Revolutionizing microbial treasure troves: innovative strategies for natural products discovery
Abstract
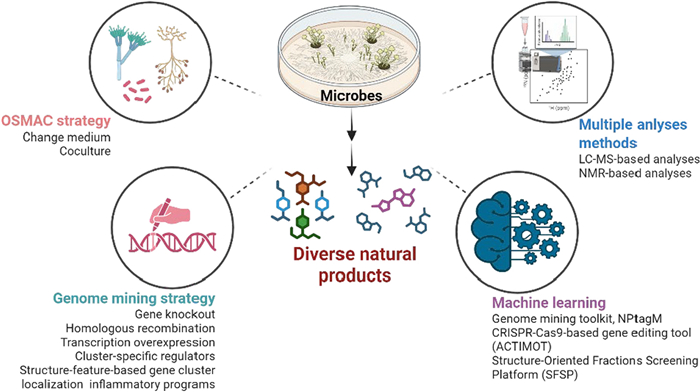
Microorganisms represent Earth's most abundant biological resource, producing metabolites of immense value across medicine, agriculture, and industry. Conventional cultivation and screening techniques, however, suffer from inefficiency and fail to meet contemporary demands. Providing a comprehensive overview, this review details how the One Strain Many Compounds (OSMAC) strategy—addressing cultivation bottlenecks—and genomics-driven mining approaches are revolutionizing the discovery of novel microbial metabolites. Crucially, it underscores the broad adoption of innovative technologies like machine learning to enable faster, more effective gene and structure targeting. Synthesizing case studies from 2019 to 2025, the review catalogs newly identified compounds and their bioactivities, while outlining future research directions to establish a theoretical framework for efficient microbial natural product exploration. These advanced discovery strategies are significantly accelerating the identification of structurally diverse lead compounds with novel mechanisms of action, thereby revitalizing pipelines for new antibiotic, anticancer, and therapeutic drug development.Graphical Abstract
Keywords
Microbial natural products OSMAC strategy Genome mining Machine learning1 Introduction
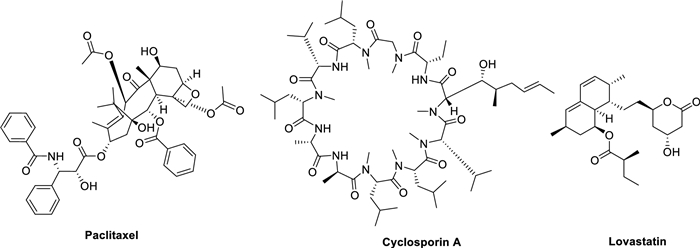
Microbial natural products, produced by diverse microorganisms including bacteria, fungi and actinomycetes, are renowned for their unique structural characteristics and broad spectrum of biological activities. These compounds have long been recognized as a crucial source of lead compounds in pharmaceutical development [1]. Statistical data reveal that approximately 70% of clinically used antibiotics originate from microbial sources [2, 3]. Beyond their antibiotic applications, microbial metabolites have also contributed significantly to the development of various therapeutic agents, including anticancer drugs. A notable example is paclitaxel, which was initially isolated from plants but is now commercially produced through microbial fermentation [4, 5]. Furthermore, microbial-derived compounds such as the immunosuppressant cyclosporin A (essential in organ transplantation) and the cholesterol-lowering agent lovastatin [6, 7] have become indispensable in modern medicine (Fig. 1).
Chemical structures of significant microbial natural drugs (paclitaxel, cyclosporin A, and lovastatin)
Despite the remarkable potential of microbial metabolites, researchers face several persistent challenges: (1) Cultivation limitations: More than 90% of environmental microorganisms resist cultivation under standard laboratory conditions, leaving an enormous reservoir of potentially novel compounds unexplored—particularly from unculturable microbes inhabiting extreme environments such as deep-sea ecosystems. (2) High rediscovery rate: Frequently studied microbial strains have been extensively characterized, resulting in inefficient screening processes that predominantly yield known compounds. (3) Silent biosynthetic gene clusters (BGCs): While microbial genomes typically harbor numerous BGCs, the majority remain transcriptionally inactive under conventional laboratory conditions, rendering their associated metabolic products undetectable. These challenges underscore the urgent need for paradigm-shifting innovations that transcend traditional research approaches, demanding the development of novel methodologies to unlock the full potential of microbial resources.
Unlike previous reviews limited to singular methods (e.g., synthetic biology or metabolomics) [8–11] or compound classes (e.g., peptides/polyketides) [12], this work provides a comprehensive analysis of challenges in discovering structurally diverse, bioactive microbial natural products and the innovative strategies addressing them. It systematically examines three transformative approaches: "One Strain Many Compounds" (OSMAC) cultivation, genome mining (CRISPR-enhanced), and machine learning-driven platforms (NPtagM/SFSP), curating advances (2019–2025) yielding over 300 novel bioactive compounds. The review concludes by proposing a roadmap integrating single-cell analysis, multi-omics, and synthetic biology to unlock microbial "dark matter" for next-generation drug/agrochemical discovery.
2 Innovative strategies
2.1 OSMAC strategy
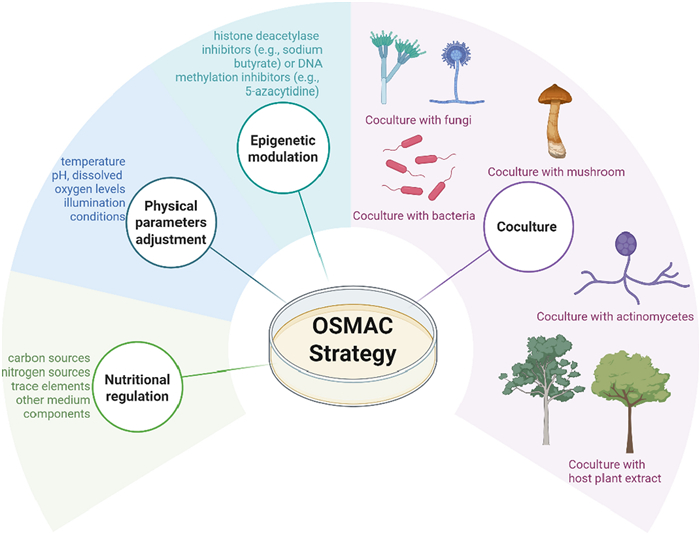
The "One Strain Many Compounds" (OSMAC) strategy represents an innovative approach in microbial cultivation optimization, which systematically modulates culture conditions to activate silent BGCs in microorganisms, thereby significantly expanding the diversity of their secondary metabolites [13]. In conventional microbial natural product research, a single strain typically yields only a limited array of known compounds under standard laboratory conditions. In contrast, the OSMAC approach substantially enhances chemical diversity through strategic manipulation of cultivation parameters. The core principles of OSMAC involve: (1) Nutritional regulation: Modifying carbon sources, nitrogen sources, trace elements, and other medium components to redirect metabolic flux; (2) Physical parameter adjustment: Optimizing temperature, pH, dissolved oxygen levels, and illumination conditions; (3) Biological interactions: Employing co-culture systems to simulate natural ecological interactions, thereby inducing the production of defensive metabolites through competitive or symbiotic relationships; (4) Epigenetic modulation: Utilizing histone deacetylase inhibitors (e.g., sodium butyrate) or DNA methylation inhibitors (e.g., 5-azacytidine) to activate dormant gene clusters. This multi-parameter synergistic regulation strategy establishes a novel paradigm for the exploration of microbial natural products (Fig. 2).
The description of OSMAC strategy
2.1.1 Modification of cultivation methods
2.1.1.1 Variations in culture medium
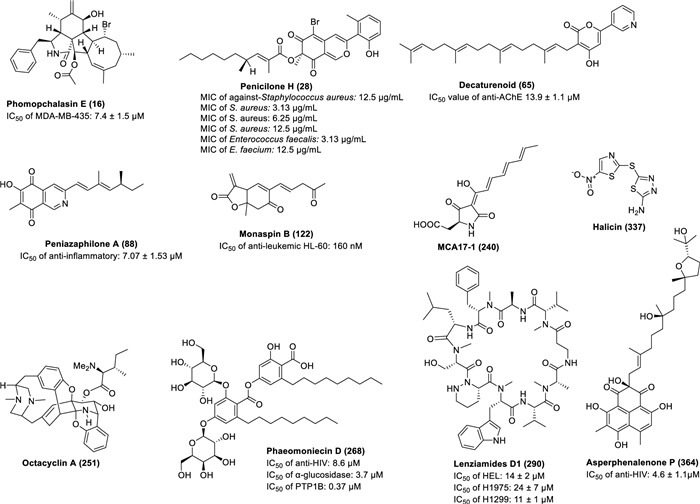
Targeted modification of culture medium components can specifically activate fungal secondary metabolic pathways. Research demonstrates that: the marine fungus Pseudallescheria boydii F44-1 produces novel spirocyclic bisindole alkaloids pseudboindoles A-B (1‒2) in amino acid-supplemented GPY medium [14], while Trichocladium sp. yields the first-reported bismacrolactone 13-N-(2-carboxyphenyl) colletoketol (3) in rice medium containing 2% tryptophan [15]. In the cultivation of the endophytic fungus Aplosporella javeedii, the addition of 3.5% NaNO3 induced the production of seven new compounds, aplosporellins A-C (4‒6), E–G (7‒9) and J (10), while 3.5% monosodium glutamate yielded four structurally distinct compounds, aplosporellins D (11) /H (12) /I (13) /K (14), highlighting the specific regulatory effects of nitrogen sources on metabolic pathways [16]. The mangrove fungus Phomopsis sp. QYM-13 produced brominated cytochalasins, phomopchalasins D (15), E (16), I (17), K (18), and N (19) in medium containing 3% NaBr, while iodinated derivatives, phomopchalasins F (20), G (21), H (22), J (23), L (24), M (25), O (26) were generated in 3% KI medium. Notably, compounds 16, 20, and 22 represented the first brominated and iodinated cytochalasins, exhibiting significant cytotoxicity against MDA-MB-435 cancer cells [17]. Similarly, Penicillium janthinellum HK1-6 produced two novel brominated azaanthraquinones penicilones G-H (27‒28) (Fig. 3 and Additional file 1: Fig. S1) and two new tricyclic polyketides penijanthinones A-B (29‒30) under NaBr induction, with penicilone H (28) showing antimicrobial activity against various pathogens (MIC: 3.13–12.5 μg/mL) [18]. These findings systematically reveal how medium components (amino acids, nitrogen sources, halide salts) precisely regulate the structural diversity and bioactivity of microbial metabolites, providing crucial strategies for targeted discovery of novel bioactive lead compounds.
Structures of representative bioactive compounds (other compounds are shown in supplementary files) obtained from the coculture
The coral-derived fungus Stachybotrys chartarum produced six new phenyldrimanes, distachydrimanes A-F (31‒36), with unprecedented C-18-C-23 connectivity in medium containing 0.1% Fe2(SO4)3 and 3% sea salt. The compounds 31, 35, and 36 inhibited L1210 cells through multiple mechanisms: > 70% inhibition rate, G0/G1 phase arrest, senescence promotion, and mitochondrial apoptosis pathway activation [19]. Meanwhile, Preussia isomera yielded the rare tetrahydro-2H-1,2-oxazine alkaloid (±)-preisomide (37) when cultured in rice medium supplemented with wheat bran liquid medium. In MEB-containing rice medium, it produced the sesquiterpenoid enantiomers (±)-preuisolactone A (38), featuring a novel tricyclo [4.4.01,6.02,8] decane skeleton, with a MIC of 10.2 μM against Micrococcus luteus [20, 21]. These findings systematically demonstrate how the mixed composition of culture medium precisely modulates both the structural diversity and bioactivities of microbial metabolites, thereby establishing an innovative research paradigm for targeted discovery of bioactive molecules through optimized culture conditions.
