Diverse phloroglucinols with hAChE inhibitory and anti-VRE effects from Rhodomyrtus tomentosa fruits
Abstract
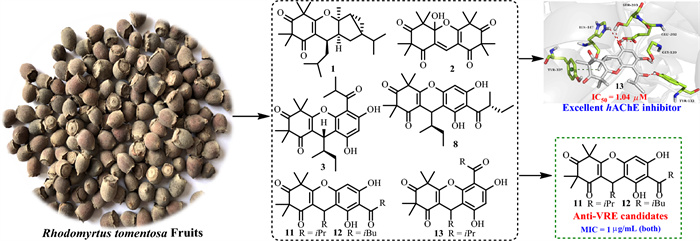
Rhodomyrtus tomentosa fruits serve as both functional food and medicinal resources due to their rich bioactive constituents and manifold pharmacological effects. Phytochemical exploration of the R. tomentosa fruits led to the identification of eight new polymethylated phloroglucinols, designated as rhodotomentodione F (1) and rhodotomentodimers H‒N (2‒8), along with six previously described congeners (9‒14). Based on the detailed inspection of comprehensive spectroscopic data, electronic circular dichroism (ECD) simulations, and nuclear magnetic resonance (NMR) calculations, and DP4+ analyses, the structures of phloroglucinols 1‒8 were determined. Heterodimeric phloroglucinols 3‒14 exhibited human acetylcholinesterase (hAChE) inhibitory activities, with 13 exhibiting the highest potency (IC50 = 1.04 μM). Moreover, molecular docking analysis clarified the potential binding interactions between the most active phloroglucinol 13 with hAChE. In addition, phloroglucinols 11 and 12 displayed significant anti-VRE (vancomycin-resistant Enterococci) activities, with MIC values reaching as low as 1 μg/mL.Graphical Abstract
Keywords
Rhodomyrtus tomentosa fruits Phloroglucinols hAChE inhibitory activity Anti-VRE activity Molecular docking1 Introduction
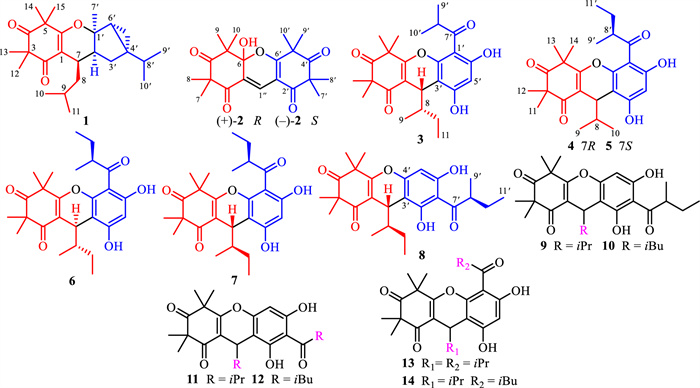
Phloroglucinols derived from species within the Myrtaceae family have garnered significant attention from scientific community due to their complex molecular architectures and notable biological activities [1]. Rhodomyrtus tomentosa, a medicinal and edible plant of this family, is extensively cultivated across Thailand, Vietnam, and southeastern China [2]. The R. tomentosa fruits are popularly used as ingredients of pies and jams in these regions [3]. Furthermore, local populations traditionally infuse the fruits into white spirits to prepare herbal liquors, which are valued for their enhanced flavor and health-promoting and disease-preventive properties. Historically, the fruits, have also been utilized for the management of diarrhea and dysentery [3]. Previous studies have demonstrated that crude extracts and phenolic constituents from R. tomentosa fruits exhibit antioxidant [4–6], anti-inflammatory [7, 8], hepatoprotective effects against nonalcoholic fatty liver disease [9], and general health-promoting benefits [10]. Despite these findings, only one phloroglucinol homodimer with anti-inflammatory activity, namely watsonianone A [11], has been identified from the fruits of title plant, leaving the broader phloroglucinol composition largely unexplored. In contrast, phloroglucinol compounds isolated from R. tomentosa leaves in tropical regions of China have been characterized for both structures and their bioactivities [12–16]. Therefore, further elucidation of novel bioactive phloroglucinol components from its fruits represents a valuable research direction. To obtain additional active phloroglucinol constituents, the fruits were subjected to ultraviolet-guided employing a DAD detector. Consequently, eight newly reported polymethylated phloroglucinols (1–8) and six known analogues (9–14) were obtained (Fig. 1). Selected compounds were examined for their hAChE inhibitory and anti-VRE activities. For the most potent hAChE inhibitor, molecular docking analysis was conducted to elucidate the possible interaction mechanism. This study presents the isolation, comprehensive structural characterization, and preliminary pharmacological assessment of the isolated metabolites.
Chemical structures of phloroglucinols 1–14
2 Results and discussion
The phloroglucinol containing moiety of R. tomentosa fruits was subjected to sequential chromatographic separation on silica gel, reversed-phase C18, Sephadex LH-20, and semi-preparative HPLC, leading to the isolation of one new polymethylated phloroglucinol meroterpenoid (PPM, 1), one new polymethylated phloroglucinol homodimer (PPHMD, 2), and six polymethylated phloroglucinol heterodimers (PPHTDs, 3–8). In addition, six previously reported congeners were structurally identified as myrciarone B (9) [17, 18], rhodotomentodimer D (10) [13], isomyrtucommulone B (11) [17, 18], rhodomyrtone (12) [19], myrtucommulone B (13) [20], and callistenone A (14) [21], based on detailed comparison of their NMR and mass spectrometry (MS) profiles with reference data previously documented in the literature.
