Advances in natural product discovery: strategies, technologies, and insights
Abstract
Natural products (NPs) and their analogues have long underpinned therapies in humans, animals, and plants health, yet, discovering truly novel scaffolds remains a formidable challenge, even with the enormous diversity offered. Over the last two decades, breakthroughs in bioinformatics, cheminformatics, advanced analytical methods, synthetic biology toolkits, and optimized microbial culture have surmounted many of the bottlenecks that stalled NP research in the 1990s and 2000s. Researchers now deploy innovative extraction and purification protocols alongside high-throughput dereplication tools to fish trace metabolites out of complex matrices. These combined approaches not only enable the discovery and rigorous characterization of biosynthesized metabolites, bio-transformed analogues and new chemical entities but also allow precise tuning of biosynthetic gene clusters (BGCs) and culture conditions- modulation and optimization, dramatically improving yield, scalability, and cost-efficiency. Several of these newly unearthed compounds exhibit unique bioactivities that directly inspire drug-development programs against metabolic disorders, cancer drug resistance, and infectious diseases. In this review, we present an up-to-date, concise roadmap of natural product discovery (NPD), majorly covering strategies for awakening silent BGCs, genome mining, and late-stage diversification systems, and we discuss the current limitations and perspectives of rational NPD.Graphical Abstract
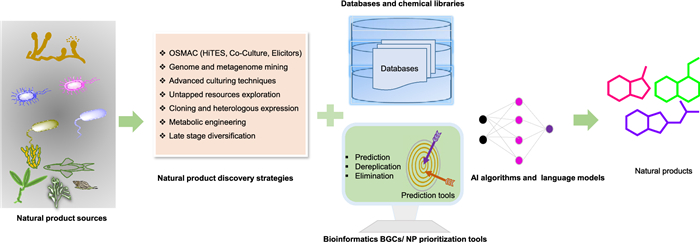
Keywords
Natural products Culturing modulation Unexplored reservoirs Genome mining Natural product diversification1 Introduction
Natural products (NPs) are small organic molecules such as peptides, polyketides, saccharides, terpenes, and alkaloids, produced by plants, microbes, invertebrates and animals for self-defense or as metabolic byproducts. While non-essential for growth, NPs play a crucial role in chemical ecology, from defense against predators and competitors to sensing environmental cues like light [1–3]. Their biosynthesis proceeds via enzyme cascades and precursors from nutrient sources or primary metabolic pools, especially amino acids and tricarboxylic acid cycle intermediates, to assemble structurally diverse compounds [2, 4].
NPs and their semi-synthetic analogs form a rich reservoir of pharmacologically potent compounds whose structural diversity underlies a broad spectrum of bioactivities, from antimicrobial and antiparasitic to anticancer effects. This chemical versatility has driven drug discovery for millennia, delivering new therapeutic leads long before modern screening platforms existed. That legacy endures today with over half of the US Food and Drug Administration-approved drugs from 1939 to 2019 derived from NPs or their derivatives [5].
Tracing back to history, "morphine" from Principium somniferum and the semi-synthetic drug "aspirin" based on salicin, an NP from Salix, being the first proof as a pure and NPs derivative practiced for human disease treatment [6, 7] Over the eons, classical trial-and-error top-down approaches, such as structure-, bioactivity, and affinity- guided bioactive molecules isolation [8], contributed to significant early milestones in drug approval. For instance, 70–80% of antibiotics discovered were directly NPs based or inspired by NPs entities. The period from 1940 to 1960s is often regarded as the "golden age" in natural products drug discovery history [9, 10]. During this time, intense research into microbial sources, especially soil-dwelling bacteria like Streptomyces, led to the rapid identification of numerous antibiotic classes [11], antipsychotic agents [12], and anticancer agents [13] that are still in clinical use today.