Research on the medium type-dependent activation of fungal secondary metabolites demonstrates that: the corn-derived Fusarium sp. YMX-C33 produced six novel sesquiterpenoids, fusarchlamols A-F (39‒44), in PDB medium, while coffee medium specifically produced methyltricinonoate (45), a structurally novel natural sesquiterpenoid, along with compounds 41 and 42. Bioactivity assessment demonstrated that compounds 39, 40, and 43/44 exhibited significant inhibitory activity against the coffee pathogenic fungus Alternaria alternata (MIC: 1 μg/mL) [22]. Research on Penicillium canescens found that solid rice medium produced penicanesins A-B (46‒47), and E–G (48‒50), while liquid medium specifically yielded penicanesins C-D (51–52). Remarkably, (±)-penicanesins A (46) and B (47) featured a rare 6/6/6/6 heterotetracyclic skeleton, with (+)-penicanesin A (46) showing IC50 values of 8.2 and 9.8 μM against HL-60 and SW480 cancer cells, respectively [23]. Similarly, Pleotrichocladium opacum produced compounds 53–54 in JSA liquid medium, compound 55 in rice medium, and compound 56 in wheat medium [24]. These findings systematically demonstrate how medium type and physical state (solid/liquid) precisely regulate the structural diversity and bioactivity of microbial metabolites. This approach provides a strategic framework for the targeted discovery of novel bioactive lead compounds.
2.1.1.2 Chemical epigenetic modifications
Chemical epigenetics refers to the exogenous addition of epigenetic modifiers, including histone deacetylase (HDAC) inhibitors and DNA methyltransferase (DNMT) inhibitors, to specifically alter the epigenetic modification state of microbial chromatin [25]. This approach alleviates gene silencing effects, activates dormant BGCs, and promotes the biosynthesis of structurally novel bioactive compounds [26, 27]. Within the OSMAC strategy, chemical epigenetic regulation primarily functions through the following molecular mechanisms: HDAC inhibitors (such as sodium butyrate, suberoylanilide hydroxamic acid, SAHA and suberic bis-hydroxamic acid, SBHA) increase histone acetylation levels, loosening condensed chromatin structures and significantly enhancing the transcriptional activity of secondary metabolite genes [28, 29]; DNMT inhibitors (such as 5-azacytidine, 5-Aza) reduce DNA methylation levels, effectively reversing gene silencing and activating cryptic metabolic pathways [30, 31]; histone methylation modulators finely regulate gene expression by dynamically adjusting histone methylation marks (e.g., activating H3K4me3 or repressive H3K27me3 modifications) [32].
The research team successfully activated silent metabolic pathways in the endophytic fungus Chaetomium globosporum isolated from Euphorbia humifusa using 5-Aza, leading to the isolation of two novel cyclopentenones globosporins A-B (57‒58) and two monoterpene indole alkaloids globosporines C-D (59‒60). Notably, compound 60 exhibited significant inhibitory activity against Xanthomonas oryzae (MIC: 14‒72 μg/mL), demonstrating excellent potential for development as a biopesticide in both in vitro and in vivo tests [33].
Treatment of the endophytic fungus Penicillium sp. KMU18029 with SAHA yielded two pairs of diterpene hybrids, pyrandecarurins A-B (61‒62) and pileotins A-B (63–64), which feature unique pyridopyrrolidone-decaturene/oxalic acid hybrid scaffolds. Of particular interest, the potential precursor decaturenoid (65) showed moderate activity against AChE with an IC50 value of 13.9 ± 1.1 μM [34].
The combined application of SBHA and 5-Aza on the deep-sea-derived fungus Eutypella sp. MCCC 3A00281 resulted in the production of 17 novel sesquiterpenoids eutypeterpenes A-Q (66–82). Among these, compound 66 represents the first bergamotane-type sesquiterpene containing a dioxolane unit, while compounds 80–82 constitute an entirely new subclass of sesquiterpenes with cyclopentane structures. All compounds showed weak cytotoxic activity with IC50 > 100 μM on RAW 264.7 cells, indicating safety to macrophages [35]. Similarly, the combined treatment of marine-derived fungus Aspergillus versicolor XS-20090066 with SAHA and 5-Aza, leading to enhanced production of diverse secondary metabolites. Through an integrated approach combining GNPS with bioactivity-guided fractionation, three previously undescribed compounds were characterized: the nucleoside derivatives kipukasins K (83) and L (84), along with the sesquiterpenoid aspergillusene E (85). Antimicrobial evaluation revealed remarkable potency, with compounds 83 and 85 demonstrated significant antibacterial activity against Staphylococcus aureus and other pathogens (MIC: 8‒16 μg/mL), providing promising candidates for anti-infective drug development [36].
The collective findings demonstrate that precisely designed epigenetic strategies can effectively activate silent biosynthetic pathways in microorganisms across diverse ecological niches, yielding structurally novel secondary metabolites with significant biological relevance.
2.1.2 Coculture
2.1.2.1 Fungal-fungal co-culture
Recent advances in fungal co-culture technology have revolutionized natural product discovery by effectively mimicking microbial ecological interactions [37–39]. This approach activates silent BGCs in fungi, enabling the production of structurally novel secondary metabolites that are typically undetectable in monoculture systems. Notably, the method not only expands chemical diversity but also enhances yields of target compounds, thereby generating valuable resources for pharmaceutical lead development. As a cutting-edge strategy in microbial natural product research, this technology demonstrates considerable potential for drug discovery applications.
(1) Co-culture with Penicillium fungi
Penicillium species have been extensively studied as prolific producers of structurally diverse and biologically active natural products [40, 41]. Recent genomic analyses reveal these filamentous fungi possess numerous silent biosynthetic gene clusters encoding potential secondary metabolites [42]. Through culture optimization and epigenetic modulation strategies, researchers have successfully activated these cryptic pathways to discover novel antibiotics, antitumor agents, and other pharmacologically relevant compounds from Penicillium strains [43].
Co-culture of the marine-derived P. bilaiae MA-267 and P. chermesinum EN-480 produced two new terpenoid derivatives, chermebilaenes A-B (86–87). Compound 86, as the first hybrid of acorane-type sesquiterpene and octadecadienoic acid, exhibited strong inhibitory activity against Ceratobasidium cornigerum and Edwardsiella tarda[44]. Meanwhile, the co-culture system of mangrove endophytic fungi P. sclerotiorum THSH-4 and ZJHJJ-18 produced nine new azaphilone derivatives, peniazaphilones A-I (88–97). Compound 88, the first naturally occurring isoquinolinequinone-structured azaphilone, demonstrated superior anti-inflammatory activity compared to indomethacin and showed significant cytotoxicity against A549 and MDA-MB-435 tumor cells [45] (Fig. 3 and Additional file 1: Fig. S1). Additionally, co-culture of the extremophilic P. fuscum with P. camembertii/clavigerum activated the production of berkeleypenostatins A-G (98–103), among which compound 101 exhibited broad-spectrum antitumor activity (TGI 1–10 μM) and the ability to inhibit the migration of human pancreatic cancer cells [46]. Another study involving P. bialowiezense and Pestalotiopsis sp. co-culture yielded six new prenylated chromane derivatives, including two pairs of enantiomeric ones (104a/104b-105a/105b) and two optical pure ones (106–107), as well as two new isoprenylated phenol glucoside derivatives (108–109), along with previously reported enantiomers 104a/104b and 105a/105b, with 104a/104b showing significant GUS inhibitory activity [47]. Furthermore, co-culture of P. brasilianum MST-FP1927 and Aspergillus nomius MST-FP2004 led to the discovery of miktospiromide A (110) and kitrinomycin A (111), the latter of which exhibited notable inhibitory effects against mouse melanoma cells and bovine parasites [48].
(2) Co-culture with Aspergillus fungi
The Aspergillus genus encompasses a diverse and ubiquitous group of fungi renowned as prolific producers of secondary metabolites [42]. They are particularly adept at generating structurally diverse natural products with significant bioactivity [49], including renowned antibiotics, cholesterol-lowering drugs, mycotoxins, and various industrial enzymes [50–52]. Their rich repertoire of BGCs renders them a valuable treasure trove for drug discovery and biotechnological applications.
The co-culture of A. fischeri and Xylaria flabelliformis produced wheldone (112), which exhibited notable cytotoxicity against breast, ovarian, and melanoma cancer cells [53]. The interaction between A. oryzae and Epicoccum dendrobii not only increased kojic acid production to 1.10 g/L, but also activated the production of four novel metabolites, epiclactones A-B (113–114), epioxochromane (115) and aoergostane (116), with multi-omics analysis revealing that this process is closely linked to fungal defense mechanisms [54]. In the co-culture system of marine-derived A. insulicola IMB18-072 and Alternaria angustiovoidea IMB20-805, the cyclic tetrapeptides violaceotides B-E (117–120) were isolated and identified. Among these, compounds 118–119 demonstrated selective inhibitory activity against aquatic pathogens (Edwardsiella tarda and E. ictaluri), while 117–120 significantly suppressed the expression of the inflammatory factor IL-6 [55]. The co-culture system of A. oryzae and Monascus purpureus successfully yielded two novel cyclohexyl-furan compounds, monaspins A-B (121–122). Compound 122 demonstrated remarkable anti-leukemic HL-60 cell activity (IC50: 160 nM) and exhibited significant in vivo anti-leukemic effects in a mouse leukemia model by reducing white blood cell, lymphocyte, and neutrophil counts [56] (Fig. 3 and Additional file 1: Fig. S1). Additionally, the co-culture study of Amphichorda sp. KMM 4639 and A. carneus KMM 4638 yielded important results, leading to the discovery of five new quinazolinone alkaloids, felicarnezolines A-E (123–127), and a novel highly oxygenated chromene derivative, oxirapentyn M (128). Notably, compound 124 exhibited potent cardioprotective and neuroprotective effects, effectively mitigating CoCl2-induced damage in H9c2 cardiomyocytes and SH-SY5Y neuroblastoma cells [57]. On the other hand, the marine-derived co-culture system of A. aculeatinus WHUF0198 and Penicillium sp. DM27 achieved a major breakthrough: not only were aculeaquamides B-C (129–130) obtained, featuring a rare 7/6/5/5/6/5 hexacyclic scaffold, but 17 structurally novel pyranopyridone alkaloids, aculeapyridones A-Q (131–147), were also isolated. Among these, compounds 142–145 were particularly distinctive due to their unique N-methoxy group, while compounds 131–147, 139–140, and 142–145 demonstrated significant protective effects against acetaminophen-induced liver injury [58, 59].
(3) Co-culture of macrofungal
Macrofungi hold significant value in the fields of food, medicine, and biotechnology. In terms of food applications, they are rich in high-quality protein, dietary fiber, and various bioactive compounds (e.g., polysaccharides and triterpenoids), making them nutritious functional food ingredients [60, 61]. In pharmaceutical applications, the secondary metabolites of macrofungi exhibit remarkable pharmacological activities. For example, Ganoderma polysaccharides have immunomodulatory effects [62, 63], and cordycepin shows antitumor activity [64]. These properties have led to their widespread use in both traditional medicine and modern drug development. Moreover, biotechnological approaches, such as co-culture techniques, can significantly enhance the production of bioactive compounds, further expanding their application potential [65].