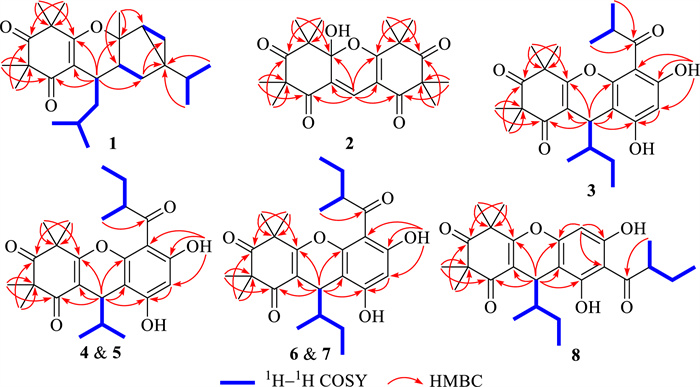
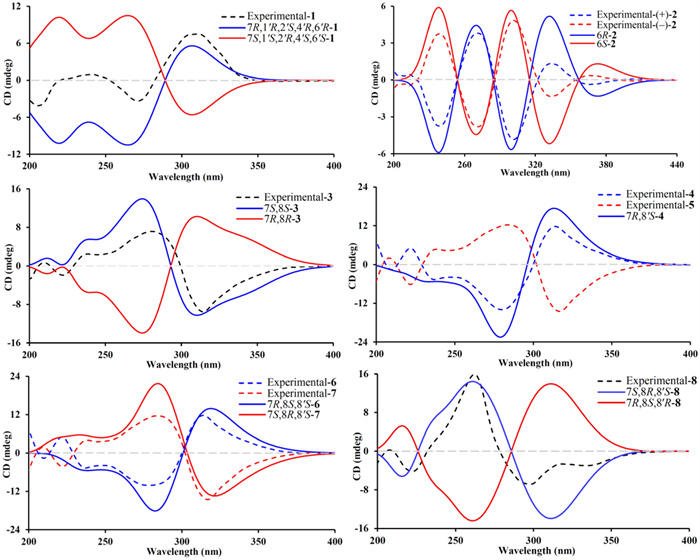
Phloroglucinol 1 was assigned the molecular formula of C25H38O3, as determined by an high-resolution electrospray ionization mass spectrometry (HRESIMS) with an [M + H]+ ion at m/z 387.2904 (calcd for C25H39O3, 387.2894), indicating seven degrees of unsaturation. The 1H NMR spectrum (Table 1) displayed signals for nine methyl groups at δH 0.85 (d, J = 6.8 Hz, H3-10'), 0.86 (d, J = 6.8 Hz, H3-9'), 0.89 (d, J = 6.6 Hz, H3-11), 0.95 (d, J = 6.6 Hz, H3-10), 1.33 (s, H3-13), 1.34 (s, H3-12), 1.38 (s, H3-15), 1.40 (s, H3-14), and 1.42 (s, H3-7′), along with two strongly shielded methylene protons at δH 0.51 (2H, brd, J = 5.4 Hz, H2-5'), characteristic of a cyclopropane moiety. Analysis of the 13C NMR data of phloroglucinol 1 (Table 1), supported by DEPT and HSQC experiments, disclosed the presence of 25 distinct carbon signals, including two keto carbonyl groups at δC 198.3 (C-2) and 213.6 (C-4), two olefinic quaternary carbons at δC 109.2 (C-1) and 168.8 (C-6), and additional four quaternary carbons, one of which was oxygenated at δC 87.3 (C-1'). The remaining signals were attributed to five methines, three methylenes, and nine methyl groups. Three of the seven degrees of unsaturation were attributed to the presence of two ketone functionalities and a single olefinic bond, while the remaining unsaturation implied the existence of a tetracyclic core structure. The planar architecture of phloroglucinol 1 was determined through comprehensive analyses of 1H–1H COSY and HMBC spectra (Fig. 2). The 1H–1H COSY experiment established an isopentyl unit of δH 2.64 (H-7)/1.29 (H2-8a) and 1.23 (H2-8b)/1.67 (H-9)/0.95 (H3-10) and 0.89 (H3-11). Key HMBC interactions, from δH 2.64 (H-7) to δC 109.2 (C-1), 168.8 (C-6), and 198.3 (C-2); from δH 1.34 (H3-12) and 1.33 (H3-13) to δC 55.5 (C-3), 98.3 (C-2) and 213.6 (C-4); and from δH 1.40 (H3-14) and 1.38 (H3-15) to δC 47.7 (C-5), 168.8 (C-6), and 213.6 (C-4), strongly supported the presence of a β-triketone core. Likewise, apart from three structural fragments of δH 1.85 (H-2')/1.34 and 1.74 (H2-3'), 0.51 (2H, H2-5')/1.36 (H-6'), and 1.29 (H-8')/0.85 (H3-9') and 0.86 (H3-10') as indicated by the 1H–1H COSY spectrum, HMBC correlations (Fig. 2), specifically from δH 1.74 (H-3'a) to δC 14.2 (C-5') and 32.4 (C-4'), from δH 1.42 (H3-7') to δC 34.9 (C-6'), 37.6 (C-2'), and 87.3 (C-1'), and from δH 0.86 (H3-9') and 0.85 (H3-10') to δC 32.4 (C-4') confirmed the presence of a thujene unit. Additional HMBC cross-peaks from δH 2.64 (H-7) to δC 34.0 (C-3') and 87.3 (C-1') indicated that the β-triketone moiety and the thujene fragment was connected through a C-6‒O‒C-1' ether linkage and a C-7‒C-2' bond. ROSEY correlations between δH 1.85 (H-2') and 0.51 (H-5'), 1.42 (H3-7'), and 2.64 (H-7) supported an α orientation of all these protons. The assignment of absolute configuration for phloroglucinol 1 was achieved by comparing its measured ECD spectrum with that simulated by TDDFT B3LYP/6–31++ G(2d, p) using Gaussian 16. The experimental curve (Fig. 4) exhibited negative Cotton effects at 205 and 271 nm and a positive signal at 311 nm, closely matching the calculated spectrum, thereby confirming the (7R,1'R,2'S,4'R,6'R) configuration. Based on combined spectroscopic and computational evidence, phloroglucinol 1, was structurally identified as a novel β-triketone‒thujene meroterpenoid and designated rhodotomentodione F.
