2) Laboratory for Marine Biology and Biotechnology, Qingdao National Laboratory for Marine Science and Technology, Qingdao, 266237, China;
3) Ningde Marine Environment Monitoring Center, Ningde, 352100, China;
4) Key Laboratory of Marine Ecological Monitoring and Restoration Technology, Shanghai, 200137, China;
5) East China Sea Branch, Ministry of Natural Resources, Shanghai, 200137, China
The sea potato (Acaudina spp.), a species of sea cucumber, widely distributes in the waters along some Asian counties including Philippine and China. Most of these species live in the intertidal zone, and only a few of them live in the coastal sediment. The sea potato has simple body structure with a fast reproductive ability. Every summer, it becomes blooming in the East Sea of China, causing serious problems on the maintenance and operation of marine facilities. For example, large numbers of adults and larvae of sea potato caused the blockage of the pipes for seawater intake and drainage for nuclear power plants. It is necessary to carry out species identification and establish the risk evaluation system for this species. The classic species identification is mainly based on morphology observation. However, this approach is hard to identify the larvae of sea potato from a large number of plankton. The molecular methods based on the mitochondria genome have become an effective way to solve this problem.
Mitochondria are subcellular organelles unique to eukaryotes, playing an important role in a series of cytological processes, such as energy metabolism (Brand, 1997; Morenosánchez et al., 2010), cellular aging and diseases (Wallace et al., 1995). Mitochondrial genome data have been utilized to study phylogeographic (Scribner et al., 2003), evolution and phylogenetic relationships of the animals in the metazoans (Boore et al., 2005). Mitochondrial DNA (mtDNA) is a double-stranded circular DNA molecule that is independent of the chromosome. Its size is about 16 kb (Boore, 1999). Except for a few species, most of animals' mitochondrial genomes consist of 37 genes: 13 protein-coding genes, 2 ribosomal RNAs (rRNAs), 22 transfers RNAs (tRNAs) (Boore and Brown, 1994; Peregrino-Uriarte et al., 2009), and a non-coding putative control region, in which signal sequences exist for transcription and replication (Takata et al., 2001). mtDNA is essential for the process of protein synthesis, transcription, and translation. In addition, it has the characteristics of the small genome (Behera et al., 2018), compact structure, high copy number (Ingman et al., 2000) and no rearrangement during cell meiosis (Fan et al., 2011). It is widely used in evolution analysis, genetic diversity study and species identification (Curole and Kocher, 1999). The full mitochondrial sequences of most species of the sea cucumber have been determined (Shen et al., 2009; Perseke et al., 2010).
In our previous study, the mitochondrial DNA sequence of Acaudina spp. has been detected through polymerase chain reaction (PCR) amplification and Sanger sequencing (Wang et al., 2019). In this study, based on the complete mitochondrial genome sequences, we screened and identified the partial sequence of nd1 gene as a specific molecular marker for the identification of Acaudina spp. from the collected plankton samples. With the molecular marker, we developed an effective method to identify and evaluate the population dynamic of Acaudina spp. through PCR.