Co-cultivation of Pleurotus ostreatus SY10 and P. eryngii SY302 yielded the monoterpene pleurotusin A (148) and cyclopentenone pleurotusin B (149), with 148 exhibiting antibacterial activity against Staphylococcus aureus [66]. In another study, the co-culture of Pleurotus ostreatus SY10 and Trametes robiniophila SY636 yielded new terpenoids postredienes D-H (150–154), among which compound 150 exhibited inhibitory activity against Candida albicans and Cryptococcus neoformans comparable to amphotericin B [67]. When Trametes versicolor SY630 was co-cultured with Vanderbylia robiniophila SY341 or Ganoderma gibbosum SY1001, antimicrobial compounds tramevandins A-C (155‒157) and sphingolipids 17-ene-1-deoxyPS (158) and 1-deoxyPS (159), which downregulate ergosterol synthesis in C. albicans, were discovered [68].
(4) Co-culture of other fungal species
In the co-culture system of Clonostachys rosea B5-2 and Nectria pseudotrichia B69-1, three terpenoid derivatives—furanocochlioquinol (160), furanocochlioquinone (161), and nectrianolin D (162)—were isolated and characterized. In vitro antitumor activity evaluation demonstrated their significant cytotoxic effects against human leukemia HL60 cells, with IC50 values ranging from 0.47 to 10.16 μM [69].
The investigation of immunomodulatory compounds showed that the co-culture of Phellinus orientoasiaticus and Xylodon flaviporus yielded three structurally novel sesquiterpenoids: 6-nor-4-hydroxy-1,3(4)-distepuren-14-oic acid (163), 6-hydroxysterpuric acid (164), and 13-hydroxy-7-deoxypaneolilludinic acid (165). Notably, compound 163 exhibited significant enhancement of NO production in LPS-induced RAW264.7 macrophage models, suggesting its potential immunomodulatory activity [70].
In the study of antifungal activity against plant pathogens, the co-culture system of Nigrospora oryzae and Irpex lacteus produced a variety of bioactive metabolites, including two squalene-type compounds, irpenigirins A-B (166–167); a novel azaphilone derivative, isonigirpexin C (168); and two sesquiterpenoids, 5-demethyl conocenol C (169) and nigrosirpexin A (170). All of these compounds displayed potent antifungal activity against phytopathogens [71]. In addition, a structurally unique isoquinoline alkaloid, irpexine (171), was also isolated from the co-culture of Irpex lacteus and Phaeosphaeria oryzae [72].
The co-culture of Colletotrichum pseudomajus and Daldinia eschscholtzii yielded 16 structurally new polyketide metabolites, assigning as coldaldols A-C (172–174), collediol (175), and daldinrins A-L (176–187). Among these, compounds 177 and 178 featured a rare benzopyran-C7 polyketide hybrid skeleton, while compounds 179–182 exhibited novel benzopyran dimer structures. Bioactivity assessment revealed that compound 174 had significantly stronger antifungal activity than the positive control drug nystatin [73].
Furthermore, phenolic compounds phexandiols A-B (188–189) and ester derivatives phomesters A-C (190–192) were produced by the co-culture of Phoma sp. YUD17001 and Armillaria sp. [74]. Additionally, the marine-derived fungus Cosmospora sp. and plant pathogen Magnaporthe oryzae co-culture successfully induced two new derivatives, soudanones H-I (193–194) [75].
The co-culture of Herpotrichia sp. SF09 and Trametes versicolor SF09A generated 12 novel metabolites, hertramycines A-L (195–206). Compounds 195 and 196 were sesquiterpene-saccharide conjugates containing a xylopyranosyl group, (±)-197 represented the first naturally occurring linear sesquiterpene racemate, while 201 was an α-pyrone derivative bearing a xylopyranosyl unit. In the MPP⁺-induced oxidative damage PC12 cells model, all compounds displayed dose-dependent neuroprotective effects, with compound 199 showing the most potent protection at 5 μM concentration. Mechanistic studies uncovered that compound 199 likely exereted its neuroprotective effects through modulation of the PI3K-Akt/MAPK signaling pathway, making it a potential candidate for developing novel Parkinson's disease therapeutics [76].
2.1.2.2 Fungal-bacterial co-culture
Currently, multiple studies have shown that the co-culture of fungi and bacteria can significantly induce the production of cryptic metabolites, which are often undetectable under monoculture conditions.
(1) Co-culture of fungi and Bacillus subtilis
Bacillus subtilis, a ubiquitous Gram-positive soil and rhizosphere bacterium, is a model organism prized for its prolific secondary metabolism. It efficiently produces diverse bioactive natural products, including key antibiotics (e.g., surfactin, iturin), antimicrobial peptides, antifungal polyketides, and industrial enzymes [77]. Its remarkably modular nonribosomal peptide synthetase (NRPS) and polyketide synthase (PKS) gene clusters make it an ideal platform for both natural product discovery and synthetic biology applications [78].
In 2023, researchers isolated a new terpenoid derivative, andrastin I, from the marine fungus Penicillium ochrochloron. A subsequent co-culture with B. subtilis led to the discovery of a new butyrolactone homologue, ochrochloronic acid (207) [79]. Similarly, when the sponge-associated fungus Aspergillus versicolor was co-cultured with B. subtilis, a new cyclic pentapeptide, cotteslosin C (208), an aflaquinolone-type compound, 22-epi-aflaquinolone B (209), and two new anthraquinones, isoversicolorin B (210) and 6,8-O-dimethylbipolarin (211) were produced [80]. Furthermore, co-culture of the endophytic fungus Trichocladium sp. from Houttuynia cordata roots with B. subtilis induced the production of a new compound, 5-epi-pestafolide A (212) [15].
(2) Co-culture of fungi with other Bacteria
Beyond B. subtilis, co-culturing fungi with other microorganisms can also activate novel metabolic pathways. For example, when the wheat root-associated bacterium Pantoea agglomerans was co-cultured with the date palm leaf-derived fungus Penicillium citrinum, it produced new pulicatin derivatives, pulicatins H-I (213‒214), which exhibited strong antifungal activity, explaining the observed fungal growth inhibition in co-culture [81].
Additionally, co-culture of the actinobacterium Streptomyces sp. 13F051 with the fungus Leohumicola minima 15S071 yielded a novel polyketide-peptide hybrid natural product, ulleungdolin (215). This compound features a complex structure with rare glycosylation modifications and showed significant anti-migratory effects against MDA-MB-231 breast cancer cells [82].
2.1.2.3 Bacterial-bacterial co-culture
When Streptomyces sp. GA02 isolated from mountain soil was cocultured with Pandoraea sp. GA02N, it could efficiently synthesize the aromatic metabolite, gwanakoside B (216) containing a 6-deoxy-α-L-talopyranose structure. Particularly noteworthy is that the yield of this compound under coculture conditions reached 100 times that of axenic culture. This remarkable enhancement highlights the advantages of microbial coculture in secondary metabolite production [83].
By co-culturing Streptomyces hygroscopicus HOK021 and Tsukamurella pulmonis TP-B0596, a structurally novel bifunctional antimicrobial conjugate, harundomycin A (217) were identified. This conjugate was formed through a thioester bond between the 2,4-dihydroxy-3-aminobenzoic acid moiety of platensimycin and N, N′-bis(2,3-dihydroxybenzoyl)-O-seryl-cysteine (bisDHBA-Ser-Cys). Genomic analysis revealed that strain HOK021 possesses biosynthetic gene clusters for both platensimycin and enterobactin, while T. pulmonis TP-B0596 acts as an inducer to activate these metabolic pathways. Compound 217 not only retained platensimycin-like antibacterial activity against drug-resistant pathogens such as MRSA and VRE but also exhibited enterobactin-like iron-chelating capacity [84].
Two Red Sea-derived actinomycetes, Actinokineospora spheciospongiae EG49 and Rhodococcus sp. UR59 were cocultured, leading to the isolation of several angucycline-class compounds [actinosporins E (218), H (219), G (220), tetragulol (221)] and the anthraquinone compound capillasterquinone B (222), which were undetectable in monocultures. These compounds displayed high affinity for Plasmodium lysyl-tRNA synthetase (PfKRS1) and exhibited significant in vitro antimalarial activity [85].
In summary, co-cultivation strategy, by mimicking natural microbial interactions, has proven to be an effective approach for activating silent metabolic gene clusters and discovering novel natural products. Despite its remarkable success, the underlying molecular mechanisms remain poorly understood. Consequently, future research paradigms should shift toward integrating multi-omics and real-time analysis technologies to systematically elucidate interaction networks, ultimately enabling a paradigm shift from empirical methods to predictable and rational design.
2.1.3 Emerging cultivation technologies and tools
While the OSMAC strategy has successfully stimulated microbial metabolic potential by altering culture conditions, its limitation lies in its continued reliance on laboratory pure culture techniques, making it difficult to access the vast majority of unculturable microorganisms in nature. This constitutes a more fundamental challenge in the field of natural product discovery. To overcome this limitation, a series of novel cultivation methods and tools designed to simulate natural habitats have emerged, significantly expanding the diversity of cultivable microorganisms.
In situ cultivation is a technological strategy that overcomes the limitations of traditional laboratory pure culture. Its core principle involves placing microorganisms in their native natural environment for cultivation, thereby directly utilizing various ecological factors from the habitat to induce microbial growth [86]. The technique operates by using physical barriers such as semi-permeable membranes to allow free diffusion of essential chemical factors like nutrients and signaling molecules from the environment, while confining the target microorganisms within specific spaces, thus creating a controlled yet nearly natural growth microenvironment.
To achieve higher throughput and standardized cultivation, novel cultivation devices such as the iChip have been developed. The iChip is a high-throughput in situ cultivation device whose basic operational workflow involves: serially diluting environmental samples and mixing them with molten agarose, then injecting the mixture into a chip containing multiple microwells so that each microwell theoretically contains only a single microbial cell; subsequently sealing the chip with semi-permeable membranes and returning it to the source environment for cultivation [87]. The semi-permeable membranes allow free diffusion of growth factors and signaling molecules from the environment, providing the encapsulated microbial cells with a nearly natural chemical environment while physically isolating them within the microwells to form monoclonal colonies. Research has shown that the microbial isolation rate achieved using the iChip can be several times to an order of magnitude higher than traditional plate culture methods, thereby successfully unlocking a vast reservoir of previously "uncultured" microbial taxa and leading to the discovery of active molecules with novel structures, including antibiotics [87, 88].
Beyond the iChip, other microenvironment control technologies such as microfluidic cultivation chips [89], and microbial encapsulation [90] are rapidly developing. These emerging technologies, combined with OSMAC strategies and genome mining, together form a powerful toolkit that is systematically addressing the challenge of microbial uncultivability, paving a broad pathway for the discovery of novel natural products from microbial "dark matter."
2.2 Genome mining strategy
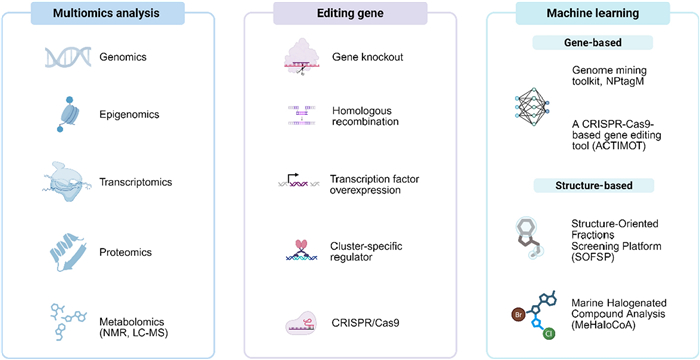
Genome mining is an integrated research strategy for natural product discovery based on microbial genomic data. The genome mining strategy is not a single research method, but rather a combinatorial research approach that continues to evolve. Depending on the specific circumstances of different research subjects, genome mining employs various methods. In 2009, Christophe Corre systematically elaborated on the genome mining strategy in a review published in Natural Product Reports [91]. It is notably that the rapid development of gene-editing technologies such as CRISPR [92] and Multiplex Base-Editing [93], combined with metabolic engineering modifications, has significantly enhanced both the discovery and production capacity of microbial secondary metabolites, providing robust technical support for natural product development (Fig. 4).