13C (150 MHz) and 1H (600 MHz) NMR data of phloroglucinol 1 in CDCl3
Key 1H–1H COSY, and HMBC correlations of phloroglucinols 1‒8
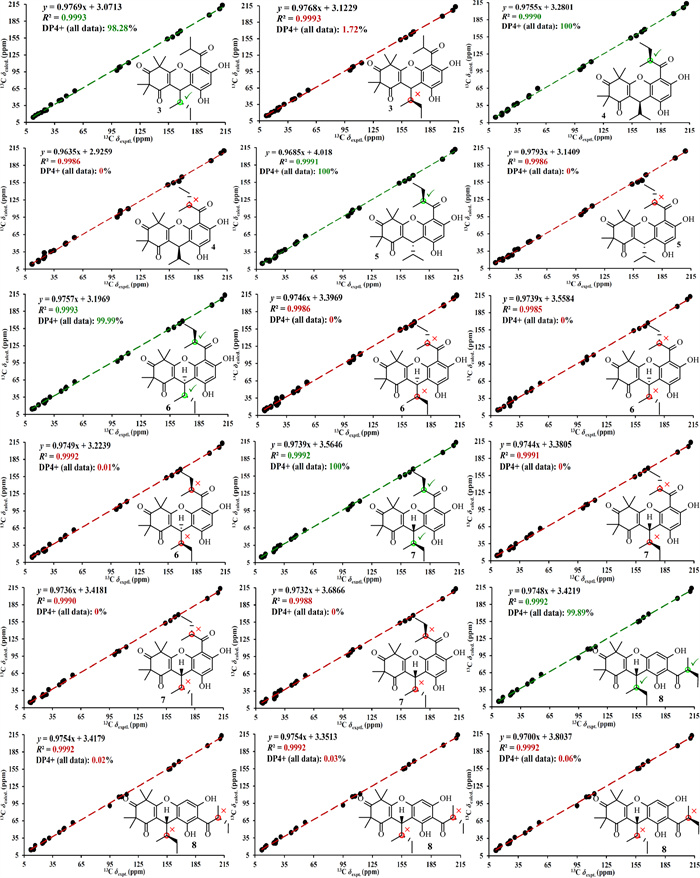
NMR calculations and DP4+ analyses for phloroglucinols 3–8
Calculated and experimental ECD spectra of phloroglucinols 1‒8
Phloroglucinol 2 was determined to possess the molecular formula C21H26O6, as indicated by HRESIMS, which displayed a [M + H]+ ion at m/z 375.1802 (cacld for C21H27O6, 375.1802). The 1H NMR spectrum of phloroglucinol 2 (Table 2) featured a downfield singlet at δH 8.00 (1H, s, H-1”) and eight singlet methyl groups at δH 1.08 (H3-9), 1.38 (H3-8), 1.40 × 2 (6H, H3-7'/H3-8'), 1.41 (H3-7), 1.50 (H3-10), and 1.53 × 2 (6H, H3-9'/H3-10'). The 13C NMR data revealed four ketone carbonyl signals at δC 194.9 (C-2'), 197.1 (C-2), 210.5 (C-4'), and 210.7 (C-4), two double bonds at δC 108.3 (C-1'), 122.1 (C-1), 127.7 (C-1”), and 172.9 (C-6'), a hemiketal carbon at δC 100.8 (C-6), four quaternary carbons at δC 48.0 (C-5'), 53.9 (C-5), 54.8 (C-3), and 56.1 (C-3'), and eight methyl groups, indicating a tricyclic scaffold. Key HMBC correlations (Fig. 2) from δH 1.41 (H3-7) and 1.38 (H3-8) to δC 54.8 (C-3), 197.1 (C-2), and 210.7 (C-4), from δH 1.50 (H3-10) and 1.08 (H3-9) to δC 53.9 (C-5), 100.8 (C-6), and 210.7 (C-4), and from δH 8.00 (H-1”) to δC 100.8 (C-6), 122.1 (C-1), and 197.1 (C-2) supported a β-triketone motif incorporating a hemiketal at C-6 and a C-1 and C-1” double bond. Similarly, HMBC cross-peaks from δH 1.40 (6H, H3-7' and H3-8') to δC 56.1 (C-3'), 194.9 (C-2'), and 210.5 (C-4'), from δH 1.53 (6H, H3-9'/ H3-10') to δC 48.0 (C-5'), 172.9 (C-6'), and 210.5 (C-4'), and from δH 8.00 (H-1”) to δC 108.3 (C-1'), 172.9 (C-6'), and 194.9 (C-2') indicated a second β-triketone fragment. Additional HMBC data indicated linkage of the two fragments via a C-6‒O‒C-6' ether bond and a C-1‒C-1”‒C-1' connection. Phloroglucinol 2, identified as a racemic mixture, was resolved into enantiomers (+)-2 and (−)-2 using a Daicel CHIRALPAK IC column. ECD spectral analysis (Fig. 4) established the absolute configurations for (+)-2 and (−)-2 to be (6R) and (6S), respectively. Thus, phloroglucinol 2 was structurally characterized as a PPHMD and designated rhodotomentodimer H.
The 13C (150 MHz) and 1H (600 MHz) NMR data of phloroglucinol 2 in CDCl3
Phloroglucinol 3 was determined to possess the molecular formula C25H32O6, as established by its HRESIMS, which exhibited a [M + H]+ ion at m/z 429.2279 (calcd for C25H33O6, 429.2272). The 1H NMR spectrum of phloroglucinol 3 (Table 3) indicated four singlet methyl groups at δH 1.39 (H3-12), 1.42 (H3-14), 1.43 (H3-13), and 1.61 (H3-15), three doublet methyl groups at δH 0.74 (J = 6.8 Hz, H3-9), 1.24 (J = 7.3 Hz, H3-10'), and 1.25 (J = 6.7 Hz, H3-9'), a triplet methyl group at δH 0.90 (J = 7.3 Hz, H3-11). An aromatic proton at δH 6.26 (s, H-5'), and a strongly deshielded hydroxyl proton at δH 13.30 (s, OH-6') were also observed. The 13C NMR spectrum (Table 3) displayed 25 carbon resonances, including eight methyl groups at δC 12.3 (CH3-11), 14.9 (CH3-9), 17.7 (CH3-9'), 20.8 (CH3-10'), 24.3 (CH3-12), 24.5 (CH3-13), 24.9 (CH3-14) and 25.1 (CH3-15), one methylene group at δC 26.1 (CH2-10), three methine groups at δC 30.9 (CH-7), 39.7 (CH-8'), and 42.1 (CH-8), two aliphatic quaternary carbons at δC 47.2 (C-5) and 56.2 (C-3), a pair of olefinic carbons at δC 112.7 (C-1) and 167.5 (C-6), six aromatic carbons at δC 100.6 (CH-5′), 103.0 (C-3′), 104.0 (C-1′), 153.8 (C-2′), 159.3 (C-4′), and 164.6 (C-6′), and three ketone carbonyl groups at δC 197.9 (C-2), 208.9 (C-7′), and 211.7 (C-4). Comparison of the abovementioned spectral data with those of myrtucommulone B (phloroglucinal 13) [20] revealed close similarity, with the exception that a 2-methylbutyl unit replaced an isobutyl at C-1 in myrtucommulone B. This structural feature was supported by a 4.35 (H-7)/1.56 (H-8)/1.50 (H2-10a) and 1.03 (H2-10b) and 0.74 (H3-9)/0.90 (H3-11) spin system identified through 1H–1H COSY analysis, along with key HMBC cross-peaks (Fig. 2) from δH 4.35 (H-7) to δC 103.0 (C-2'), 112.7 (C-1), 153.8 (C-2'), 159.3 (C-4'), 167.5 (C-6), and 197.9 (C-2). The H-7 was stochastically assigned as a β configuration, and the relative configuration for the side chain at C-7 was determined by a theoretical NMR calculation method (Fig. 3) and DP4+ probability analysis. Finally, the absolute configuration of compound 3 was established as (7S,8S) based on the ECD calculations (Fig. 4). Thus, phloroglucinol 3 was identified as a PPHTD and designated rhodotomentodimer I.