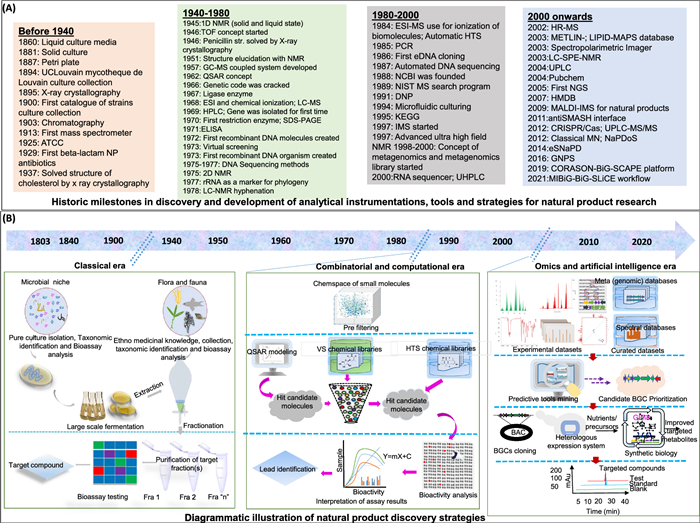
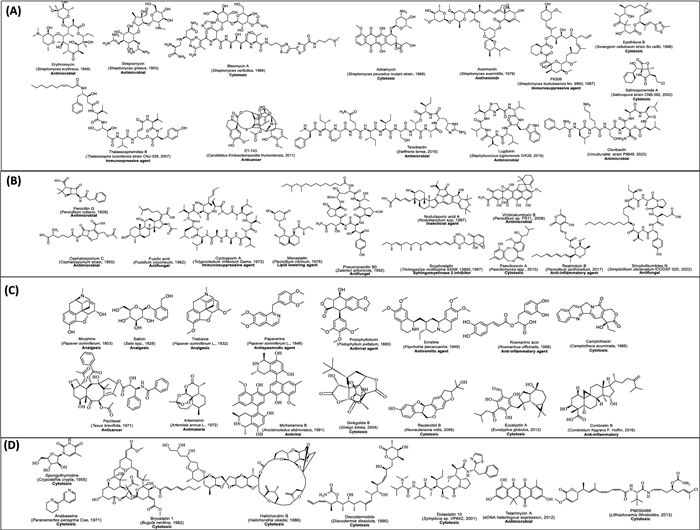
Figure 1 illustrates the chronological progression of essential tools, methodologies, and landmark discoveries that have propelled natural product discovery (NPD) from 1800 to 2024. It highlights major technological milestones, such as instruments, databases, and dereplication tools, alongside key isolation strategies exploited maximum during that period. Figure 2 displays notable NPs discovered across this timeline, emphasizing their origins from microbial and plant sources and detailing their biological activities.
Progress time trend in hallmark tools and strategies workflow for accelerating natural product discovery (NPD). A Significant discoveries time of instruments, databases and dereplication tools in NPD research. B Time trend in isolation techniques majorly, ancient to late-twentieth century period (Top-Down Approach), late 20th to early 21th century period (Virtual Screening, High Throughput Screening and Combinatorial-/biosynthesis) and modern early 21th century to now (Omics, Artificial Intelligence and Bottom-Up Approach)
Representative revolutionary natural product discovered during the 1800–2023, usually in every decade. A Natural products (NPs) isolated from bacterial source, B NPs isolated from fungal source, C NPs isolated from plant source and D NPs from miscellaneous sources. Each metabolite is listed with its name, source of first-time isolation (provided in parentheses), and biological activity (highlighted in bold)
Ethno medicine, phenotypic screening and bioactivity-guided trial-and-error methods have traditionally served as widely adopted protocols for NPs identification and discovery in the classical era. However, these trial-and-error methods constraints such as poor cultivability under standard culture settings, high rediscovery rates, labor and time-intensive, and unnecessary financial burden diminished the natural products scientists stake and pharmaceutical industries on NPs research [14]. In this scenario, high throughput screening (HTS) and combinatorial synthesis intensively exploited between the 1980s to early 2000s improved the lead discovery by increasing the hit rate to approximately 10–40% [15], but these systems have their own set of restrictions, such as limited chemical diversity and screening library chemspace size [16]. Therefore, they failed to meet continuous demand for new chemical entities (NCEs), new scaffolds and drugs as highlighted by a survey reporting only two Food and Drug Administration-approved combination drugs over a 39-year period [5].