Fundamental methods of genome mining strategy
2.2.1 Gene knockout
Precise elimination of specific gene functions through frame-shift mutations or premature stop codons, serving as foundational tools for metabolic flux redirection. In the polar strain Eutytella sp. D-1, researchers employed gene knockout technology to delete the key gene responsible for triterpene cyclase biosynthesis, inducing metabolic flux reprogramming and activating dormant secondary metabolic gene clusters. In this study, 8 novel structural compounds, 223–230 were isolated and identified, among which compound 229 contained a unique 5/10 macrocyclic ether skeleton. In-depth functional studies demonstrated that the acorane-type sesquiterpenoid 230 exhibited significant anti-inflammatory activity in both cellular and transgenic zebrafish models by regulating the MAPK and NLRP3/caspase-1 signaling pathways [94].
Meanwhile, a novel antifungal compound alligamycin A (231) was obtained from the rapamycin-producing strain Streptomyces iranensis through genome mining. Targeted gene editing was used to knockout the aliA and aliH genes within the polyketide synthase (PKS) gene cluster, confirming the biosynthesis origin of the cluster. Compound 231 features a unique β-lactone-[6, 6]-spiroketal hybrid scaffold (with 13 chiral centers) and a novel polyketide extender unit (7-oxo-octylmalonyl-CoA). The gene knockout directly abolished the biosynthesis of the target compound. Subsequent structural derivatization (disruption of the β-lactone ring in alligamycin B (232)) demonstrated that this moiety is the essential pharmacophore for inhibiting drug-resistant fungi (including Aspergillus and Talaromyces species). Proteomics analysis revealed that compound 231 exerted its antifungal effects by compromising fungal cell wall integrity, providing a promising candidate to address the clinical limitations of current antifungal therapeutics [95].
2.2.2 Homologous recombination
Template-directed DNA repair mechanism achieving scarless gene replacement or insertion, critical for stable pathway refactoring. In the fungal strain Calcarisporium arbuscula NRRL 3705, researchers pioneered a novel approach by employing homologous recombination to generate the ΔaurA knockout mutant of the aurovertins biosynthetic gene cluster. Through high-throughput screening, two highly efficient promoters, aurBp and A07068p were identified. This integrated strategy accomplished dual objectives: it successfully activated the expression of endogenous polyketide synthase (PKS) gene clusters while simultaneously enabling high-efficiency heterologous expression of three exogenous fungal PKS gene clusters. The methodology substantially broadened the structural diversity of aurovertin-class polyketide 233, thereby establishing an innovative technological platform for natural product structural engineering [96].
2.2.3 Transcription factor overexpression
Amplification of rate-limiting transcriptional actuators to overcome metabolic bottlenecks, particularly effective for silent cluster awakening. By employing whole-genome scanning and bioinformatic prediction techniques, researchers identified a gene cluster encoding a distinctive tandem polyketide synthase in Ascomycete sp. F53. The isolated azaphilone compound, lijiquinone (234), featured a unique cyclohexenone-bicyclic hybrid scaffold and showed remarkable cytotoxicity along with broad-spectrum antifungal activity [97]. Notably, in Talaromyces sp., overexpression of the transcription factor LutB successfully activated a cryptic metabolic pathway, yielding five structurally novel sclerotiorin-type azaphilones (235–239). Subsequent gene knockout and isotope labeling experiments elucidated the pivotal role of the LutC-LutD oxidoreductase system in the biosynthetic pathway [98].
A genetic engineering strategy was used to activate silent biosynthetic gene clusters in filamentous fungi: Using an aurovertin-deficient mutant strain as the host, overexpression of a pathway-specific zinc-finger transcription factor successfully activated the silent gene cluster mca17 in Calcarisporium arbuscula, leading to the isolation of a novel tetramic acid-type MCA17-1 (240). Biosynthesis of this compound was catalyzed by a PKS-NRPS hybrid enzyme, and it exhibited significant pharmacological activity in a liver fibrosis model: In the TGF-β-induced LX-2 hepatic stellate cell activation assay, compound 240 exhibited significant inhibitory activity to the clinical positive control drug obeticholic acid (OCA) [99] (Fig. 3 and Additional file 1: Fig. S1).
2.2.4 Engineered activation of cluster-specific regulators
Artificial triggering of pathway-specific transcriptional switches via synthetic promoters or inducer molecules, exemplified by: through comprehensive genomic mining of Aspergillus terreus, researchers identified a cryptic PKS-NRPS hybrid gene cluster. Activation of cluster-specific regulatory factors enabled the isolation of ten structurally novel pyranterreone, pyranterrones A-H (241–248), cordylactam (249), and 7-hydroxycordylactam (250). By integrating gene knockout experiments, bioinformatic analysis, and stable isotope labeling studies, the unique biosynthetic mechanism of thePKS-NRPS hybrid system was veiled in the fungal species [100].
2.2.5 Structure-feature-based gene cluster localization strategies
A fragment-guided strategy, focusing on strained cyclophane as a key biosynthetic fragment, was employed to develop a novel approach for natural product discovery. Cyclophane-containing natural products (CNPs), as a unique class of molecules widely distributed in nature, are formed through the cyclization of precursors such as polyketides or peptide chains. Their distinctive structural features confer important biological activities, like vancomycin. Of particular interest is the biosynthetic pathway of the antiviral compound herquline A [101], in which three key enzymes—HqlA (NRPS), HqlB (SDR), and HqlC (P450)—work synergistically to convert tyrosine into a dityrosine piperazine precursor and ultimately construct the strained cyclophane scaffold. Leveraging the high reactivity and structural plasticity of this cyclophane module, the researchers proposed that the HqlABC gene cluster constitutes a "minimal cyclophane-forming cassette, " serving as an ideal target for discovering structurally innovative natural products. Using this method, octacycline A (251) was isolated from conserved gene clusters. This compound contains a complex octacyclic structure and an unprecedented hetero-[3.3.1] bicyclic framework, providing a new paradigm for expanding the structural diversity of natural products [102] (Fig. 3 and Additional file 1: Fig. S1).
2.2.6 Conservativity mining of core biosynthetic enzymes
Four important agricultural pathogenic fungi: wheat pathogens (Bipolaris sorokiniana and Zymoseptoria tritici), maize leaf spot pathogen (Cercospora zeae-maydis), and tomato leaf mold pathogen (Fulvia fulva) were selected for the subjects. Based on the biosynthetic characteristics of toxins such as fumonisins, the researchers targeted BGCs anchored by highly reducing polyketide synthases (HRPKSs) and identified a highly conserved BGC among these pathogens and Trichoderma species (including the biocontrol strain T. afroharzianum t-22). This BGC encoded five core enzymes: HRPKS, ABH, P450, α-glucosidase, and a predicted terpene cyclase with EH-like features. The tre BGC from T. afroharzianum t-22 additionally contained P450 and O-methyltransferase (O-MeT) genes. By analyzing the unique combination of these enzymes, the researchers predicted that the metabolic products might possess novel structures. The corresponding metabolite, treconorin (252), was successfully identified. Structural analysis revealed that this compound features a terpene-like trans-fused 5,7-bicyclic core skeleton, which the researchers speculate may originate from a (4 + 3) cycloaddition reaction mechanism. This discovery not only expands the structural diversity of fungal-derived natural products but also provides new insights to subsequent biosynthetic mechanism studies [103].
A "biosynthetic enzyme-guided genome mining strategy" was innovatively applied to systematically analyze conserved key biosynthetic enzymes (NRPS and IDO) of marine fungus Neosartorya pseudofischeri F27-1 genomes, leading to the discovery of the novel 1-benzazepine-structured alkaloid-pseudofisnins A and B (253 and 254) [104]. The development of this strategy stemmed from an in-depth understanding of the L-kynurenine (Kyn) metabolic pathway: although Kyn, as a key intermediate in tryptophan metabolism, is widely involved in various physiological processes, its reported use as a structural unit in natural products is extremely rare. Preliminary research revealed that all known Kyn-derived natural products (e.g., nanangelenin A and aspcandine) [105, 106] share a critical feature—their biosynthetic gene clusters invariably contain genes encoding NRPS and IDO. Building on this pattern, the research team used NRPS and IDO as molecular markers to systematically mine fungal genome databases. This approach not only successfully identified the biosynthetic gene cluster for pseudofisnins but also fully elucidated its synthetic pathway through in vivo and in vitro experiments, while characterizing the novel catalytic function of the iterative methyltransferase PseC. The successful implementation of this strategy underscores the significant value of characteristic biosynthetic enzymes as "signposts" for genome mining, providing a paradigm for the targeted discovery of rare scaffold natural products. With the continuous accumulation of fungal genome data, this mining method based on conserved features of key enzymes holds promise as a powerful tool for discovering more Kyn-derived and other types of novel bioactive molecules.
2.3 Integration of genome mining and metabolism analysis
To bridge the gap between genomic predictions and tangible chemical entities, natural products uncovered through genome mining must be synergistically integrated with advanced structural analysis platforms-such as tandem mass spectrometry (LC–MS/MS), and high-resolution NMR [107–109]. This multidisciplinary convergence not only accelerates the dereplication of known compounds but also enables rapid structural elucidation of novel scaffolds, significantly compressing the discovery timeline from months to weeks. Critically, by minimizing resource-intensive isolation of redundant metabolites, this approach optimizes the path to high-value target molecules, thereby enhancing the cost-efficiency of natural product pipelines in drug development [110] (Fig. 5).
Overview of genome mining and metabolism analysis strategies
2.3.1 Integrating homology-based genome mining with HPLC–MS validation techniques
The research team developed an enzyme-guided genome mining strategy based on the key biosynthetic enzyme C-glycosyltransferase (AuCGT) involved in the antibiotic stromemycin pathway. By integrating homology-based genome mining with HPLC–MS validation techniques, they identified multiple BGCs across diverse fungal strains, leading to the characterization of 20 novel aromatic polyketide C/O-glycosides, talarocellmycins A-I (255‒263), carnemycin I (264), phaeomoniecins A-H (265–272) and verapyrones A-B (273–274). Functional characterization of three novel α-pyrone C-glycosyltransferases and their heterologous expression in Aspergillus nidulans enabled combinatorial biosynthesis of unnatural glycosides. Notably, these glycosides demonstrated significant antiviral, antibacterial, and antidiabetic activities [111].
Two polyketide synthase gene clusters (syw1/syw2) were identified from the marine fungus Aspergillus sydowii PKU374 through antiSMASH analysis. The key gene syw1G showed high homology with the known xanthone biosynthesis gene mdpG, indicating the strain's potential for xanthone production. Furthermore, researchers employed UPLC-HRESIMS/MS coupled with GNPS molecular networking to annotate molecular cluster nodes and preliminarily identified several known xanthone compounds. Notably, multiple unresolved nodes in the molecular network suggested the possible existence of xanthone hybrid metabolites containing unknown structural units. Targeted fractionation of xanthone-enriched components ultimately yielded 11 xanthone alkaloids, sydoxanthones F-M (275–282). Among these, compound 277a exhibited significant neuroprotective activity by effectively scavenging H2O2-induced ROS and improving the viability of SH-SY5Y neuronal cells [112].
In addition, the genome mining techniques was used to systematically screen mangrove-derived fungi. Using the polyketide synthase gene AusA from Aspergillus calidoustus as a molecular probe, researchers identified a homologous gene cluster (tam) in the endophytic fungus Talaromyces sp. JNQQJ-4 isolated from Kandelia candel. GNPS molecular network analysis annotated two known DMTD analogs and several unresolved nodes. Through targeted isolation, seven structurally unique 3,5-dimethylorsellinic acid-derived meroterpenoids, talaromeroterpenoids A-G (283‒289) were obtained. Among them, compound 283 featured an unprecedented 7/6/6/6/5/5 hexacyclic system based on a 2,8,18,21-tetraoxa-hexacyclo [12.5.2.13,12.01,16.04,10.016,22] docosane core scaffold, which represented the first novel natural products. Bioactivity evaluation demonstrated that compound 284 significantly inhibited the expression of pro-inflammatory factors and exerted anti-inflammatory effects by modulating the NF-κB signaling pathway [113].
