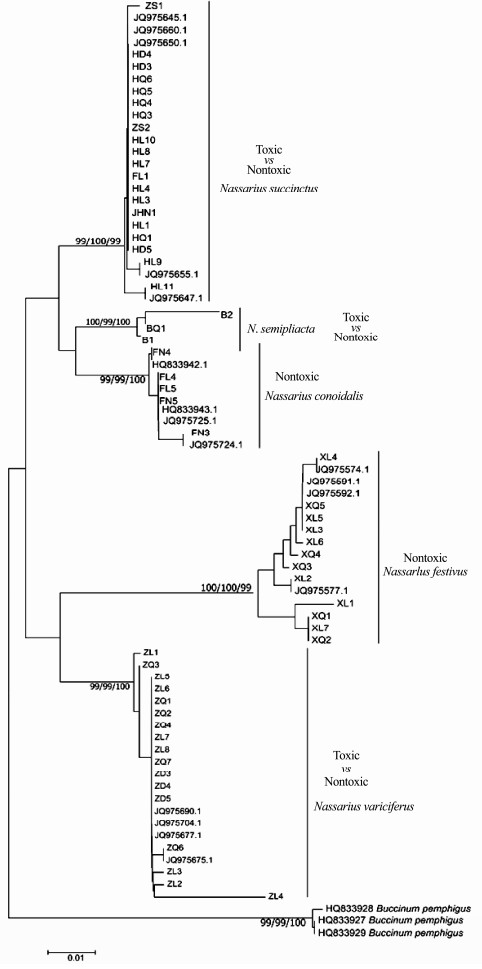
|
Fig. 6 Bayesian tree of the 16S gene sequences. Left to right, maximum likelihood bootstraps/NJ bootstraps and posterior probabilities. |
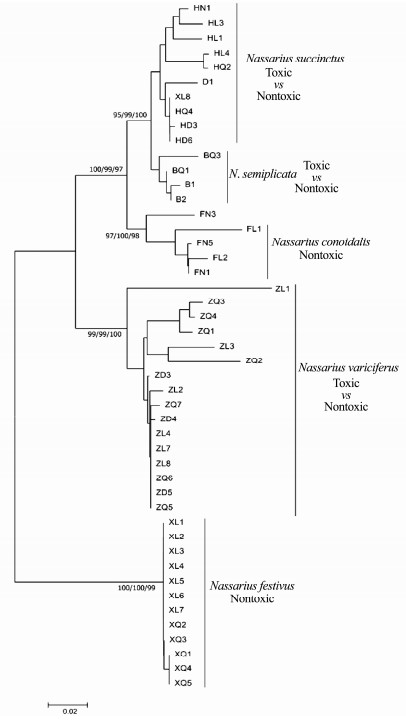
|
Fig. 7 Bayesian tree of the ITS gene sequences. Left to right, maximum likelihood bootstraps/NJ bootstraps and posterior probabilities. |
It was indicated that the five Nassarius species, N. variciferus, N. conoidalis, N. semiplicata, N. succinctus and N. festivus, were clearly separated in NJ, ML and Bayesian trees of COI, 16S and ITS genes (Figs. 5, 6 and 7). The published sequence data of each species also clustered together with the sequences got in this study. This indicated that the different toxic species could be distinguished clearly by multiple gene sequences. However, the different toxic populations of N. succinctus and N. variciferus could not be separated from each other, which just clustered together. For example, for both N. succinctus and N. variciferus, the toxic populations from Lianyungang and the nontoxic populations from Dalian and Qingdao were not distinguished. Additionally, the toxic and nontoxic species did not cluster together separately.
4 DiscussionConsumption of gastropod Nassarius spp. caused many poisoning incidents in the last several decades in China since Nassarius (Gastropoda, Nassariidae) could accumulate tetrodotoxin (TTX) (Xu et al., 2007; Zhang et al., 2009). It has been proved that the toxicity of Nassarius is related to species and population. Nevertheless, the identification of Nassarius is extremely difficult, which could deteriorate the danger of eating Nassarius samples. Accurate taxonomy assignment of different toxic Nassarius samples is important for controlling the poison incidents and is helpful to well understand the adaptational evolution of toxin-resistance (Bricelj et al., 2005). In this study, we combined the toxicity, toxin detection and the molecular sequences to determine the species assignment of different toxic Nassarius samples and analyze the potential phylogenetic relationships among them.