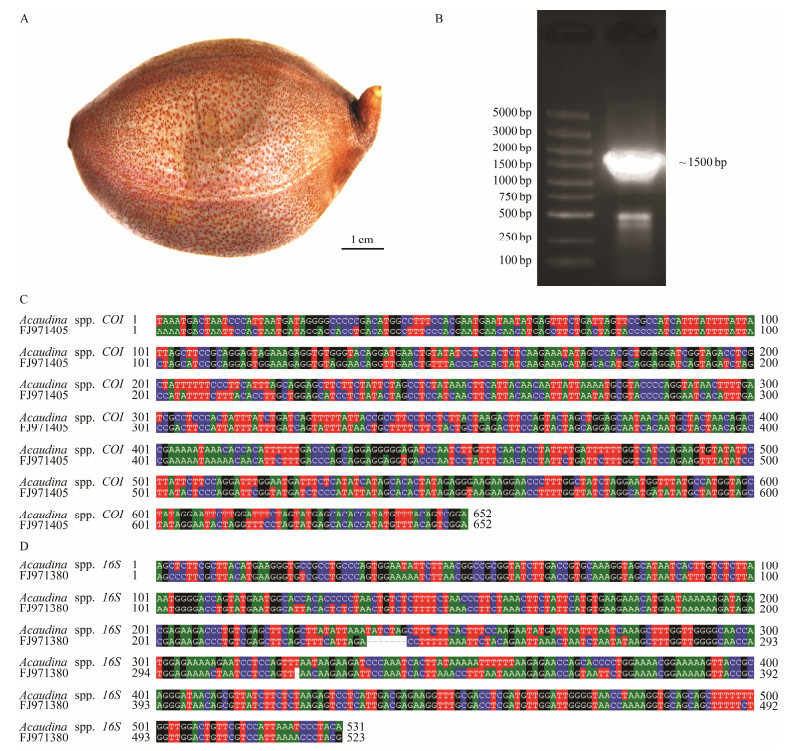
2 Materials and Methods 2.1 Animal CollectionFour adults of Acaudina spp. (Fig. 1A) were collected from the coast in Fujian Province, China. Species were identified by morphology (Xiao, 2015) and 16S rRNA sequencing based on the published sequences (Wen et al., 2011). The samples were then fixed in 75% ethanol and stored at 4℃ until DNA extraction.
|
Fig. 1 Morphological and molecular identification of sea potato Acaudina spp. (A) Image of adult Acaudina spp.. Bar is 1 cm. (B) Polymerase chain reaction (PCR) product with about 1500 bp size. The length of the markers from top to bottom is 5000 bp, 3000 bp, 2000 bp, 1500 bp, 1000 bp, 750 bp, 500 bp, 250 bp, and 100 bp, respectively. (C) Sequence alignment between amplicon and Am-COI (FJ971405). The identity is 85%. (D) Sequence alignment between amplicon and Am-16S (FJ971380). The identity is 90%. |
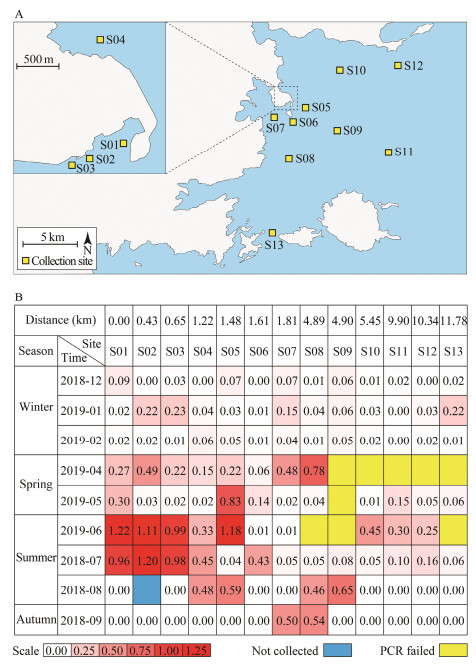
The larvae of Acaudina spp. were collected using a type Ⅱ plankton collection net (diameter 31.6 cm, length 140 cm, mesh size 0.16 mm, rope length 10 m) from 13 stations. The plankton samples were concentrated in a 500 mL plastic sampling bottle and then fixed with 75% ethanol. The positions of sampling stations were indicated in Fig. 3A.
|
Fig. 3 Quantitative evaluation of the density of Acaudina spp. larvae. (A) Sampling stations in this study (B) Relative quantity of Acaudina spp. larval density. Relative densities of Acaudina spp. larvae are indicated through a gradient change from white to red. White color indicates lower density of larvae. Red color indicates higher density of the larvae. Blue color indicates that the sample was not collected. Yellow color indicates failures of PCR reaction (no band on the gel). |
Total genomic DNA was extracted from the gonad of adult Acaudina spp. and the planktonic samples, respectively, using a modified phenol/chloroform/isoamyl alcohol method (Wei et al., 2020). Firstly, 45 mL STE buffer (100 mmol L-1 NaCl, 10 mmol L-1 Tris-HCl, 100 mmol L-1 EDTA, pH 8.0) and 4.5 mL 10% SDS (dissolved in ddH2O) were mixed in a 50 mL tube to make the lysis buffer. The gonads were dissected from the animal and were put into a 1.5 mL tube. Then 700 μL lysis buffer and 2.8 μL Proteinase K (Merck, dissolve in ddH2O with the concentration of 50 mg mL-1) were added into the tube to make a final concentration of 200 μg mL-1 proteinase K. The mixture was shaken gently and incubated in a water bath at 58℃ for 3 h. Finally, DNA was extracted with phenol-chloroform-isoamyl alcohol and chloroform-isoamyl alcohol, respectively, and then precipitated by alcohol and dissolved in ddH2O. The dissolved DNA was stored at -20℃.