2.3.2 Coupling NMR metabolomics with genome mining
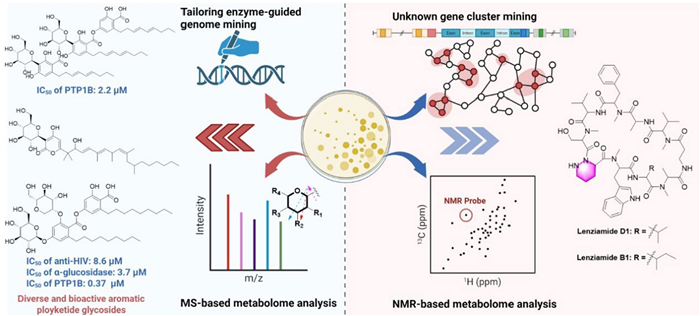
An innovative strategy combining metabolomics and genomics was used to systematically investigate natural products of the marine-derived Streptomyces sp. S063. Through NMR-based metabolomic analysis, rare characteristic signals with negative chemical shifts (δH -0.34, corresponding to a methylene carbon at δC 21.3) were observed for the first time in the bacterial extracts. Meanwhile, the characteristic peaks for amino acid α-hydrogens (δH 4.0–6.0/δC 45.0–75.0) and N-methylation signals (δH 2.7–3.5/δC 25.0–40.0). These findings collectively suggested the presence of structurally novel N-methylated peptide metabolites. Further genome mining revealed a unique nonribosomal peptide synthetase gene cluster (Cluster 20), which not only contained key piperazic acid biosynthesis genes (lenE and lenF) but also exhibited distinctive structural features (including 13 NRPS genes, 10 adenylation domains, and 5 N-methyltransferase domains), with less than 48% homology to known peptide gene clusters. Based on these discoveries, researchers successfully isolated and identified two novel piperazic acid-containing cyclic decapeptides—lenziamides D1 (290) and B1 (291). Notably, both compounds demonstrated moderate growth inhibitory activity against multiple human cancer cell lines (including HEL, H1975, and H1299) as well as the paclitaxel-resistant strain A549-taxol, with IC50 values ranging from 8 to 24 μM. This study not only uncovered a structurally unique class of microbial secondary metabolites but also provided potential candidate molecules for the development of novel antitumor drugs [114] (Fig. 3 and Additional file 1: Fig. S1).
2.4 Machine-learning-driven strategy
2.4.1 AI-facilitated enzyme tailoring platform
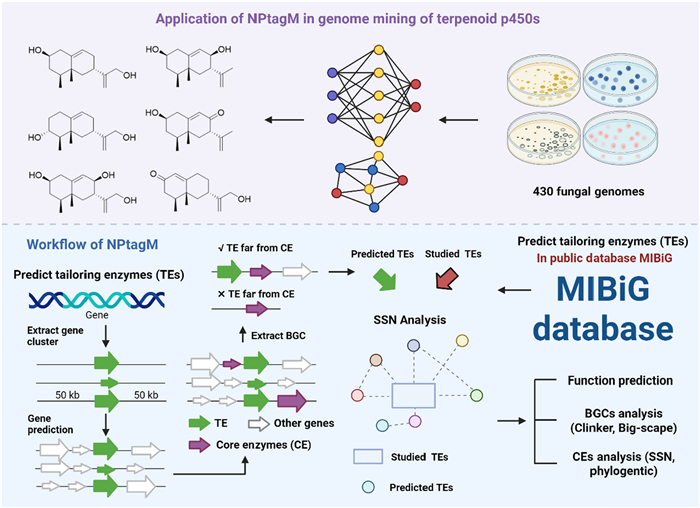
Machine-learning-driven identification of glycosyltransferase/P450 enzyme signatures for tailored natural product diversification. To overcome the limitations of traditional genome mining tools, the research team developed a novel genome mining toolkit, NPtagM, for modifying enzymes. By optimizing algorithmic architecture and workflows, this tool improved the prediction efficiency of P450 enzymes to three times that of the existing tool antiSMASH while reducing runtime by eightfold. Systematic analysis of fungal genomes using NPtagM identified a P450 enzyme family potentially involved in the biosynthesis of eremophilane-type sesquiterpenoids from 1,189 deduplicated terpenoid P450 enzymes. Heterologous expression in Aspergillus oryzae successfully yielded two novel eremophilane-type compounds (292–293) and four known analogues [115] (Fig. 6).
Framework of an AI-facilitated enzyme tailoring system
2.4.2 Intelligent CRISPR-guided genome engineering
Combinatorial deployment of Cas9 nickase with deaminases for DSB-free multiplexed engineering of biosynthetic machineries. Performance metrics in: in gene editing technology, the team developed a revolutionary multiplex base-editing (MBE) platform. By integrating the CRISPR-Cas9 system with base-editing tools, this platform achieved high-throughput gene editing in filamentous fungi, enabling simultaneous editing of up to eight genes in a single transformation. Using this technology, the team combinatorially regulated three epigenetic regulators (CclA, ClrD, and HdaA) in Aspergillus nidulans, successfully activating eight silent BGCs and isolating four structurally novel compounds, 8-oxycichorine (294), andicichorine A (295), andicichorine B (296), andicichorine C (297). In-depth studies revealed that the unique hybrid scaffolds of compounds 295, 296, and 297 likely originated from cross-reactions between the cichorine and polyamine metabolic pathways [116].
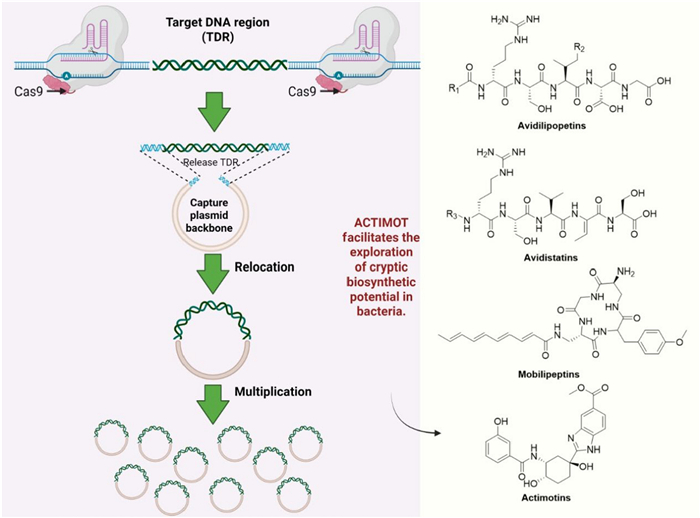
In addition, a CRISPR-Cas9-based gene editing tool named ACTIMOT was developed, which was inspired by the horizontal transfer mechanism of antibiotic resistance genes. This method enables the in situ excision and transfer of large chromosomal DNA fragments (such as silent biosynthetic gene clusters) directly within bacterial cells, efficiently integrating them into autonomously replicating plasmids. This circumvents the limitations of traditional cloning techniques in manipulating large DNA fragments. Applying this technology, the researchers successfully mobilized four cryptic gene clusters from the Streptomyces chromosome. Plasmid-based overexpression activated their biosynthetic pathways, ultimately leading to the discovery of 39 structurally diverse novel natural products (encompassing four distinct classes of compounds, avidistatins A1-A2, B1-B2, C1-C2, D (298–304), avidilipopeptides A1-A5, B1-B4, A6-A11 (305–319), mobilipeptins A-E (320–324), actimotins A-L (325–336)). his work demonstrates that ACTIMOT can accelerate the exploration of cryptic metabolic pathways and the discovery of natural products [117] (Fig. 7).
Framework of intelligent CRISPR-guided genome engineering
2.4.3 Structure-oriented intelligent screening
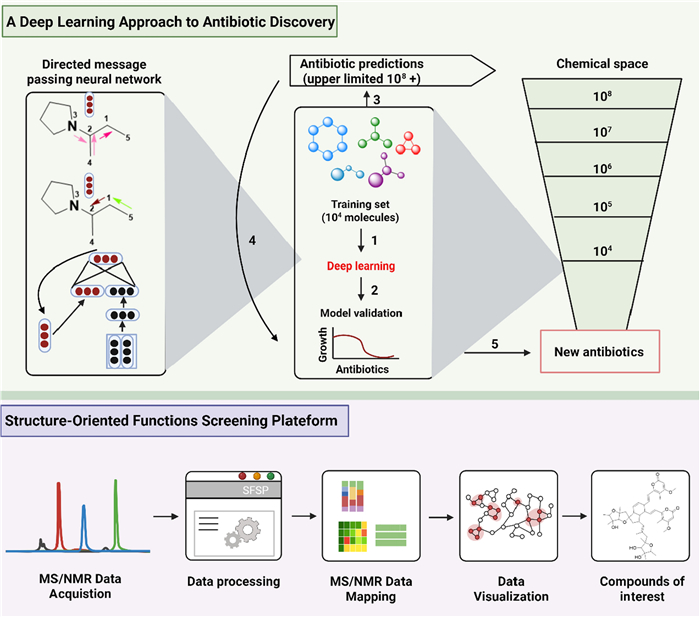
In this study, an antimicrobial activity prediction model was developed based on the deep neural networks. Initially, 2,335 known anti-Escherichia coli active molecules were trained. Then, the model was employed to direct message-passing neural networks to construct molecular representations and integrated multi-source features. After that, its performance was significantly enhanced due to the hyperparameter optimization and ensemble learning. The model was innovatively applied in a dual-track screening strategy: on one hand, it virtually screened over 107 million molecules from the ZINC15 database, leading to the experimental validation of 23 candidates and the identification of 8 novel antibacterial compounds after prediction score ranking; on the other hand, combined with targeted drug repositioning technology, it efficiently identified the structurally innovative antibiotic halicin (337) from the "Drug Repurposing Hub" library. Compound 337 possessing a markedly different structure from conventional antibiotics, exhibited broad-spectrum of antimicrobial activity against clinically critical pathogens including Mycobacterium tuberculosis and carbapenem-resistant Enterobacteriaceae. Especially, compound 337 can be used to surpress Clostridioides difficile and treat pan-drug-resistant Acinetobacter baumannii infections in mouse models [118] (Fig. 8).
Workflow of structure-based intelligent screening
Structure-Oriented Fractions Screening Platform (SFSP) was established to be an efficient natural product discovery system by integrating machine learning algorithms with multidimensional spectroscopic data. Specifically, the SFSP platform enables direct prediction of compound node chemical shifts from extract library MS1 mass spectrometry data and 1H-PSYCHE-NMR spectra, achieving database-independent structural feature analysis. Using this innovative technological platform, we successfully identified two structurally novel classes of natural products from the deep-sea hydrothermal vent-derived fungus Aspergillus sp. GE2-6: Flavipidin-type compounds identified through 1H NMR olefin signal guidance, including flavipidins A-E (338–342), diflavipidins A-Q (343–359), and isoflavipidins A-B (360–361); Phenalenones-type compounds discovered via 1H NMR low-field hydroxy signal guidance: asperphenalenones N-P (362–364). Notably, compound 338 demonstrated significant anti-influenza A virus (PR8 strain) activity (IC50: 21.9/12.9 μM), while compounds 363–364 effectively inhibited HIV pseudovirus infection in 293FT cells (IC50: 6.1/4.6 μM) [119] (Fig. 8).