First of all, we tested the toxicity of five Nassarius species from the north China coast. The result indicated that different species and different populations within one species had various toxicity. N. festiva and N. conoidalis were nontoxic species, and the different populations within N. succinctus, N. variciferus and N. semiplicata had different toxicity. The toxin was confirmed to be TTX, which was consistent with the previous studies (Xu et al., 2007; Luo, 2008). The toxin content also varied among different samples, which may be the reason that when people eat Nassarius samples some people are safe and some people may die. For both N. succinctus and N. variciferus, the populations from Lianyungang and Ningbo had serious toxicity, but the populations from Qingdao and Dalian had no toxicity. This can be an important case for studying the adaptational evolution of TTX-resistance in Nassarius to the environmental changes. The origin of the TTX is still not clear. Some studies showed that the TTX might come from the food chain in the environment (Luo, 2008; Zhang et al., 2010). The microalgae composition in the food chain varies in different environment, which could explain that the Nassarius samples have different toxicity. However, the mechanism of the TTX resistance for toxic Nassarius samples is unknown. Some studies have indicated that the resistance to TTX is due to the amino acid mutation in sodium channel genes or the protein compound with the TTX (Geffeney et al., 2005; Venkatesh et al., 2005; Jost et al., 2008; Hanifin & Gilly, 2015). The Nassarius could be as ideal case to study the mechanism of TTX-resistance in Mollusca.
Our study firstly combined the toxicity detection and genetic tools to identify the various toxic species and study their potential phylogenetic relationships. The molecular results showed that the different Nassarius species detected were separated as monophyletic clades clearly by the COI, 16S and ITS sequences in the phylogenetic trees. The different toxic Nassarius samples have been assigned to species level by multiple gene loci, which was also consistent with our previous species assignment since the published sequence data in Zou et al. (2012) were included. However, the toxic and nontoxic species did not form two separated clades, and the toxic and nontoxic populations within N. succinctus, N. variciferus and N. semiplicata were not separated. These showed that the toxicity of Nassarius is really related with species, but not absolutely related with populations in genetic level. Our molecular identification of various toxic Nassarius samples provided the basis for controlling poisoning accidents and studying the adaptational evolution of TTX-resistance in genetic mutation level. Many studies found that the adaptation of the TTX-resistance could be due to the mutation of the amino acid in the sodium channel, e.g., the snakes (Feldman et al., 2012). Future studies about the amino acid mutation of sodium channel among the different toxic Nassarius samples can greatly contribute to the toxin-resistance study in species adaptation evolution. Our study provided the important species assignment among the different toxic Nassarius samples for the research of adaptation evolution.
5 ConclusionsDifferent toxic Nassarius samples were collected from the China coast. The toxicity detection indicated that different species and populations have different toxicity. The toxin in Nassarius was confirmed to be TTX. The COI, 16S and ITS genes assign the different toxic Nassarius samples to species level, but the different toxic populations within one species was not separated. This showed that the toxicity of Nassarius is really related with species at the genetic level, but not absolutely related with populations. As a whole, our study constructed the relationship between toxicity and taxonomy assignment of different toxic Nassarius samples at genetic level, and provided basic information for controlling poisoning and researching about TTX-resistance mechanism in mollusk.
Supporting Information
|
|
Table S1 The Nassarius toxicity test conducted by intraperitoneal injection of mice |
The support of the National Natural Science Foundation of China (No. 31600294), the Fundamental Research Funds for the Central University (No. KYZ201627) and the Fundamental Research Funds for the Central University (No. KJQN201742) is gratefully acknowledged.