Amid this backdrop, cutting-edge "bottom-up approaches" including multi-omics technologies, hyphenated analytical techniques, and bio-/cheminformatics platforms, offer exciting avenues for accelerating the novel carbon skeletons or NCEs discovery. Coupled with advanced mass spectrometry (MS)/ nuclear magnetic resonance (NMR)-based dereplication methodologies and databases (Table 1), which swiftly filter known compounds and eliminate redundancy, these innovations enable precise targeted exploration of unique metabolites [17]. This era is often regarded as a "new golden period" or "muti-omics AI era" for NPs-based drug discovery and development [18]. Concomitantly, the rapid surge of NPs, with approximately 6–7 million standardized and centralized in public databases such as Dictionary of Natural Products, COlleCtion of Open Natural prodUcTs, Natural Product Atlas (Table 1A), accounts for only 1/10th of the total NPs chemspace [2]. Yet, specialized metabolites produced in low concentrations (often less than 1% by weight) in complicated cellular compartments, a time-consuming dereplication process, silent or incomplete biosynthetic domains nature of biosynthetic gene clusters (BGCs) remain major challenges [19].
Databases and mining tools in natural product discovery
Additionally, modern bioinformatics analysis, metagenomics and next-generation sequencing illuminated that the number of BGCs is considerably underappreciated compared to expected [76], highlighting the tremendous potential for NCEs from untapped biodiversity [77]. Further, profound improvement in mass spectrometry studies disclosed that the number of secreted NPs far surpasses the number of BGCs, which paved the way for a logic for myriad unconventional NPs. Given the limitations of classical approaches, recent advancements in metabologenomics, next-generation synthetic biology, and cutting-edge technologies such as artificial intelligence (AI), machine learning (ML), and large language models have significantly improved both the hit rate and the yield of structurally diverse carbon skeletons and their analogues. Moreover, emphasis has been placed on combinatorial approaches [37], particularly with the expansion of virtual screening libraries to ultra-large scales [78] and the implementation of platforms such as VirtualFlow [79]. These strategies are often coupled with ultrasensitive automation, substantial miniaturization, or whole-cell phenotypic high-throughput screening (HTS) techniques [80], leveraging massive datasets like the L1000 [81] to enhance screening efficiency. Nevertheless, robust high-throughput methodologies for the comprehensive detection, isolation, and characterization of all encoded natural products and their full chemical diversity from complex extracts remain elusive.
NPs offer tremendous promise for developing novel therapeutics and advancing sustainability in food and agriculture. However, despite their vast potential, technical, biological, and regulatory hurdles continue to constrain their discovery and translation into practical applications. NP discovery generally begins with metabolite isolation via chromatography and bioassay-guided fractionation, followed by structural elucidation using advanced spectroscopic methods such as MS, NMR, chemical derivatization, and X-ray crystallography. Although many NPs exhibit potent bioactivity, issues like poor solubility, chemical instability, and toxicity can impede pharmacological development. Conversely, their ability to interact with multiple biological targets opens opportunities for multitarget therapies while also raising concerns about off-target effects. Early dereplication helps identify known compounds and reduce redundancy, yet it underscores the difficulty of finding truly novel entities. To overcome these challenges, advances in analytical instrumentation and informatics, especially AI-driven platforms and deep-learning tools in bioinformatics and cheminformatics, combined with next-generation synthetic biology now enable high-confidence prediction, reconstruction, and expression of BGCs and metabolic pathways. Emerging and updated tools and strategies such as antiSMASH, Global Natural Products Social Molecular Networking (GNPS), High Throughput Elicitors Screening (HiTES), resistance/phylogeny-guided genome mining, transcriptional/translational modulation, and late-stage modification are becoming indispensable components of the modern NP discovery pipeline.
2 Important pillars of natural product discovery: osmac, co-culture, and elicitors
Classical methods like bioassay-guided fractionation, top-down approaches, and structure elucidation via NMR, X-ray, and MS, alongside culture optimization techniques like one strain many compounds (OSMAC) [82] and HiTES [83], continue to thrive in unveiling NCEs maintaining its effectiveness even amid the remarkable success of modern discovery approaches.