2.3 PCR AmplificationPCR reaction was carried out using PfuS DNA polymerase (gift from Dr. Zhiyi Lv) in a 50 μL volume. The reaction solution includes 10 μL 5× Phusion HF Buffer (Thermo fisher Catalog number: F518L), 1 μL 10 mmol L-1 dNTP, 2.5 μL 10 μmol L-1 Primer F, 2.5 μL 10 μmol L-1 Primer R, 1 μL template DNA (200 ng μL), 0.5 μL PfuS DNA polymerase, and 32.5 μL ddH2O. The PCR was performed as follows: Pre-denaturation at 95℃ for 3 min; denaturation at 95℃ for 15 s; annealing at 55℃ for 15 s, followed with elongation at 72℃ for 1 min, and totally for 35 cycles; and a final extension at 72℃for 5 min. The PCR products were purified by Gene JET Gel Extraction Kit (Thermo Fisher Scientific, Lithuanian).
The primers for the amplification of nd1 from Acaudina spp. were designed based on the published mitogenomes (Wang et al., 2019). The primer sequences and annealing temperatures of PCR were shown in Table 1.
|
|
Table 1 Primer sequences and annealing temperatures |
TOPO cloning was performed using pEasy-Blunt3 kit (Transgen, Beijing, China). Firstly, 4 μL purified PCR product was mixed with 1 μL pEasy-Blunt3 vector and incubated at 25℃ for 15 min. Then they were transformed into Trans1-T1 competent cells and spray on Luria-Bertani (LB) agar plates. The LB plates were cultured overnight at 37℃. Monoclonal colonies were picked up for sequencing. Sequencing was performed by Genwiz company (Tianjin, China). Multiple sequence alignments were carried out using the ClustalW program in BioEdit software (version 7.0.4.1) with default parameters.
2.5 Image Analysis and Statistical AnalysesTotally 116 plankton samples were examined by PCR to determine whether the larvae of Acaudina spp. were in the samples. PCR products were detected on a 1% agarose gel. The electrophoresis bands of 116 samples were analyzed using ImageJ software to calculate the gray value, which can reflect the content of DNA. 18S rDNA (also known as 18S rRNA gene) was used as a reference. The distribution of Acaudina spp. larvae at 13 detected stations were evaluated based on molecular data.
3 Results 3.1 Morphological and Molecular Identification of Acaudina spp.One adult sample was collected from the coast of Fujian province. It was about eight cm long and its body presented the light brown color with dark brown spots (Fig. 1A). They were morphologically similar to sea potato. To identify the species, we designed primers 16S rRNA-F and co1-R to amplify a mitochondrial fragment including two genes 16S rRNA and cox based on previously published sequences (FJ971405 and FJ971380) of Acaudina spp. (Wen et al., 2011). A 1500 bp DNA fragment was amplified from the total DNA (Fig. 1B). After sequencing and alignment, both COI and 16S rRNA genes from our samples showed high identify with the ones from Acaudina spp.. However, the identity of COI sequence from our samples with FJ971405 was only 85% (Fig. 1C), and the 16S rRNA sequence from our samples showed 90% identity with FJ971380 (Fig. 1D). Based on these data, we identified our samples as Acaudina spp..