13C (150 MHz) and 1H (600 MHz) data for phloroglucinols 3–5 in CDCl3
Phloroglucinols 4 and 5 were both assigned shared the molecular formula C25H32O6, supported by HRESIMS [M + H]+ peaks at m/z 429.2275 and 429.2277 (both calcd for C25H33O6, 429.2272), respectively. Further examination of 1D NMR data (Table 3) with that of callistenone B (14) [20] revealed a substitution C-1', where the isovaleryl group in 14 was replaced by a 2-methylbutanoyl moiety in both compounds. This structural change was validated by HMBC correlation between Me-9' and C-7' and 1H–1H COSY cross-peaks linking H-8'/H2-10' and H3-9'/H3-11' (Fig. 2). The relative configurations for H-7 in compounds 4 and 5 were assigned as α and β configurations, respectively, and the relative configurations for the C-7′ side chains were determined according to further NMR calculations and DP4+ probability analysis (Fig. 3). In light of ECD calculations (Fig. 4), the absolute configuration of phloroglucinol 4 was established as (7R,8′S), and comparison of experimental ECD spectra of 5 and 4 led to the establishment of absolute configuration for 5 to be (7S,8′S). Based on spectral and computational evidence, the two structures were characterized as two PPHTDs and designated rhodotomentodimer J and rhodotomentodimer K, respectively.
Phloroglucinols 6 and 7 shared the molecular formula C26H34O6, as indicated by their HRESIMS [M + H]+ signals at m/z 443.2435 and 443.2438 (both calcd for C26H35O6, 443.2428), respectively. Their 1D NMR spectra (Table 4) were highly similar to those of phloroglucinol 3, suggesting structural similarity to PPHTD congeners. A distinguishing feature was the presence of a 2-methylbutyl moiety at C-1', evidenced by disclosed by HMBC correlations of H3-9' with C-7' and of OH-6' with C-1' as well as a COSY defined spin system of H3-9'/H-8'/H2-10'/H3-11' (Fig. 2). The relative configurations for H-7 in compounds 6 and 7 were randomly assigned as α and β configurations, respectively, and further NMR calculations and DP4+ probability analysis allowed the establishment of the relative configurations for the C-7′ side chains (Fig. 3). Owing to ECD calculations (Fig. 4), the absolute configurations of phloroglucinols 6 and 7 were established to be (7R,8S,8′S) and (7S,8R,8′S), respectively. Based on these spectroscopic and computational results, phloroglucinols 6 and 7 were identified as two two PPHTDs and designated rhodotomentodimer L and rhodotomentodimer M, respectively.
13C (150 MHz) and 1H (600 MHz) data for 6–8 in CDCl3
Phloroglucinol 8 was assigned the same molecular formula C26H34O6, identical to that of 7, as indicated by its HRESIMS peak at m/z 443.2431 [M + H]+ (calcd for C26H35O6, 443.2428). Analysis of the 13C NMR spectrum (Table 4), supported by HSQC and DEPT data, revealed 26 carbon resonances, including three keto carbonyls, an aromatic ring, four quaternary carbons (two olefinic ones), three methine groups, two methene groups, and eight methyl groups. Comparison of phloroglucinol 8 and rhodotomentodimer D (10) [13] indicated the replacement of the isopentyl side chain at C-1 by a 2-methylbutyl group. This substitution was corroborated by a 1H–1H COSY spin system (Fig. 2) of δH 4.32 (H-7)/1.60 (H-8)/1.45 (H2-10a) and 1.01 (H2-10b) and 0.71 (H3-9)/0.89 (H3-11) and HMBC correlations of δH 4.32 (H-7) with δC 104.4 (C-3'), 112.6 (C-1), 156.6 (C-4'), 162.3 (C-2'), 167.6 (C-6), and 197.6 (C-2). The relative configuration of H-7 in compound 8 was stochastically assigned to be β orientated, the β configuration for the C-8 and C-8' ethyl groups were determined by theoretical NMR calculations and DP4+ probability analysis (Fig. 3). Subsequent ECD calculations (Fig. 4) permitted the assignment of the absolute configuration for phloroglucinol 8 to be (7S,8R,8′S). Hence, phloroglucinol 8 was identified as a PPHTD and named rhodotomentodimer N.