2.1 OSMAC
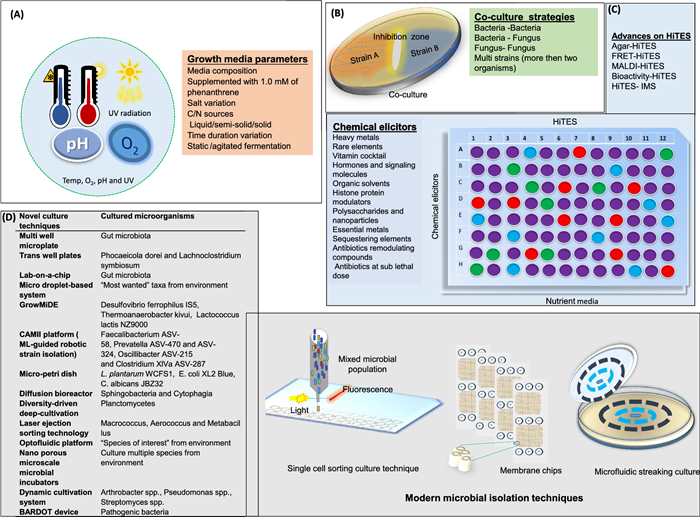
OSMAC is a modified culturing technique widely used in NP research to expand chemical diversity and trigger cryptic gene expression for cryptic specialized metabolites by systemic altering environmental and chemical triggers like media composition, carbon sources, and fermentation methods and listed in Fig. 3A. Despite its low-tech simplicity, this approach has proven remarkably powerful across diverse sources, including marine resources [84], symbiotic fungi [85], lichen [86], and Streptomyces [87] to discover plethora of novel NPs. For example, recent comprehensive reviews by Zhang et al. [88] and Zhu and Zhang [89] reported the discovery of over 284 microbial cyclic peptides from 63 endophytic strains, and 476 secondary metabolites from fungal strains, respectively, through the application of the OSMAC strategy, illuminating the strategy's huge potential for NPs discovery.
Culture modulating and untapping the unexplored reservoirs techniques. A Physical and chemical methods B Co-culture strategies. C HiTES experiment. D Examples of modern microbial cultivation techniques
One major limitation of this approach is the prioritization of compounds within a complex crude extract, which contains a vast chemical space. This complexity poses significant challenges for subsequent steps such as compound isolation and purification. In recent years, efforts to simplify the dereplication process have involved OSMAC-modified strategies, including integration with heterologous expression systems [90], molecular networking [91], genome mining [92], metabolic shunting [93] to speculate optimum conditions for discovering novel NPs, or analogues. For instance, Esposito et al. [92] isolated more than 30 new glycolipids with unusual functional groups from the marine bacterium Rhodococcus sp. I2R by combining an OSMAC approach with genome mining and advanced metabolomics analysis. Similarly, Wu et al. [94] applied genome mining alongside an OSMAC strategy to uncover 12 new alkaloids from termite-associated Streptomyces tanashiensis BYF-112.
2.2 Co-culture
Co-culture techniques leverage interactions between multiple microorganisms, either on solid or in liquid media, to enhance NPD. By mimicking ecological niches and chemical crosstalk, co-culture can activate silent biosynthetic gene clusters or trigger the scalable production of specialized metabolites via poorly understood signals [95], except with little evidence of the mimicry of ecological culture niches and interspecies/intraspecies crosstalk [96]. Unlike monoculture, which often silences key BGCs and requires complex genetic or bioprocess interventions, co-culture offers a simple promising alternative for activating and enhancing the yield of diverse NPs [97]. For example, Peng et al. [98] and Li et al. [97] reported the discovery of 93 novel bioactive natural products from co-cultured microorganisms (2017–2020) and 69 metabolites from various co-culture techniques, respectively.
Figure 3B illustrate possible combination of co-partner in co-culture strategies. Moreover, basic setups include mixed growth, spatial separation via layered or immobilized arrangements, and encapsulation in shared media to limit dominance. In other designs, partners are placed in distinct chambers that exchange metabolites through semi-permeable membranes or volatile signals across a gas interface. Alternatively, one strain spent media or extract can be used to stimulate the metabolic activity of the other [97].