3 Summary and prospects
The development strategies for microbial natural products each have their own characteristics: the OSMAC strategy can activate silent gene clusters by simply adjusting culture conditions, but it suffers from uncontrollable products and low yields; the structure-based localization strategy is highly targeted but heavily relies on known compound structure databases, limiting its predictive capability for novel structures or rare modification patterns; the conservative mining strategy of core biosynthetic enzymes can rapidly locate key genes through conserved functional domains, making it suitable for high-throughput screening of homologous gene clusters, but it may miss atypical biosynthetic enzymes or functionally differentiated enzymes, and the conservative analysis cannot directly reflect the activity of gene clusters or the structural diversity of products, requiring experimental validation; in omics integration strategies, HPLC–MS offers high sensitivity but depends on existing mass spectrometry libraries, while NMR provides precise structural resolution but has limited sensitivity; in machine learning-driven intelligent strategies, the NPtagM targeted enzyme modification platform can automatically identify glycosylation, methylation, and other modifying enzymes, accelerating structural derivatization, but it relies on training sets of known modifying enzymes and performs poorly in predicting non-classical modifications; CRISPR base-editing technology can precisely activate silent gene clusters, directly establishing gene-product associations, making it suitable for hard-to-culture microorganisms; deep neural network prediction can learn complex structure–activity relationships, enabling virtual screening and significantly reducing experimental workload, but it requires large sample sizes for stable training.
The core contradiction in current research lies in the fact that while microbial genomic data is growing rapidly (with over 200,000 genomes sequenced), the proportion of systematic studies remains extremely low [120–122]. This gap highlights both the enormous potential and the significant challenges in the development of microbial secondary metabolites [123, 124]. Notably, microbial genetic diversity far exceeds expectations [125]. Taking fungi as an example, their genome size and number of gene clusters far surpass those of bacteria, but complex regulatory networks and bottlenecks in genetic manipulation severely hinder research progress [126].
Future breakthroughs should focus on three dimensions: at the technological innovation level, new cultivation techniques and single-cell analysis methods need to be developed to overcome the challenges of accessing unculturable microbial resources; in terms of strategy optimization, deeper integration of multi-omics is required, particularly the establishment of "genome-transcriptome-metabolome" correlation networks; the key to methodological innovation lies in interdisciplinary integration, including: (1) bioinformatics-driven intelligent prediction, (2) synthetic biology-guided rational design, and (3) microfluidic technology-enabled high-throughput screening. With breakthroughs in CRISPR and other technologies [27], research on microbial secondary metabolism has entered a golden development period. The construction of a "data-driven experimentation, experimental validation, and engineering optimization" research framework holds promise for delivering transformative breakthroughs in pharmaceutical development and agricultural applications.
Notes
Acknowledgements
This work was supported by the Yunnan Revitalization Talent Support Program "Young Talent" Project (XDYC-QNRC-2022–0480), and the Basic Research Program of Yunnan Province (202301AT070332).
Author contributions
Yu-Jie Li: Conceptualization, Investigation, Formal analysis, Methodology, Data Curation, Visualization, Writing—Original Draft. Ming-Hua Qiu: Investigation. Xing-Rong Peng: Supervision, Writing—Review and Editing, Project administration, Funding acquisition. The author(s) read and approved the final manuscript.
Data availability
No data was used for the research described in the article.
Declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
-
1.Atanasov AG, Zotchev SB, Dirsch VM, et al. Natural products in drug discovery: advances and opportunities. Nat Rev Drug Discov. 2021;20: 200-16. CrossRef PubMed Google Scholar
-
2.Newman DJ, Cragg GM. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J Nat Prod. 2020;83(3): 770-803. CrossRef PubMed Google Scholar
-
3.Lewis K. The science of antibiotic discovery. Cell. 2020;181(1): 29-45. CrossRef PubMed Google Scholar
-
4.Ajikumar PK, Xiao W-H, Tyo KEJ, et al. Isoprenoid pathway optimization for taxol precursor overproduction in Escherichia coli. Science. 2010;330(6000): 70-4. CrossRef PubMed Google Scholar
-
5.Zhu L, Chen L. Progress in research on paclitaxel and tumor immunotherapy. Cell Mol Biol Lett. 2019;24(1): 40. CrossRef PubMed Google Scholar
-
6.Wang J, Liang J, Chen L, et al. Structural basis for the biosynthesis of lovastatin. Nat Commun. 2021;12(1): 867. CrossRef PubMed Google Scholar
-
7.Sheng N, Wang Y-Q, Wang C-F, et al. AGR2-induced cholesterol synthesis drives lovastatin resistance that is overcome by combination therapy with allicin. Acta Pharmacol Sin. 2022;43(11): 2905-16. CrossRef PubMed Google Scholar
-
8.Zhang Y, Zong J, Liu Y, et al. Progress on targeted discovery of microbial natural products based on the predictions of both structure and activity. Nat Prod Rep. 2025. CrossRef PubMed Google Scholar
-
9.Wang DG, Hu JQ, Wang CY, et al. Exploring microbial natural products through NMR-based metabolomics. Nat Prod Rep. 2025;42: 1459-88. CrossRef PubMed Google Scholar
-
10.Liu R, Peng XP, Newman DJ, et al. Unlocking the metabolic potential of endophytic fungi through epigenetics: a paradigm shift for natural product discovery and plant–microbe interactions. Nat Prod Rep. 2025. CrossRef PubMed Google Scholar
-
11.Kulkarni AS, Carrara GMP, Jin J, et al. Mass spectrometry-based metabolomics approaches to interrogate host–microbiome interactions in mammalian systems. Nat Prod Rep. 2025. CrossRef PubMed Google Scholar
-
12.Chen B, Bülbül EF, Bang S, et al. Toward a unified pipeline for natural product discovery: tools and strategies for NRPS and PKS pathway exploration and engineering. Nat Prod Rep. 2025. CrossRef PubMed Google Scholar
-
13.Zhang Y, Feng L, Hemu X, et al. Osmac strategy: a promising way to explore microbial cyclic peptides. Eur J Med Chem. 2024;268: 116175. CrossRef PubMed Google Scholar
-
14.Yuan MX, Qiu Y, Ran YQ, et al. Exploration of indole alkaloids from marine fungus Pseudallescheria boydii F44–1 using an amino acid-directed strategy. Mar Drugs. 2019;17(2): 77. CrossRef PubMed Google Scholar
-
15.Tran-Cong NM, Mándi A, Kurtán T, et al. Induction of cryptic metabolites of the endophytic fungus Trichocladium sp. through OSMAC and co-cultivation. RSC Adv. 2019;9: 27279-88. CrossRef PubMed Google Scholar
-
16.Gao Y, Stuhldreier F, Schmitt L, et al. Induction of new lactam derivatives from the endophytic fungus Aplosporella javeedii through an OSMAC approach. Front Microbiol. 2020;11: 600983. CrossRef PubMed Google Scholar
-
17.Chen Y, Yang W, Zou G, et al. Cytotoxic bromine- and iodine-containing cytochalasins produced by the mangrove endophytic fungus Phomopsis sp. QYM-13 using the OSMAC approach. J Nat Prod. 2022;85(5): 1229-38. CrossRef PubMed Google Scholar
-
18.Chen M, Zheng YY, Chen ZQ, et al. NaBr-induced production of brominated azaphilones and related tricyclic polyketides by the marine-derived fungus Penicillium janthinellum HK1-6. J Nat Prod. 2019;82(2): 368-74. CrossRef PubMed Google Scholar
-
19.Lin S, Huang JZ, Zeng HX, et al. Distachydrimanes A–F, phenylspirodrimane dimers and hybrids with cytotoxic activity from the coral-derived fungus Stachybotrys chartarum. Chin Chem Lett. 2022;33(10): 4587-94. CrossRef PubMed Google Scholar
-
20.Chen HL, Zhao WT, Liu QP, et al. (±)-preisomide: a new alkaloid featuring a rare naturally occurring tetrahydro-2H-1,2-oxazin skeleton from an endophytic fungus Preussia isomera by using OSMAC strategy. Fitoterapia. 2020;141: 104475. CrossRef PubMed Google Scholar
-
21.Xu LL, Chen HL, Hai P, et al. (+)- and (−)-preuisolactone A: a pair of caged norsesquiterpenoidal enantiomers with a tricyclo[4.4.01,6.02,8]decane carbon skeleton from the endophytic fungus Preussia isomera. Org Lett. 2019;21(4): 1078-81. CrossRef PubMed Google Scholar
-
22.Sun J, Yang XQ, Wan JL, et al. The antifungal metabolites isolated from maize endophytic fungus Fusarium sp. induced by OSMAC strategy. Fitoterapia. 2023;171: 105710. CrossRef PubMed Google Scholar
-
23.Zang Y, Gong Y, Shi Z, et al. Multioxidized aromatic polyketides produced by a soil-derived fungus Penicillium canescens. Phytochemistry. 2021;193: 113012. CrossRef PubMed Google Scholar
-
24.Rodríguez Martín-Aragón V, Trigal Martínez M, Cuadrado C, et al. Osmac approach and cocultivation for the induction of secondary metabolism of the fungus Pleotrichocladium opacum. ACS Omega. 2023;8: 39873-85. CrossRef PubMed Google Scholar
-
25.González-Menéndez V, Pérez-Bonilla M, Pérez-Victoria I, et al. Multicomponent analysis of the differential induction of secondary metabolite profiles in fungal endophytes. Molecules. 2016;21(2): 234. CrossRef PubMed Google Scholar
-
26.Cichewicz RH. Epigenome manipulation as a pathway to new natural product scaffolds and their congeners. Nat Prod Rep. 2010;27(1): 11-22. CrossRef PubMed Google Scholar
-
27.Rutledge PJ, Challis GL. Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat Rev Microbiol. 2015;13(8): 509-23. CrossRef PubMed Google Scholar
-
28.Williams RB, Henrikson JC, Hoover AR, et al. Epigenetic remodeling of the fungal secondary metabolome. Org Biomol Chem. 2008;6(11): 1895-7. CrossRef PubMed Google Scholar
-
29.Fisch KM, Gillaspy AF, Gipson M, et al. Chemical induction of silent biosynthetic pathway transcription in Aspergillus niger. J Ind Microbiol Biotechnol. 2009;36(9): 1199-213. CrossRef PubMed Google Scholar
-
30.Wang X, Sena Filho JG, Hoover AR, et al. Chemical epigenetics alters the secondary metabolite composition of guttate excreted by an atlantic-forest-soil-derived Penicillium citreonigrum. J Nat Prod. 2010;73(5): 942-8. CrossRef PubMed Google Scholar