Given that rhodotomentodimer D (10) [16], a structurally related analogue, exhibited inhibitory activity against human acetylcholinesterase (hAChE), further screening was conducted to assess this activity among related compounds (Table 5). All PPHTDs (3‒14) demonstrated significant hAChE inhibition, with IC50 values between 1.04 and 3.35 μM. In contrast, phloroglucinols 1 and 2 were inactive, with IC50 values exceeding 40.0 μM.
hAChE inhibitory activities of phloroglucinols 1–9 and 11–14
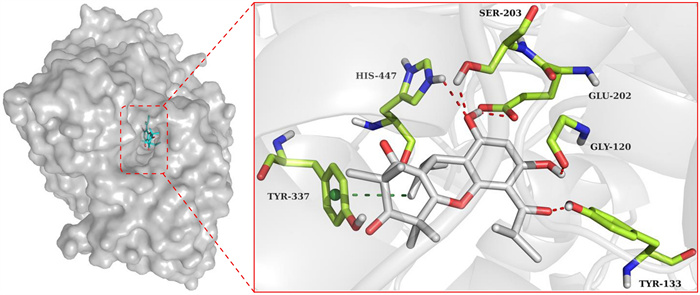
To gain insights into the binding modes of excellent hAChE inhibitor 13, molecular docking was carried out using the crystal structure of recombinant hAChE complex with (‒)-galantamine (PDB code: 4EY6, 2.40 Å resolution) [22]. The docking results indicated that phloroglucinol 13 fits well within the active site pocket of hAChE (Fig. 5). Apart from a π‒σ stacking interaction with Tyr337 (3.61 Å), five hydrogen bonds were observed between phloroglucinol 13 and the residues Gly120 (2.46 Å), Tyr133 (1.67 Å), Glu202 (2.13 Å), Ser203 (2.24 Å), and His447 (2.39 Å).
Molecular docking of phloroglucinol 13 with hAChE
In light of well-documented antibacterial activity of rhodomyrtone (12) [23–26], other phloroglucinol compounds were also assessed for their inhibitory effects on vancomycin-resistant Enterococci (VRE). Phloroglucinols 11 and 12 exhibited strong activity (MIC = 1 μg/mL), outperforming ampicillin (AMP, MIC = 2 μg/mL). Similarly, phloroglucinols 8‒10 showed comparable potency (MIC = 2 μg/mL). In contrast, phloroglucinols 1‒7, 13, and 14 were inactive (MIC > 32 μg/mL). These findings suggested that the different connection patterns of PPHTDs can significantly affect their antibacterial properties, thereby offering valuable insights for the design and synthesis of novel antibacterial agents.
3 Conclusions
A total of 14 diverse phloroglucinol derivatives (1‒14), including eight newly identified isolates (1‒8), were obtained from R. tomentosa fruits. The absolute configurations of the side chains in phloroglucinols 3‒8 were unambiguously determined through a combination of ECD computations, NMR calculations, and DP4+ probability assessments. In addition, the previously reported phloroglucinol dimers (9–14) were obtained from R. tomentosa fruits for the first time, and several of them represent characteristic and major bioactive constituents of both R. tomentosa and Myrtus communis. Notably, many of the isolated compounds demonstrated potent hAChE inhibitory activities, highlighting their potential as lead candidates for the treatment of neurological disorders, including Alzheimer's disease [27]. Furthermore, several isolates exhibited significant anti-VRE effects, underscoring the therapeutic promise of R. tomentosa fruits for combating drug-resistant bacterial infections [28, 29]. Given the limited prior knowledge of the phloroglucinol constituents of R. tomentosa fruits, the present findings significantly expand current understanding and provide a scientific basis for the future development of nutraceutical and medicinal food products.
4 Experimental procedures
4.1 General experimental procedures
The experimental procedures followed those previously reported in our recent publication [30].
4.2 Plant material
The mature fruits of R. tomentosa were collected in Wanning, Hainan province, China, in September 2023. The plant material was authenticated by Xin-Quan Yang (Hainan Branch Institute of Medicinal Plant Development, Chinese Academy of Medical Sciences). A voucher specimen (KIB-Q-202301) has been deposited at the State Key Laboratory of Phytochemistry and Natural Medicines, Kunming Institute of Botany, Chinese Academy of Sciences.
4.3 Extraction and isolation
The air-dried and powdered fruits of R. tomentosa (19.5 kg) were extracted thrice with ethanol at room temperature (48 h). The ethanol extracts were concentrated under reduced pressure, to afford a crude residue (1.56 kg), which was partitioned with petroleum ether (PE) and ethyl acetate (EtOAc), yielding PE (331 g) and EtOAc-soluble (282 g) fractions. To eliminate lipoid components, the PE extract was further partitioned with MeOH‒H2O (8:2, v/v), yielding a phloroglucinol enriched moiety (58.0 g). This enriched extract was subjected to silica gel column chromatography with a PE–EtOAc (100:1 → 1:1, v/v) gradient, yielding seven main fractions (Fr. A‒Fr. G). Fr. A (28.6 g) was further separated on silica gel (PE‒EtOAc, 100:0 → 0:100, v/v), producing three subfractions (Fr. A1–Fr. A3). Fr. A1 (6.7 g) was purified by an RP-C18 column (MeOH–H2O, 70:30 → 100:0, v/v), producing six subfractions (Fr. A1.1–Fr. A1.6). Compound 12 (4 mg) was obtained from Fr. A1.1 via recrystallization d (Acetone–MeOH, 10:1, v/v), while 10 (2.8 mg, tR = 36 min) was isolated from Fr. A1.2 (0.05 g) by semi-preparative HPLC (MeOH–H2O, 83:17, v/v). Fr. B (13 g) was separated by silica gel chromatography (PE–EtOAc, 15:1 → 0:16, v/v), resulting in seven subfractions (Fr. B1–Fr. B7). Fr. B1 (0.3 g) was further purified via semi-preparative HPLC (MeCN‒H2O, 52:48, v/v) to afford compounds 9 (3.1 mg, tR = 33 min) and 13 (10.5 mg, tR = 49 min). Fr. C (12 g) underwent RP-C18 chromatography (MeOH–H2O, 70:30 → 100:0, v/v), yielding five fractions (Fr. C1‒Fr. C5). Fr. C2 (0.18 g) was further purified by a semi-preparative HPLC (MeCN–H2O, 80:20, v/v), affording compounds 3 (1.4 mg, tR = 18 min) and 5 (3.0 mg, tR = 15 min). Fr. D (16 g) was fractionated by an RP-C18 column (MeOH–H2O, 75:25 → 100:0, v/v), resulting in six subfractions (Fr. D1‒Fr. D6). Fr. D2 (0.92 g) afforded compounds 1 (2.0 mg, tR = 35 min) and 8 (2.2 mg, tR = 40 min) by semi-preparative HPLC (MeCN–H2O, 73:27, v/v). Likewise, compound 2 (3.2 mg, tR = 42 min) was isolated from Fr. D4 (0.65 g) by a semi-prep. HPLC (MeCN–H2O, 65:35, v/v). Fr. E (2 g) was separated on Sephadex LH-20 (CHCl3–MeOH, 1:1, v/v), giving three fractions (Fr. E1‒Fr. E3). Fr. E1 (0.39 g) was subjected to semi-preparative HPLC (MeCN–H2O, 60:40, v/v), yielding compounds 4 (3 mg, tR = 32 min), 11 (7.2 mg, tR = 40 min), 6 (2.6 mg, tR = 44 min), 7 (3 mg, tR = 42 min), and 14 (7.0 mg, tR = 37 min). Chiral separation of compound 2 was achieved using a Daicel CHIRALPAK IC column (n-hexane–isopropanol, 95:5 → 85:15, v/v), furnishing enantiomers (+)-2 (1.0 mg, tR = 12 min) and (−)-2 (1.0 mg, tR = 13 min).