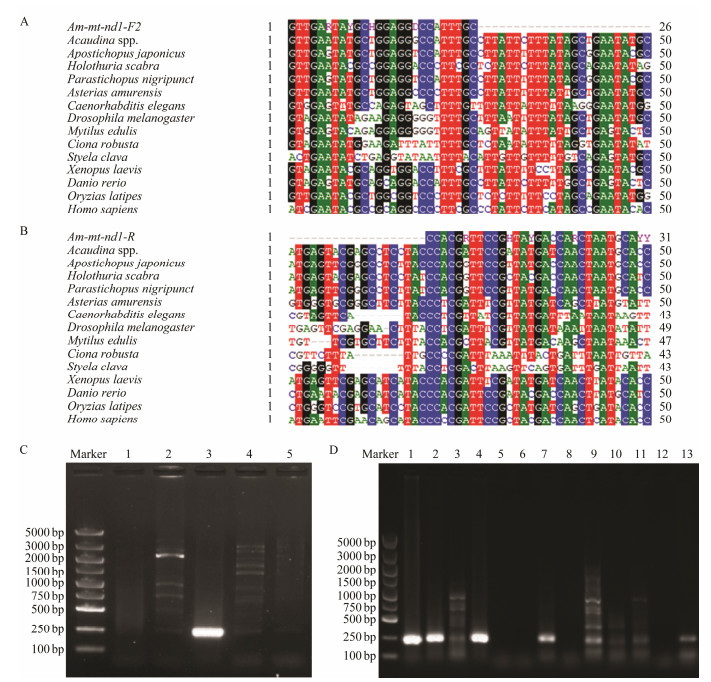
3.2 The Fragment of Mitochondrial nd1 Gene Was an Effective Species-Specific Molecular MarkerMitochondrial genes are widely used in population identification and genetic studies, such as atp6 (Das et al., 2018), co1 (Wang et al., 2018), cytb (Behera et al., 2015). In our previous work, we got the mitochondrial genome sequence of Acaudina spp.. Based on this data, we designed primers to amplify different fragments from the total DNA of Acaudina spp. and other marine species to identify the species-specific marker. The results showed that a 229 bp DNA fragment of nd1 gene could be effectively amplified through a primers pair, Am-mt-nd1-F2 (Fig. 2A) and Am-mt-nd1-R (Fig. 2B) from the total DNA of Acaudina spp., but could not from other marine species including ascidians (Ciona robusta, Styela clava), clam (Mactra quadrangularis) and scallop (Patinopecten yessoensis) (Fig. 2C). We then tested the validity of this pair of primers using total DNA extracted from the collected plankton samples. The results showed that one specific fragment could be successfully amplified from 9 of 13 samples (Fig. 2D), suggesting that the fragment of nd1 gene is an effective species-specific molecular marker for species identification and larva detection from the plankton samples.
|
Fig. 2 Screen of species-specific molecular for the identification of Acaudina spp. (A) Alignment of the forward primer with the nd1 sequences from different species. (B) Alignment of the reverse primers with the nd1 sequences from different species. (C) The amplificated bands were presented in 1% agarose gel using the nd1 gene primers and the total DNA from different marine species. M, DL5000 marker; 1, C. robusta, 2, S. clava, 3, Acaudina spp., 4, M. quadrangularis, 5, P. yessoensis. (D) The amplificated bands were presented in 1% agarose gel using the nd1 gene primers and the total DNA from the nine collected plankton samples. |
To evaluate the population of sea potato, we developed a relative quantitative analysis method based on the species-specific nd1 marker. The quality of DNA from each sample was verified by 18S rDNA with universal primers. The optical density ratio of nd1 and 18S rDNA PCR bands were used to represent the relative quantity of sea potato larvae, and compare the population variation among different samples. The 116 samples were collected from 13 stations distributed along the coast region of Fujian province (Fig. 3A). We divided all the sampling stations into four groups by the distance to the S01 station: group Ⅰ included S01, S02, and S03, which were off S01 station less than 1 km; group Ⅱ included S04, S05, S06 and S07, off S01 station 1 to 2 km; Group Ⅲ included S08, S09, and S10, off S01 station 4 to 6 km; group Ⅳ included S11, S12, and S13, off S01 station more than 9 km. The total DNA was extracted from the samples, respectively. All DNA samples were diluted to 20 ng μL-1, and 10 μL was utilized for PCR. The DNA quality of most samples (107/ 116) were sufficiently good for the amplification of the fragments of nd1 and 18S rDNA. The results showed obvious differences in the relative larval density among the samples and the groups (Fig. 3B). It clearly showed that the high-density sea potato was presented in group Ⅰ station in spring and summer near the land, suggesting that distribution of sea potato is associated to the marine sediment environment.