Bricelj, V. M., Connell, L., Konoki, K., Macquarrie, S. P., Scheuer, T., Catterall, W. A. and Trainer, V. L., 2005. Sodium channel mutation leading to saxitoxin resistance in clams increases risk of PSP. Nature, 434: 763-767. DOI:10.1038/nature03415 (  0) 0) |
Cernohorsky, W. O., 1972. Indo-Pacific nassariidae (Mollusca: Gastropoda). Records of the Auckland Institute & Museum, 9: 125-194. (  0) 0) |
Feldman, C. R., Brodie, E. D. and Pfrender, M. E., 2012. Constraint shapes convergence in tetrodotoxin-resistant sodium channels of snakes. Proceedings of the National Academy of Sciences of the United States of America, 109: 4556-4561. (  0) 0) |
Geffeney, S. L., Fujimoto, E., Brodie, E. D. and Ruben, P. C., 2005. Evolutionary diversification of TTX-resistant sodium channels in a predator-prey interaction. Nature, 434: 759-763. DOI:10.1038/nature03444 (  0) 0) |
Haasl, D. M., 2000. Phylogenetic relationships among Nassariid gastropoda. Journal of Vertebrate Paleontology, 74: 839-852. DOI:10.1017/S0022336000033047 (  0) 0) |
Hanifin, C. T. and Gilly, W. F., 2015. Evolutionary history of a complex adaptation: Tetrodotoxin resistance in salamanders. Evolution, 69: 232-244. DOI:10.1111/evo.12552 (  0) 0) |
Hwang, D. F. and Jeng, S. S., 1991. Bioassay of tetrodotoxin by using ICR strain male mouse. Journal of the Chinese Chemical Society, 20: 80-86. (  0) 0) |
Jost, M. C., Hillis, D. M., Lu, Y., Kyle, J. W., Fozzard, H. A. and Zakon, H. H., 2008. Toxin-resistant sodium channels: Parallel adaptive evolution across a complete gene family. Molecular Biology and Evolution, 25(6): 1016-1024. DOI:10.1093/molbev/msn025 (  0) 0) |
Katoh, K., Asimenos, G. and Toh, H., 2009. Multiple alignment of DNA sequences with MAFFT. Methods in Molecular Biology, 537: 39-64. (  0) 0) |
Kumar, S., Stecher, G. and Tamura, K., 2016. MEGA7: Molecular evolutionary genetics analysis Version 7.0 for bigger datasets. Molecular Biology and Evolution, 33: 1870-1874. (  0) 0) |
Lin, X. Y., Lin, J., Zhang, X. Y. and Cai, X. X., 2015. Epidemiologocal investigation of several accidental poisoning caused by nassarius. Chinese Journal of Health Laboratory Technology, 25: 1646-1646. (  0) 0) |
Luo, X., 2008. Toxicity dynamics and toxin composition of Nassarius spp. along the coast of China. PhD thesis. Institute of Oceanology, Chinese Academy of Sciences.
(  0) 0) |
Miller, M., Holder, M., Vos, R., Midford, P., Liebowitz, T., Chan, L., Hoover, P. and Warnow, T., 2009. The molecular mystique of tetrodotoxin. Toxicon, 63: 165-183. (  0) 0) |
Posada, D., 2008. JModelTest: Phylogenetic model averaging. Molecular Biology and Evolution, 25: 1253-1256. DOI:10.1093/molbev/msn083 (  0) 0) |
Stamatakis, A., 2006. RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics, 22: 2688-2690. (  0) 0) |
Stamatakis, A., Hoover, P. and Rougemont, J., 2008. A rapid bootstrap algorithm for the RAxML web servers. Systematic Biology, 57: 758-771. (  0) 0) |
Tamura, K., Dudley, J., Nei, M. and Kumar, S., 2007. Mega 4: Molecular evolutionary genetics analyses (mega) software version 4.0. Molecular Biology and Evolution, 24: 1596-1599. (  0) 0) |
Venkatesh, B., Lu, S. Q., Dandona, N., See, S. L., Brenner, S. and Soong, T. W., 2005. Genetic basis of tetrodotoxin resistance in pufferfishes. Current Biology, 15: 2069-2072. DOI:10.1016/j.cub.2005.10.068 (  0) 0) |
Wang, X. J., 2008. Studies on the origin of tetrdtoxin in gastropod (Nassarius spp.). PhD thesis. Chinese Academy of Sciences.
(  0) 0) |
Winnepenninckx, B., Backeljau, T. and De, Wachter, R, 1993. Extraction of high molecular weight DNA from molluscs. Trends Genetics, 9: 407. DOI:10.1016/0168-9525(93)90102-N (  0) 0) |
Xu, J., Xu, G., Chen, Y., Qin, P. and Yu, M., 2007. Correlation between toxicity of poisonous Nassarius sp. and their habitats. Chinese Journal of Health Laboratory Technology, 17: 63-67. (  0) 0) |
Zhang, N., Liu, H., Sun, J., Ye, S., and Li, L., 2012. Tetrodotoxin and its derivatives in gastropods (Nassarius). 2: 220-224.
(  0) 0) |
Zhang, N., Su, J., Liu, H., Ye, S. and Li, L., 2009. The species and toxicities of Nassariidae collected from the coast of Southeast China Sea. Asian Journal of Ecotoxicology, 4: 289-294. (  0) 0) |
Zhang, N., Su, J., Liu, H. and Li, L., 2010. Study on source of toxin in Nassarius semiplicatus. Marine Environmental Research, 29: 705-707. (  0) 0) |
Zou, S., Li, Q. and Kong, L., 2012. Monophyly, distance and character-based multigene barcoding reveal extraordinary cryptic diversity in Nassarius: A complex and dangerous community. PLoS One, 7: e47276. DOI:10.1371/journal.pone.0047276 (  0) 0) |
 2019, Vol. 18
2019, Vol. 18