Rhodotomentodione F (1): Colorless gum; [α] + 28.1 (c 0.1, MeOH); UV (MeOH) λmax (log ε) 268 (3.68) nm; ECD (MeOH) λmax (Δε) 205(–5.36), 241 (+ 1.25), 271 (–4.41), 311 (+ 9.76) nm; 1H (600 MHz, CDCl3) and 13C (150 MHz, CDCl3) NMR data, see Table 1; (+)-HRESIMS m/z 387.2904 [M + H]+ (calcd for C25H38O3, 387.2894).
Rhodotomentodimer H (2): Colorless gum; [α] + 49.0 (c 0.10, MeOH) for (+)-2; [α] –49.0 (c 0.10, MeOH) for (–)-2; UV (MeOH) λmax (log ε) 217 (2.68), 272 (2.52) nm; ECD (MeOH) λmax (Δε) 239 (‒4.49), 271 (+4.56), 301 (−5.83), 333 (+ 1.61) nm for (+)-2; ECD (MeOH) λmax (Δε) 239 (+4.49), 271 (−4.56), 301 (+ 5.83), 333 (−1.61) nm for (–)-2; 1H (600 MHz, CDCl3) and 13C (150 MHz, CDCl3) NMR data, see Table 2; (+)-HRESIMS m/z 375.1802 [M + H]+ (calcd for C21H27O6, 375.1802).
Rhodotomentodimer I (3): Colorless gum; [α] − 85.4 (c 0.10, MeOH); UV (MeOH) λmax (log ε) 203 (4.08), 216 (4.06), 295 (3.87) nm; ECD (MeOH) λmax (Δε) 209 (+0.34), 222 (‒1.01), 238 (+1.26), 280 (+3.70), 314 (‒4.94) nm; 1H (600 MHz, CDCl3) and 13C (150 MHz, CDCl3) NMR data, see Table 3; (+)-HRESIMS m/z 429.2279 [M + H]+ (calcd for C25H33O6, 429.2272).
Rhodotomentodimer J (4): Colorless gum; [α] + 149.4 (c 0.10, MeOH); UV (MeOH) λmax (log ε) 203 (4.27), 217 (4.26), 295 (4.10) nm; ECD (MeOH) λmax (Δε) 209 (‒1.19), 221 (+3.65), 237 (−3.38), 280 (−10.08), 313 (+8.51) nm; 1H (600 MHz, CDCl3) and 13C (150 MHz, CDCl3) NMR data, see Table 3; (+)-HRESIMS m/z 429.2275 [M + H]+ (calcd for C25H33O6, 429.2272).
Rhodotomentodimer K (5): Colorless gum; [α] –186.6 (c 0.10, MeOH); UV (MeOH) λmax (log ε) 203 (4.26), 216 (4.27), 294 (4.16) nm; ECD (MeOH) λmax (Δε) 208 (+ 1.49), 221 (‒4.50), 238 (‒3.40), 283 (+ 8.83), 316 (‒10.50) nm; 1H (600 MHz, CDCl3) and 13C (150 MHz, CDCl3) NMR data, see Table 3; (+)-HRESIMS m/z 429.2277 [M + H]+ (calcd for C25H33O6, 429.2272).
Rhodotomentodimer L (6): Colorless gum; [α] + 83.7 (c 0.10, MeOH); UV (MeOH) λmax (log ε) 203 (4.10), 217 (4.10), 295 (3.98) nm; ECD (MeOH) λmax (Δε) 209 (‒0.79), 221 (+ 2.42), 237 (‒2.24), 278 (‒4.85), 313 (+ 5.65) nm; 1H (600 MHz, CDCl3) and 13C (150 MHz, CDCl3) NMR data, see Table 4; (+)-HRESIMS m/z 443.2435 [M + H]+ (calcd for C26H35O6, 443.2428).
Rhodotomentodimer M (7): Colorless gum; [α] ‒85.4 (c 0.10, MeOH); UV (MeOH) λmax (log ε) 205 (4.25), 218 (4.27), 297 (4.15) nm; ECD (MeOH) λmax (Δε) 209 (+ 1.22), 222 (–2.88), 238 (+ 2.69), 283 (+ 7.78), 316 (–9.79) nm; 1H (600 MHz, CDCl3) and 13C (150 MHz, CDCl3) NMR data, see Table 4; (+)-HRESIMS m/z 443.2438 [M + H]+ (calcd for C26H35O6, 443.2428).
Rhodotomentodimer N (8): Colorless gum; [α] ‒60.2 (c 0.10, MeOH); UV (MeOH) λmax (log ε) 202 (4.02), 225 (3.99), 263 (3.73), 301 (3.88) nm; ECD (MeOH) λmax (Δε) 207 (+ 0.20), 222 (‒2.94), 262 (+ 11.13), 297 (‒4.81) nm; 1H (600 MHz, CDCl3) and 13C (150 MHz, CDCl3) NMR data, see Table 4; (+)-HRESIMS m/z 443.2431 [M + H]+ (calcd for C26H35O6, 443.2428).
4.4 ECD and NMR calculations
ECD and NMR computations were carried out based on previously reported methods using the Gaussian 16 software package [14]. Three-dimensional molecular structures were constructed based on ROESY spectra (for compound 1) or randomly generated (for compounds 2–8), followed by conformational analysis using CONFLEX with the MMFF94S force field. ECD and NMR chemical shift calculations were conducted at the B3LYP/6–311++ G(2d, p) and mPW1PW91/6–31+ G(d, p) levels, respectively. The DP4+ probability analyses were accomplished utilizing a published computational template [31].