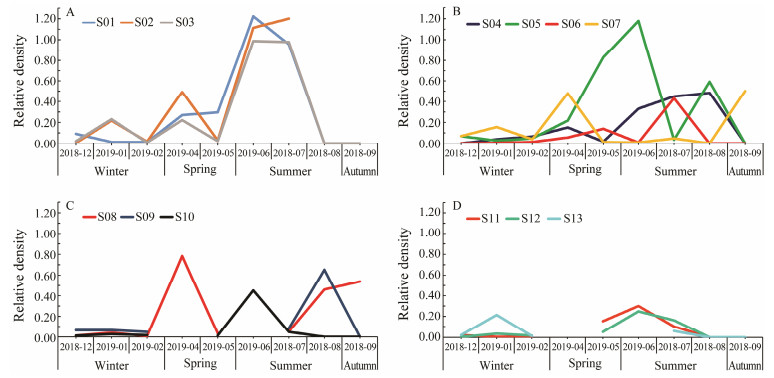
3.4 Season Variation of Acaudina spp. LarvaeThe samples were collected in four seasons. The data showed that the highest larval density appeared in June and July. For stations in group Ⅰ, there was a significant increase from May to July, and then a rapid decline after September (Fig. 4A). For stations in groups Ⅱ and Ⅲ, they showed similar patterns: the lowest density in Winter, and other seasons showed the moderate density (Figs. 4B and 4C). For group Ⅳ, the relative density was globally low and two small peaks were observed in January and June (Fig. 4D).
|
Fig. 4 Relative density of Acaudina spp. larvae at different stations. Variation trend of relative density plot in stations of group Ⅰ (A), group Ⅱ (B), group Ⅲ (C), and group Ⅳ (D), respectively. |
In this study, we successfully identified a 229 bp fragment of gene nd1 as Acaudina spp. specific molecular marker, which provides an effective molecular tool for species identification and population dynamic evaluation. By analyzing the relative quantity of this fragment with PCR, we quantitatively detected the larval density of Acaudina spp. in a marine coast, and found that their density decreased gradually from offshore to deeper waters. In the samples that were collected from about 20 m deep of the seawaters, the larval density is quite low. In addition, the higher density of larvae was presented in the samples collected in May, June, and July, suggesting that the larvae of Acaudina spp. might propagate from April to July.
It is worthy to note that the method that we developed in this study cannot distinguish the species within one genus. CRISPR-based specific high sensitivity enzymatic reporter unlocking technology (Gootenberg et al., 2017; Gootenberg et al., 2018) and DNA endonuclease targeted CRISPR trans reporter technology (Chen et al., 2018) can be the alternative ways to accurately quantify target molecules. Loop-mediated isothermal amplification is another potentially useful technology with high specificity and efficiency under isothermal conditions (Notomi et al., 2015).
Our results showed that the larval density of Acaudina spp. is dynamic and tightly correlates with the temperature and the sediment in the offshore seawaters. These results therefore provide useful information to understand the seasonal and regional distribution of Acaudina spp. larvae. Based on this information, population dynamics can be effectively evaluated, early warning mechanisms can be established, and the population density of Acaudina spp. can be regulated in the specific sea region.
AcknowledgementsThis work was supported by the National Key Research and Development Program of China (No. 2018YFD090 0705), the Key Laboratory of Integrated Marine Monitoring and Applied Technologies for Harmful Algal Blooms Funds (No. MATHAB201706), and the Fundamental Research Funds for the Central Universities (No. 201822016).