-
31.Brakhage AA, Schroeckh V. Fungal secondary metabolites: strategies to activate silent gene clusters. Fungal Genet Biol. 2011;48(1): 15-22. CrossRef PubMed Google Scholar
-
32.Gacek A, Strauss J. The chromatin code of fungal secondary metabolite gene clusters. Appl Microbiol Biotechnol. 2012;95(6): 1389-404. CrossRef PubMed Google Scholar
-
33.Cai XY, Li N, Li Y, et al. An epigenetic modifier enhances the generation of anti-phytopathogenic compounds from the endophytic fungus Chaetomium globosporum of Euphorbia humifusa. Phytochemistry. 2022;203: 113426. CrossRef PubMed Google Scholar
-
34.Li C, Shao Y, Li W, et al. Hybrid diterpenic meroterpenoids from an endophytic Penicillium sp. induced by chemical epigenetic manipulation. J Nat Prod. 2022;85(6): 1486-94. CrossRef PubMed Google Scholar
-
35.Niu S, Liu D, Shao Z, et al. Chemical epigenetic manipulation triggers the production of sesquiterpenes from the deep-sea derived Eutypella fungus. Phytochemistry. 2021;192: 112978. CrossRef PubMed Google Scholar
-
36.Wu J-S, Yao G-S, Shi X-H, et al. Epigenetic agents trigger the production of bioactive nucleoside derivatives and bisabolane sesquiterpenes from the marine-derived fungus Aspergillus versicolor. Front Microbiol. 2020;11: 85. CrossRef PubMed Google Scholar
-
37.Netzker T, Flak M, Krespach MKC, et al. Microbial interactions trigger the production of antibiotics. Curr Opin Microbiol. 2018;45: 117-23. CrossRef PubMed Google Scholar
-
38.Adnani N, Rajski SR, Bugni TS. Symbiosis-inspired approaches to antibiotic discovery. Nat Prod Rep. 2017;34(7): 784-814. CrossRef PubMed Google Scholar
-
39.Bertrand S, Bohni N, Schnee S, et al. Metabolite induction via microorganism co-culture: a potential way to enhance chemical diversity for drug discovery. Biotechnol Adv. 2014;32(6): 1180-204. CrossRef PubMed Google Scholar
-
40.Nielsen JC, Grijseels S, Prigent S, et al. Global analysis of biosynthetic gene clusters reveals vast potential of secondary metabolite production in Penicillium species. Nat Microbiol. 2017;2(6): 17044. CrossRef PubMed Google Scholar
-
41.Brakhage AA. Regulation of fungal secondary metabolism. Nat Rev Microbiol. 2013;11(1): 21-32. CrossRef PubMed Google Scholar
-
42.Keller NP. Fungal secondary metabolism: regulation, function and drug discovery. Nat Rev Microbiol. 2019;17(3): 167-80. CrossRef PubMed Google Scholar
-
43.Henrikson JC, Hoover AR, Joyner PM, et al. A chemical epigenetics approach for engineering the in situbiosynthesis of a cryptic natural product from Aspergillus niger. Org Biomol Chem. 2009;7(3): 435-8. CrossRef PubMed Google Scholar
-
44.Meng LH, Li XM, Li HL, et al. Chermebilaenes A and B, new bioactive meroterpenoids from co-cultures of marine-derived isolates of Penicillium bilaiae MA-267 and Penicillium chermesinum EN-480. Mar Drugs. 2020;18: 339. CrossRef PubMed Google Scholar
-
45.Yang W, Yuan J, Tan Q, et al. Peniazaphilones A—I, produced by co-culturing of mangrove endophytic fungi, Penicillium sclerotiorum THSH-4 and Penicillium sclerotiorum ZJHJJ-18. Chin J Chem. 2021;39: 3404-12. CrossRef PubMed Google Scholar
-
46.Stierle AA, Stierle DB, Decato D, et al. Cryptic biosynthesis of the berkeleypenostatins from coculture of extremophilic Penicillium sp. J Nat Prod. 2021;84: 1656-65. CrossRef PubMed Google Scholar
-
47.Li F, Yan S, Huang Z, et al. Inducing new bioactive metabolites production from coculture of Pestalotiopsis sp. and Penicillium bialowiezense. Bioorg Chem. 2021;110: 104826. CrossRef PubMed Google Scholar
-
48.Cowled MS, Kalaitzis JA, Crombie A, et al. Fungal duel between Penicillium brasilianum and Aspergillus nomius results in dual induction of miktospiromide A and kitrinomycin A. J Nat Prod. 2023;86: 2398-406. CrossRef PubMed Google Scholar
-
49.de Vries RP, Riley R, Wiebenga A, et al. Comparative genomics reveals high biological diversity and specific adaptations in the industrially and medically important fungal genus Aspergillus. Genome Biol. 2017;18(1): 28. CrossRef PubMed Google Scholar
-
50.El-hawary SS, Moawad AS, Bahr HS, et al. Natural product diversity from the endophytic fungi of the genus Aspergillus. RSC Adv. 2020;10(37): 22058-79. CrossRef PubMed Google Scholar
-
51.Fleißner A, Dersch P. Expression and export: recombinant protein production systems for Aspergillus. Appl Microbiol Biotechnol. 2010;87(4): 1255-70. CrossRef PubMed Google Scholar
-
52.Barrios-González J, Miranda RU. Biotechnological production and applications of statins. Appl Microbiol Biotechnol. 2010;85(4): 869-83. CrossRef PubMed Google Scholar
-
53.Knowles SL, Raja HA, Isawi IH, et al. Wheldone: characterization of a unique scaffold from the coculture of Aspergillus fischeri and Xylaria flabelliformis. Org Lett. 2020;22: 1878-82. CrossRef PubMed Google Scholar
-
54.Shen X, Lei C, Zhang A, et al. Stimulating novel and bioactive metabolite production by cocultivation of two fungi─Aspergillus oryzae and Epicoccum dendrobii. J Agric Food Chem. 2024;72: 16825-34. CrossRef PubMed Google Scholar
-
55.Li Q, Li S, Li S, et al. Antimicrobial and anti-inflammatory cyclic tetrapeptides from the co-cultures of two marine-derived fungi. J Nat Prod. 2024;87: 365-70. CrossRef PubMed Google Scholar
-
56.Meng Z, Tan Y, Duan Y-L, et al. Monaspin B, a novel cyclohexyl-furan from cocultivation of Monascus purpureus and Aspergillus oryzae, exhibits potent antileukemic activity. J Agric Food Chem. 2024;72(2): 1114-23. CrossRef PubMed Google Scholar
-
57.Belousova EB, Zhuravleva OI, Yurchenko EA, et al. New anti-hypoxic metabolites from co-culture of marine-derived fungi Aspergillus carneus KMM 4638 and Amphichorda sp. KMM 4639. Biomolecules. 2023;13(5): 741. CrossRef PubMed Google Scholar
-
58.Edjah PP, Lu M-F, Chen W-C, et al. Aculeaquamides B and C, two new paraherquamides from the co-culture of Aspergillus aculeatinus WHUF0198 and Penicillium sp. DM27. Fitoterapia. 2024;179: 106251. CrossRef PubMed Google Scholar
-
59.Chen W-C, Song X, Wu J, et al. Aculeapyridones A-Q, pyranopyridone alkaloids with protective effects against acetaminophen-induced acute liver injury discovered from a coculture of Aspergillus aculeatinus WHUF0198 and a Penicillium sp. J Nat Prod. 2025;88(2): 336-48. CrossRef PubMed Google Scholar
-
60.Pérez-Montes A, Rangel-Vargas E, Lorenzo JM, et al. Edible mushrooms as a novel trend in the development of healthier meat products. Curr Opin Food Sci. 2021;37: 118-24. CrossRef PubMed Google Scholar
-
61.Singdevsachan SK, Auroshree P, Mishra J, et al. Mushroom polysaccharides as potential prebiotics with their antitumor and immunomodulating properties: a review. Bioact Carbohydr Diet Fibre. 2016;7(1): 1-14. CrossRef PubMed Google Scholar
-
62.Wu ZW, Liu XC, Quan CX, et al. Novel galactose-rich polysaccharide from Ganoderma lucidum: structural characterization and immunomodulatory activities. Carbohydr Polym. 2025;362: 123695. CrossRef PubMed Google Scholar
-
63.Guo C, Guo D, Fang L, et al. Ganoderma lucidum polysaccharide modulates gut microbiota and immune cell function to inhibit inflammation and tumorigenesis in colon. Carbohydr Polym. 2021;267: 118231. CrossRef PubMed Google Scholar
-
64.Das J, Fatmi MQ, Devi M, et al. Antitumor activity of cordycepin in murine malignant tumor cell line: an in vitro and in silico study. J Mol Struct. 2024;1297: 136946. CrossRef PubMed Google Scholar
-
65.Guo L, Xi B, Lu L. Strategies to enhance production of metabolites in microbial co-culture systems. Bioresour Technol. 2024;406: 131049. CrossRef PubMed Google Scholar
-
66.Ge X, Wang Y, Sun C, et al. Secondary metabolites produced by coculture of Pleurotus ostreatus SY10 and Pleurotus eryngii SY302. Chem Biodivers. 2022;19: e202100832. CrossRef PubMed Google Scholar
-
67.Yu G, Ge X, Wang Y, et al. Discovery of novel terpenoids from the basidiomycete Pleurotus ostreatus through genome mining and coculture optimization. J Agric Food Chem. 2023;71: 11110-23. CrossRef PubMed Google Scholar
-
68.Ji L, Tan L, Shang Z, et al. Discovery of new antimicrobial metabolites in the coculture of medicinal mushrooms. J Agric Food Chem. 2024;72: 5247-57. CrossRef PubMed Google Scholar
-
69.Sofian FF, Suzuki T, Supratman U, et al. Cochlioquinone derivatives produced by coculture of endophytes, Clonostachys rosea and Nectria pseudotrichia. Fitoterapia. 2021;155: 105056. CrossRef PubMed Google Scholar
-
70.Pham HT, Doan TP, Kim HW, et al. Cyclohumulanoid sesquiterpenes induced by the noncompetitive coculture of Phellinus orientoasiaticus and Xylodon flaviporus. J Nat Prod. 2022;85: 511-8. CrossRef PubMed Google Scholar
-
71.Wu YM, Zhou QY, Yang XQ, et al. Induction of antiphytopathogenic metabolite and squalene production and phytotoxin elimination by adjustment of the mode of fermentation in cocultures of phytopathogenic Nigrospora oryzae and Irpex lacteus. J Agric Food Chem. 2019;67: 11877-82. CrossRef PubMed Google Scholar
-
72.Sadahiro Y, Kato H, Williams RM, et al. Irpexine, an isoindolinone alkaloid produced by coculture of endophytic fungi, Irpex lacteus and Phaeosphaeria oryzae. J Nat Prod. 2020;83: 1368-73. CrossRef PubMed Google Scholar
-
73.Chen JX, Yang XQ, Wang XY, et al. Antifeedant, antifungal cryptic polyketides with six structural frameworks from tea endophyte Daldinia eschscholtzii propelled by the antagonistic Coculture with phytopathogen Colletotrichum pseudomajus and different culture methods. J Agric Food Chem. 2023;72: 378-89. CrossRef PubMed Google Scholar
-
74.Li H-T, Zhou H, Duan R-T, et al. Inducing secondary metabolite production by co-culture of the endophytic fungus Phoma sp. and the symbiotic fungus Armillaria sp. J Nat Prod. 2019;82: 1009-13. CrossRef PubMed Google Scholar
-
75.Oppong-Danquah E, Blümel M, Scarpato S, et al. Induction of isochromanones by co-cultivation of the marine fungus Cosmospora sp. and the phytopathogen Magnaporthe oryzae. Int J Mol Sci. 2022;23: 782. CrossRef PubMed Google Scholar
-
76.Wang ZJ, Hong YR, Wang XY, et al. Fungal coculture of Herpotrichia sp. and Trametes versicolor induces production of diverse metabolites with anti-parkinson’s neuroprotective activity. J Nat Prod. 2024;87(9): 2180-93. CrossRef PubMed Google Scholar
-