4.5 Human AChE inhibitory assay
hAChE inhibition assays were performed according to a reported protocol, with galantamine as the positive control [32]. All assays were performed in triplicate. Compounds with IC50 values exceeding 40 μM were considered inactive.
4.6 Anti-VRE assay
As described in earlier studies, the antibacterial activities against vancomycin-resistant Enterococci (VRE) were achieved by a standard broth microdilution method in 96-well plates [33]. Ampicillin (AMP) was a positive control, and all tests were performed three times.
4.7 Molecular docking
Molecular docking simulations were performed utilizing the AutoDock software suit [34], following established protocols. The docking complexes of phloroglucinol 13 with hAChE (PDB ID: 4EY6) were generated, and the resulting binding poses were visualized utilizing the PyMOL Molecular Graphics System.
Notes
Acknowledgements
This study was financially supported by the National Natural Science Foundation of China (32270425 and 31970377), the Yunnan Fundamental Research Projects (202501AS070093, 202201AT070181, and 202401AT070191), the Top Young Talent of Ten Thousand Talents Program of Yunnan Province (YNWR-QNBJ-2020-253), the Reserve Talents of Young and Middle-Aged Academic and Technical Leaders of Yunnan Province (202105AC160023), and CAS Light of West China Program. We are grateful to Analysis and Testing Centre (Kunming Institute of Botany, Chinese Academy of Sciences) for NMR, HRESIMS, CD, and bioactive measurement.
Author contributions
Ling-Yun Chen carried out the experiments and prepared the original draft; Mu-Yuan Yu participated in the main experiments and revised the manuscript; E-E Luo and Wen-Ying Zong performed the ECD calculations; Shu-Mei Lei and Yu Pan performed the NMR calculations; Ai-Chun Lu and Cheng-Qin Liang revised the manuscript; Xu-Jie Qin conceived and supervised the study, revised the manuscript, and secured project funding. All authors reviewed and approved the final version of the manuscript.
Data availability
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
Declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
-
1.Celaj O, Duarán AG, Cennamo P, Scognamiglio M, Fiorentino A, Esposito A, et al. Phloroglucinols from Myrtaceae: attractive targets for structural characterization, biological properties and synthetic procedures. Phytochem Rev 2021;20(1): 259-99. CrossRef PubMed Google Scholar
-
2.Saising J, Ongsakul M, Voravuthikunchai SP. Rhodomyrtus tomentosa (Aiton) Hassk. ethanol extract and rhodomyrtone: a potential strategy for the treatment of biofilm-forming staphylococci. J Med Microbiol 2011;60(12): 1793-800. CrossRef PubMed Google Scholar
-
3.Hu X, Chen Y, Dai J, Yao L, Wang L. Rhodomyrtus tomentosa fruits in two ripening stages: chemical compositions, antioxidant capacity and digestive enzymes inhibitory activity. Antioxidants 2022;11(7): 1390. CrossRef PubMed Google Scholar
-
4.Cui C, Zhang S, You L, Ren J, Luo W, Chen W, et al. Antioxidant capacity of anthocyanins from Rhodomyrtus tomentosa (Ait.) and identification of the major anthocyanins. Food Chem 2013;139(1–4): 1-8. CrossRef PubMed Google Scholar
-
5.Lai TNH, André C, Rogez H, Mignolet E, Nguyen TBT, Larondelle Y. Nutritional composition and antioxidant properties of the sim fruit (Rhodomyrtus tomentosa). Food Chem 2015;168: 410-6. CrossRef PubMed Google Scholar
-
6.Wu P, Ma G, Li N, Deng Q, Yin Y, Huang R. Investigation of in vitro and in vivo antioxidant activities of flavonoids rich extract from the berries of Rhodomyrtus tomentosa (Ait.) Hassk. Food Chem 2015;173: 194-202. CrossRef PubMed Google Scholar
-
7.Shiratake S, Nakahara T, Iwahashi H, Onodera T, Mizushina Y. Rose myrtle (Rhodomyrtus tomentosa) extract and its component, piceatannol, enhance the activity of DNA polymerase and suppress the inflammatory response elicited by UVB-induced DNA damage in skin cells. Mol Med Rep 2015;12(4): 5857-64. CrossRef PubMed Google Scholar
-
8.Wang R, Yao L, Lin X, Hu X, Wang L. Exploring the potential mechanism of Rhodomyrtus tomentosa (Ait.) Hassk fruit phenolic rich extract on ameliorating nonalcoholic fatty liver disease by integration of transcriptomics and metabolomics profiling. Food Res Int 2022;151: 110824. CrossRef PubMed Google Scholar
-
9.Wang R, Yao L, Meng T, Li C, Wang L. Rhodomyrtus tomentosa (Ait.) Hassk fruit phenolic-rich extract mitigates intestinal barrier dysfunction and inflammation in mice. Food Chem 2022;393: 133438. CrossRef PubMed Google Scholar
-
10.Lai TNH, Herent MF, Leclercq JQ, Nguyen TBT, Rogez H, Larondelle Y, et al. Piceatannol, a potent bioactive stilbene, as major phenolic component in Rhodomyrtus tomentosa. Food Chem 2013;138(2–3): 1421-30. CrossRef PubMed Google Scholar
-
11.Zhuang L, Chen LF, Zhang YB, Liu Z, Xiao XH, Tang W, et al. Watsonianone a from Rhodomyrtus tomentosa fruit attenuates respiratory-syncytial-virus-induced inflammation in vitro. J Agric Food Chem 2017;65(17): 3481-9. CrossRef PubMed Google Scholar
-
12.Chen LY, Luo EE, Pan Y, Liang CQ, Yu MY, Qin XJ. Acetylcholinesterase inhibitory phloroglucinols from tropic Rhodomyrtus tomentosa. Phytochemistry 2024;228: 114254. CrossRef PubMed Google Scholar