Behera, B. K., Baisvar, V. S., Rout, A. K., Pakrashi, S., Kumari, K., Panda, D., Das, P., Parida, P. K., Meena, D. K., Bhakta, D., Das, B. K. and Jena, J., 2018. The population structure and genetic divergence of Labeo gonius (Hamilton, 1822) analyzed through mitochondrial DNA cytochrome b gene for conservation in Indian waters. Mitochondrial DNA. Part A, DNA Mapping, Sequencing, and Analysis, 29: 543-551. (  0) 0) |
Behera, B. K., Kunal, S. P., Paria, P., Das, P., Meena, D. K., Pakrashi, S., Sahoo, A. K., Panda, D., Jena, J. and Sharma, A. P., 2015. Genetic differentiation in Indian Major Carp, Cirrhinus mrigala (Hamilton, 1822) from Indian Rivers, as revealed by direct sequencing analysis of mitochondrial Cytochrome b re-gion. Mitochondrial DNA Part A, DNA Mapping, Sequencing, and Analysis, 26: 1-3. (  0) 0) |
Boore, J. L., 1999. Animal mitochondrial genomes. Nucleic Acids Research, 27: 1767-1780. DOI:10.1093/nar/27.8.1767 (  0) 0) |
Boore, J. L. and Brown, W. M., 1994. Mitochondrial genomes and the phylogeny of molluscs. The Nautilus, 108(supp. 2): 61-78. (  0) 0) |
Boore, J. L., Macey, J. R. and Medina, M., 2005. Sequencing and comparing whole mitochondrial genomes of animals. Methods in Enzymology, 395: 311. (  0) 0) |
Brand, M. D., 1997. Regulation analysis of energy metabolism. Journal of Experimental Biology, 200: 193-202. DOI:10.1242/jeb.200.2.193 (  0) 0) |
Chen, J. S., Ma, E., Harrington, L. B., Da, Costa M., Tian, X., Palefsky, J. M. and Doudna, J. A., 2018. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science, 360: 436. DOI:10.1126/science.aar6245 (  0) 0) |
Curole, J. P. and Kocher, T. D., 1999. Mitogenomics: Digging deeper with complete mitochondrial genomes. Trends in Ecology & Evolution, 14: 394-398. (  0) 0) |
Das, S. P., Swain, S., Jena, J. and Das, P., 2018. Genetic diversity and population structure of Cirrhinus mrigala revealed by mitochondrial ATPase 6 gene. Mitochondrial DNA. Part A, DNA Mapping, Sequencing, and Analysis, 29: 495-500. (  0) 0) |
Fan, S., Hu, C., Wen, J. and Zhang, L., 2011. Characterization of mitochondrial genome of sea cucumber Stichopus horrens: A novel gene arrangement in Holothuroidea. Science China. Life Sciences, 54: 434-441. DOI:10.1007/s11427-011-4168-8 (  0) 0) |
Gootenberg, J. S., Abudayyeh, O. O., Kellner, M. J., Joung, J., Collins, J. J. and Zhang, F., 2018. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science, 360: 439. DOI:10.1126/science.aaq0179 (  0) 0) |
Gootenberg, J. S., Abudayyeh, O. O., Lee, J. W., Essletzbichler, P., Dy, A. J., Joung, J., Verdine, V., Donghia, N., Daringer, N. M., Freije, C. A., Myhrvold, C., Bhattacharyya, R. P., Livny, J., Regev, A., Koonin, E. V., Hung, D. T., Sabeti, P. C., Collins, J. J. and Zhang, F., 2017. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science, 356: 438. DOI:10.1126/science.aam9321 (  0) 0) |
Ingman, M., Kaessmann, H., Pääbo, S. and Gyllensten, U., 2000. Mitochondrial genome variation and the origin of modern humans. Nature, 408: 708-713. DOI:10.1038/35047064 (  0) 0) |
Morenosánchez,, R, ., Rodríguezenríquez,, S, ., Marínhernández,, A, . and Saavedra, E., 2010. Energy metabolism in tumor cells. FEBS Journal, 274: 1393-1418. (  0) 0) |
Notomi, T., Mori, Y., Tomita, N. and Kanda, H., 2015. Loopmediated isothermal amplification (LAMP): Principle, features, and future prospects. Journal of Microbiology, 53: 1-5. DOI:10.1007/s12275-015-4656-9 (  0) 0) |