77.Put H, Gerstmans H, Vande Capelle H, et al. Bacillus subtilis as a host for natural product discovery and engineering of biosynthetic gene clusters. Nat Prod Rep. 2024;41(7): 1113-51. CrossRef PubMed Google Scholar
-
78.Keller L, Surette MG. Communication in bacteria: an ecological and evolutionary perspective. Nat Rev Microbiol. 2006;4(4): 249-58. CrossRef PubMed Google Scholar
-
79.Eze PM, Liu Y, Simons VE, et al. Two new metabolites from a marine-derived fungus Penicillium ochrochloron. Phytochem Lett. 2023;55: 101-4. CrossRef PubMed Google Scholar
-
80.Abdel-Wahab N, Scharf S, Özkaya F, et al. Induction of secondary metabolites from the marine-derived fungus Aspergillus versicolor through co-cultivation with Bacillus subtilis. Planta Med. 2019;85: 503-12. CrossRef PubMed Google Scholar
-
81.Thissera B, Alhadrami HA, Hassan MHA, et al. Induction of cryptic antifungal pulicatin derivatives from Pantoea agglomerans by microbial co-culture. Biomolecules. 2020;10: 268. CrossRef PubMed Google Scholar
-
82.Hwang GJ, Jang M, Son S, et al. Ulleungdolin, a polyketide–peptide hybrid bearing a 2,4-Di-O-methyl-β-d-antiarose from Streptomyces sp. 13F051 co-cultured with Leohumicola minima 15S071. J Nat Prod. 2022;85: 2445-53. CrossRef PubMed Google Scholar
-
83.Huynh TH, Lee J, Moon DH, et al. Gwanakosides A and B, 6-Deoxy-α-l-talopyranose-bearing aromatic metabolites from a Streptomyces sp. and coculture with Pandoraea sp. J Nat Prod. 2021;85: 83-90. CrossRef PubMed Google Scholar
-
84.Asamizu S, Pramana AAC, Kawai S-J, et al. Comparative metabolomics reveals a bifunctional antibacterial conjugate from combined-culture of Streptomyces hygroscopicus HOK021 and Tsukamurella pulmonis TP-B0596. ACS Chem Biol. 2022;17: 2664-72. CrossRef PubMed Google Scholar
-
85.Alhadrami HA, Thissera B, Hassan MHA, et al. Bio-guided isolation of antimalarial metabolites from the coculture of two red sea sponge-derived Actinokineospora and Rhodococcus spp. Mar Drugs. 2021;19(2): 109. CrossRef PubMed Google Scholar
-
86.Kaeberlein T, Lewis K, Epstein SS. Isolating “uncultivable” microorganisms in pure culture in a simulated natural environment. Science. 2002;296(5570): 1127-9. CrossRef PubMed Google Scholar
-
87.Nichols D, Cahoon N, Trakhtenberg EM, et al. Use of ichip for high-throughput in situ cultivation of microbial species. Appl Environ Microbiol. 2010;76(8): 2445-50. CrossRef PubMed Google Scholar
-
88.Ling LL, Schneider T, Peoples AJ, et al. A new antibiotic kills pathogens without detectable resistance. Nature. 2015;517(7535): 455-9. CrossRef PubMed Google Scholar
-
89.Jiang CY, Dong L, Zhao JK, et al. High-throughput single-cell cultivation on microfluidic streak plates. Appl Environ Microbiol. 2016;82(7): 2210-8. CrossRef PubMed Google Scholar
-
90.Zengler K, Toledo G, Rappé M, et al. Cultivating the uncultured. Proc Natl Acad Sci U S A. 2002;99(24): 15681-6. CrossRef PubMed Google Scholar
-
91.Corre C, Challis GL. New natural product biosynthetic chemistry discovered by genome mining. Nat Prod Rep. 2009;26: 977-86. CrossRef PubMed Google Scholar
-
92.Woodcraft C, Chooi Y-H, Roux I. The expanding CRISPR toolbox for natural product discovery and engineering in filamentous fungi. Nat Prod Rep. 2023;40: 158-73. CrossRef PubMed Google Scholar
-
93.Zhao F, Sun C, Liu Z, et al. Multiplex base-editing enables combinatorial epigenetic regulation for genome mining of fungal natural products. J Am Chem Soc. 2022;145(1): 413-21. CrossRef PubMed Google Scholar
-
94.Ning Y, Gu Q, Zheng T, et al. Genome mining leads to diverse sesquiterpenes with anti-inflammatory activity from an arctic-derived fungus. J Nat Prod. 2024;87: 1426-40. CrossRef PubMed Google Scholar
-
95.Yang Z, Qiao Y, Strøbech E, et al. Alligamycin A, an antifungal β-lactone spiroketal macrolide from Streptomyces iranensis. Nat Commun. 2024;15(1): 9259. CrossRef PubMed Google Scholar
-
96.Cheng JT, Yu JH, Sun CF, et al. A cell factory of a fungicolous fungus Calcarisporium arbuscula for efficient production of natural products. ACS Synth Biol. 2021;10: 698-706. CrossRef PubMed Google Scholar
-
97.Cain JW, Miller KI, Kalaitzis JA, et al. Genome mining of a fungal endophyte of Taxus yunnanensis (Chinese yew) leads to the discovery of a novel azaphilone polyketide, lijiquinone. Microb Biotechnol. 2020;13(5): 1415-27. CrossRef PubMed Google Scholar
-
98.Huang X, Li D, Long B, et al. Activation of a silent gene cluster from the endophytic fungus Talaromyces sp. unearths cryptic azaphilone metabolites. J Agric Food Chem. 2024;72: 15801-10. CrossRef PubMed Google Scholar
-
99.Cheng JT, Wang HM, Yu JH, et al. Discovery of a potential liver fibrosis inhibitor from a mushroom endophytic fungus by genome mining of a silent biosynthetic gene cluster. J Agric Food Chem. 2021;69: 11303-10. CrossRef PubMed Google Scholar
-
100.Tang S, Zhang W, Li Z, et al. Discovery and characterization of a PKS-NRPS hybrid in Aspergillus terreus by genome mining. J Nat Prod. 2020;83: 473-80. CrossRef PubMed Google Scholar
-
101.Yu X, Liu F, Zou Y, et al. Biosynthesis of strained piperazine alkaloids: uncovering the concise pathway of herquline A. J Am Chem Soc. 2016;138(41): 13529-32. CrossRef PubMed Google Scholar
-
102.Zhang Y, Wu L, Kerr TA, et al. Fragment-guided genome mining of octacyclic cyclophane alkaloids from fungi. J Am Chem Soc. 2024;146: 23933-42. CrossRef PubMed Google Scholar
-
103.Yan C, Han W, Zhou Q, et al. Genome mining from agriculturally relevant fungi led to a d-glucose esterified polyketide with a terpene-like core structure. J Am Chem Soc. 2023;145: 25080-5. CrossRef PubMed Google Scholar
-
104.Xue XX, Chen L, Tang MC. Genome mining discovery of a new benzazepine alkaloid pseudofisnin A from the marine fungus Neosartorya pseudofischeri F27–1. Antibiotics. 2022;11: 1444. CrossRef PubMed Google Scholar
-
105.Li H, Gilchrist CLM, Phan C-S, et al. Biosynthesis of a new benzazepine alkaloid nanangelenin a from Aspergillus nanangensis involves an unusual l-kynurenine-incorporating NRPS catalyzing regioselective lactamization. J Am Chem Soc. 2020;142(15): 7145-52. CrossRef PubMed Google Scholar
-
106.Chen L, Tang JW, Liu YY, et al. Aspcandine: a pyrrolobenzazepine alkaloid synthesized by a fungal nonribosomal peptide synthetase-polyketide synthase hybrid. Org Lett. 2022;24(26): 4816-9. CrossRef PubMed Google Scholar
-
107.Wang M, Carver JJ, Phelan VV, et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat Biotechnol. 2016;34(8): 828-37. CrossRef PubMed Google Scholar
-
108.Wishart DS, Sayeeda Z, Budinski Z, et al. NP-MRD: the natural products magnetic resonance database. Nucleic Acids Res. 2022;50(D1): D665-77. CrossRef PubMed Google Scholar
-
109.Sigrist R, Paulo BS, Angolini CFF, et al. Mass spectrometry-guided genome mining as a tool to uncover novel natural products. J Vis Exp. 2020;157: e60825. CrossRef PubMed Google Scholar
-
110.van der Hooft JJJ, Mohimani H, Bauermeister A, et al. Linking genomics and metabolomics to chart specialized metabolic diversity. Chem Soc Rev. 2020;49(11): 3297-314. CrossRef PubMed Google Scholar
-
111.Chen D, Song Z, Han J, et al. Targeted discovery of glycosylated natural products by tailoring enzyme-guided genome mining and MS-based metabolome analysis. J Am Chem Soc. 2024;146: 9614-22. CrossRef PubMed Google Scholar
-
112.Ge R, Guo X, Jia H, et al. Nucleobase-coupled xanthones with anti-ROS effects from marine-derived fungus Aspergillus sydowii. J Org Chem. 2024;89: 7692-704. CrossRef PubMed Google Scholar
-
113.Wang G, Li Z, Wu J, et al. Mining of meroterpenoids with anti-inflammatory activity from the mangrove endophytic fungus Talaromyces sp. JNQQJ-4 by genome analysis and molecular network strategy. Org Lett. 2025;27(7): 1662-7. CrossRef PubMed Google Scholar
-
114.Huang H, Yue L, Deng F, et al. NMR-metabolomic profiling and genome mining drive the discovery of cyclic decapeptides from a marine Streptomyces. J Nat Prod. 2023;86: 2122-30. CrossRef PubMed Google Scholar
-
115.Zhanren C, Njeru Joe M, Qiang Y, et al. NPtagM: a tailoring enzyme genome mining toolkit and its application in terpenoid P450s from phytopathogenic fungi. J Agric Food Chem. 2024;72: 27225-34. CrossRef PubMed Google Scholar
-
116.Zhao F, Sun C, Liu Z, et al. Multiplex base-editing enables combinatorial epigenetic regulation for genome mining of fungal natural products. J Am Chem Soc. 2023;145: 413-21. CrossRef PubMed Google Scholar
-
117.Xie F, Zhao H, Liu J, et al. Autologous DNA mobilization and multiplication expedite natural products discovery from bacteria. Science. 2024;386(6727): eabq7333. CrossRef PubMed Google Scholar
-
118.Stokes JM, Yang K, Swanson K, et al. A deep learning approach to antibiotic discovery. Cell. 2020;180(4): 688-702.e13. CrossRef PubMed Google Scholar
-
119.Ge Y, Zhou C, Ma Y, et al. Advancing natural product discovery: a structure-oriented fractions screening platform for compound annotation and isolation. Anal Chem. 2024;96(14): 5399-406. CrossRef PubMed Google Scholar
-
120.Kautsar SA, Blin K, Shaw S, et al. MIBiG 2.0: a repository for biosynthetic gene clusters of known function. Nucleic Acids Res. 2020;48(D1): D454-8. CrossRef PubMed Google Scholar
-
121.Nayfach S, Roux S, Seshadri R, et al. A genomic catalog of earth’s microbiomes. Nat Biotechnol. 2021;39(4): 499-509. CrossRef PubMed Google Scholar
-
122.Gavriilidou A, Kautsar SA, Zaburannyi N, et al. Compendium of specialized metabolite biosynthetic diversity encoded in bacterial genomes. Nat Microbiol. 2022;7(5): 726-35. CrossRef PubMed Google Scholar
-
123.Ziemert N, Alanjary M, Weber T. The evolution of genome mining in microbes: a review. Nat Prod Rep. 2016;33(8): 988-1005. CrossRef PubMed Google Scholar
-
124.Crits-Christoph A, Diamond S, Butterfield CN, et al. Novel soil bacteria possess diverse genes for secondary metabolite biosynthesis. Nature. 2018;558(7710): 440-4. CrossRef PubMed Google Scholar
-
125.Naranjo-Ortiz MA, Gabaldón T. Fungal evolution: cellular, genomic and metabolic complexity. Biol Rev. 2020;95(5): 1198-232. CrossRef PubMed Google Scholar
-
126.Kjærbølling I, Vesth TC, Frisvad JC, et al. Linking secondary metabolites to gene clusters through genome sequencing of six diverse Aspergillus species. Proc Natl Acad Sci U S A. 2018;115(4): E753-61. CrossRef PubMed Google Scholar
Copyright information
© The Author(s) 2026
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.