-
13.Luo EE, Liu SN, Wang ZJ, Chen LY, Liang CQ, Yu MY, et al. Oligomeric phloroglucinols with hAChE inhibitory and antibacterial activities from tropic Rhodomyrtus tomentosa. Bioorg Chem 2023;141: 106836. CrossRef PubMed Google Scholar
-
14.Qin XJ, Liu H, Li PP, Ni W, He L, Khan A, et al. Polymethylated acylphloroglucinols from Rhodomyrtus tomentosa exert acetylcholinesterase inhibitory effects. Bioorg Chem 2021;107: 104519. CrossRef PubMed Google Scholar
-
15.Qin XJ, Rauwolf TJ, Li PP, Liu H, McNeely J, Hua Y, et al. Isolation and synthesis of novel meroterpenoids from Rhodomyrtus tomentosa: Investigation of a reactive enetrione intermediate. Angew Chem Int Ed 2019;58(13): 4291-6. CrossRef PubMed Google Scholar
-
16.Yu MY, Liu SN, Luo EE, Jin Q, Liu H, Liu HY, et al. Phloroglucinols with hAChE and α-glucosidase inhibitory activities from the leaves of tropic Rhodomyrtus tomentosa. Phytochemistry 2022;203: 113394. CrossRef PubMed Google Scholar
-
17.Kaneshima T, Myoda T, Toeda K, Fujimori T, Nishizawa M. Antimicrobial constituents of peel and seeds of camu-camu (Myrciaria dubia). Biosci Biotechnol Biochem 2017;81(8): 1461-5. CrossRef PubMed Google Scholar
-
18.Tanaka N, Jia Y, Niwa K, Imabayashi K, Tatano Y, Yagi H, et al. Phloroglucinol derivatives and a chromone glucoside from the leaves of Myrtus communis. Tetrahedron 2018;74(1): 17-123. CrossRef PubMed Google Scholar
-
19.Salni D, Sargent MV, Skelton BW, Soediro I, Sutisna M, White AH, et al. Rhodomyrtone, an antibiotic from Rhodomyrtus tomentosa. Aust J Chem 2002;55(3): 229-32. CrossRef PubMed Google Scholar
-
20.Charpentier M, Hans M, Jauch J. Enantioselective synthesis of myrtucommulone A. Eur J Org Chem 2013;2013(19): 4078-84. CrossRef PubMed Google Scholar
-
21.Rattanaburi S, Mahabusarakam W, Phongpaichit S, Carroll AR. Acylphloroglucinols from Callistemon lanceolatus DC. Tetrahedron 2013;69(30): 6070-5. CrossRef PubMed Google Scholar
-
22.Cheung J, Rudolph MJ, Burshteyn F, Cassidy MS, Gary EN, Love J, et al. Structures of human acetylcholinesterase in complex with pharmacologically important ligands. J Med Chem 2012;55(22): 10282-6. CrossRef PubMed Google Scholar
-
23.Leejae S, Taylor PW, Voravuthikunchai P. Antibacterial mechanisms of rhodomyrtone against important hospital-acquired antibiotic-resistant pathogenic bacteria. J Med Microbiol 2013;62(1): 78-85. CrossRef PubMed Google Scholar
-
24.Limsuwan S, Hesseling-Meinders A, Voravuthikunchai SP, Dijl JM, Kayser O. Potential antibiotic and anti-infective effects of rhodomyrtone from Rhodomyrtus tomentosa (Aiton) Hassk. on Streptococcus pyogenes as revealed by proteomics. Phytomedicine 2011;18(11): 934-40. CrossRef PubMed Google Scholar
-
25.Rani R, Righetto GM, Schäfer AB, Wenzel M. The diverse activities and mechanisms of the acylphloroglucinol antibiotic rhodomyrtone: antibacterial activity and beyond. Antibiotics 2024;13(10): 936. CrossRef PubMed Google Scholar
-
26.Srisuwan S, Mackin KE, Hocking D, Lyras D, Bennett-Wood V, Voravuthikunchai SP, et al. Antibacterial activity of rhodomyrtone on Clostridium difficile vegetative cells and spores in vitro. Int J Antimicrob Agents 2018;52(5): 724-9. CrossRef PubMed Google Scholar
-
27.Singh M, Kaur M, Kukreja H, Chugh R, Silakari O, Singh D. Acetylcholinesterase inhibitors as Alzheimer therapy: from nerve toxins to neuroprotection. Eur J Med Chem 2013;70: 165-88. CrossRef PubMed Google Scholar
-
28.Kumar G, Engle K. Natural products acting against S. aureus through membrane and cell wall disruption. Nat Prod Rep 2023;40(10): 1608-46. PubMed Google Scholar
-
29.Wang ZJ, Zhu YY, Bai LY, Tang DM, Zhou ZS, Wei MZ, et al. A new therapeutic strategy for infectious diseases against intracellular multidrug-resistant bacteria. J Control Release 2024;375: 467-77. CrossRef PubMed Google Scholar
-
30.Luo EE, Yang WF, Wang ZJ, Chen LY, Yu MY, Luo XD, et al. Phytocannabinoid-like meroterpenoids from twigs and leaves of Rhododendron spinuliferum. Phytochemistry 2024;228: 114241. CrossRef PubMed Google Scholar
-
31.Grimblat N, Zanardi MM, Sarotti AM. Beyond DP4: an improved probability for the stereochemical assignment of isomeric compounds using quantum chemical calculations of NMR shifts. J Org Chem 2015;80: 12526-34. CrossRef PubMed Google Scholar
-
32.Liu H, He XZ, Feng MY, Zeng Y, Rauwolf TJ, Shao LD, et al. Acylphloroglucinols with acetylcholinesterase inhibitory effects from the fruits of Eucalyptus robusta. Bioorg Chem 2020;103: 1104127. CrossRef PubMed Google Scholar
-
33.Shi YZ, Wang ZJ, Shi N, Bai LY, Jiang YM, Jiang L, et al. Anti-MRSA mechanism of spirostane saponin in Rohdea pachynema F.T. Wang & tang. J Ethnopharmacol 2024;331: 118327. CrossRef PubMed Google Scholar
-
34.Yu MY, Liu SN, Liu H, Meng QH, Qin XJ, Liu HY. Acylphloroglucinol trimers from Callistemon salignus seeds: isolation, configurational assignment, hAChE inhibitory effects, and molecular docking studies. Bioorg Chem 2021;117: 105404. CrossRef PubMed Google Scholar
Copyright information
© The Author(s) 2026.
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.