Peregrino-Uriarte, A. B., Varela-Romero, A., Muhlia-Almazan, A., Anduro-Corona, I., Vega-Heredia, S., Gutierrez-Millan, L. E., De la Rosa-Velez, J. and Yepiz-Plascencia, G., 2009. The complete mitochondrial genomes of the yellowleg shrimp Far-fantepenaeus californiensis and the blue shrimp Litopenaeus stylirostris (Crustacea: Decapoda). Comparative Biochemistry and Physiology. Part D, Genomics & proteomics, 4: 45-53. (  0) 0) |
Perseke, M., Bernhard, D., Fritzsch, G., Brümmer,, F, ., Stadler, P. F. and Schlegel, M., 2010. Mitochondrial genome evolution in Ophiuroidea, Echinoidea, and Holothuroidea: Insights in phylogenetic relationships of Echinodermata. Molecular Phylog enetics & Evolution, 56: 201. (  0) 0) |
Scribner, K. T., Talbot, S. L., Pearce, J. M., Pierson, B. J., Bollinger, K. S. and Derksen, D. V., 2003. Phylogeography of Canada Geese (Branta canadensis) in western North America. The Auk, 120: 889-907. DOI:10.1642/0004-8038(2003)120[0889:POCGBC]2.0.CO;2 (  0) 0) |
Shen, X., Tian, M., Liu, Z., Cheng, H., Tan, J., Meng, X. and Ren, J., 2009. Complete mitochondrial genome of the sea cucumber Apostichopus japonicus (Echinodermata: Holothuroidea): The first representative from the subclass Aspidochirotacea with the echinoderm ground pattern. Gene, 439: 79-86. DOI:10.1016/j.gene.2009.03.008 (  0) 0) |
Takata, K., Yoshida, H., Hirose, F., Yamaguchi, M., Kai, M., Oshige, M., Sakimoto, I., Koiwai, O. and Sakaguchi, K., 2001. Drosophila mitochondrial transcription factor A: Characterization of its cDNA and expression pattern during development. Biochemical and Biophysical Rresearch Communications, 287: 474-483. DOI:10.1006/bbrc.2001.5528 (  0) 0) |
Wallace, D. C., Shoffner, J. M., Trounce, I., Brown, M. D., Ballinger, S. W., Corraldebrinski, M., Horton, T., Jun, A. S. and Lott, M. T., 1995. Mitochondrial DNA mutations in human degenerative diseases and aging. Biochimica et Biophysica Acta Molecular Basis of Disease, 1271: 141-151. DOI:10.1016/0925-4439(95)00021-U (  0) 0) |
Wang, G., Li, X., Wang, J., Zhang, J., Liu, W., Lu, C., Guo, Y. and Dong, B., 2019. The complete mitochondrial genome and phylogenetic analysis of Acaudina molpdioides. Mitochondrial DNA Part B-Resources, 4: 668-669. DOI:10.1080/23802359.2019.1572476 (  0) 0) |
Wang, X., Han, X., Zhang, Y., Liu, S. and Lin, Q., 2018. Phylogenetic analysis and genetic structure of the seahorse, Hippocampus fuscus from the Arabian and Red Sea based on mitochondrial DNA sequences. Mitochondrial DNA. Part A, DNA Mapping, Sequencing, and Analysis, 39: 165-171. (  0) 0) |
Wei, J., Zhang, J., Lu, Q., Ren, P., Guo, X., Wang, J., Li, X., Chang, Y., Duan, S., Wang, S., Yu, H., Zhang, X., Yang, X., Gao, H. and Dong, B., 2020. Genomic basis of environmental adaptation in the leathery sea squirt (Styela clava). Molecular Ecology Resources, 20: 1414-1431. DOI:10.1111/1755-0998.13209 (  0) 0) |
Wen, J., Hu, C., Zhang, L. and Fan, S., 2011. Genetic identification of global commercial sea cucumber species on the basis of mitochondrial DNA sequences. Food Control, 22: 72-77. DOI:10.1016/j.foodcont.2010.06.010 (  0) 0) |
Xiao, N., 2015. Echinoderm in the Yellow Sea and the Bohai Sea. Science Press, Beijing, 100pp (in Chinese).
(  0) 0) |
 2021, Vol. 20
2021, Vol. 20