Recent technological advances have further empowered co-culture. Biosensor-assisted cell selection strategy [99], optogenetics circuit system [100], microbe-laden hydrogel system [101] and metabolic flux analysis [101] allow real-time monitoring and dynamic control of species interactions. For instance, Guo et al. [99] engineered two separate Escherichia coli–Escherichia coli co-culture systems, one channeling 4-hydroxybenzoate into phenol, the other tyrosine, each equipped with a phenol biosensor incorporating a biosensor to monitor phenol production, which resulted in 2.3- and 3.9-fold increases in phenol titers, respectively.
Furthermore, integration with advanced analytics, for instance, combined with imaging mass spectrometry techniques such as MALDI-TOF–MS and Nano-DESI-IMS [102], as well as high-throughput elicitor screens (HiTES) [103], have significantly uncovered target NPs and their derivatives. In one study, coupling a bioassay with HiTES in a co-culture system led by Moon et al. [103] to isolate the novel lanthipeptide antibiotic cebulantin.
More recently, a new approach termed "modular co-culture engineering" has been introduced to address the shortcomings of monoculture fermentation and improve the enhance NPs biosynthesis. In this strategy, a complex biosynthetic pathway into distinct functional modules, each assigned to a different microbial strain. Each strain is independently engineered and optimized prior to co-culturing (either together in shared media or within compartmentalized systems), to produce the target compound. By balancing population ratios and monitoring metabolite exchange, this approach reduces metabolic stress on individual microbes and improves overall biosynthetic efficiency [104, 105]. For example, applying this strategy, Marsafari et al. [106] co-cultured engineered Yarrowia lipolytica Po1f and Po1g strains, and reported an increase of amorphadiene titer of 60–70 mg/mL, compared to 40 mg/mL in monoculture.
2.3 HiTES
HiTES is an innovative technique designed to unlock the hidden biosynthetic potential of microorganisms by activating silent or cryptic BGCs with libraries of small-molecule elicitors. Unlike traditional genetic manipulation methods, HiTES operates without the need for cloning or genome editing, making it a rapid and versatile platform [83] (Fig. 3C). However, no established methodology currently exists for identifying small-molecule elicitors that selectively activate a specific silent BGC. This shortfall has slowed efforts to unravel the complex regulatory networks controlling secondary metabolite expression and left many potentially valuable natural products dormant. Reporter constructs offer a partial solution by linking induction of a target BGC to an easily measurable signal. In one early demonstration, inserted a reporter into multiple gene clusters and screened small-molecule libraries, identifying sub lethal trimethoprim as a global inducer of at least five BGCs [107]. Building on this, Xu et al. (2017) engineered an eGFP reporter into the sur BGC, which led to the characterization of 14 novel surugamide-family metabolites following elicitor treatment [108].
Another major challenge in HiTES workflows is managing and interpreting the exceptionally complex LC–MS datasets that each elicitor screen produces. Automated dereplication against spectral libraries (e.g. GNPS) [109] and in-silico fragmentation tools (e.g. SIRIUS/CSI: FingerID) [110] help to annotate known compounds, but unknowns remain pervasive. GNPS, conceptually initiated in 2011 and introduced in 2014, is an open-access, web-based platform that facilitates the community-driven sharing and analysis of MS/MS data. It supports a variety of advanced workflows that enhance analytical resolution and throughput by integrating complementary modalities [111]. It have been widely applied across diverse sources including plants, microorganisms, and extremophiles for the discovery of all classes of NPs [112, 113]. GNPS advances include feature-based molecular networking (FBMN) which improves molecular comparisons by incorporating fragmentation spectra, isotope patterns, retention times, and ion mobility data [114]. Ion Identity Molecular Networking extends FBMN by linking ion species of the same molecule based on known mass differences, adding an MS1-level connectivity layer [115]. Building Blocks-Based Molecular Networking combines neutral loss scanning with molecular networking to identify biogenetically relevant metabolites and streamline MS2 datasets through feature filtering [116]. Substructure-Based Molecular Networking applies unsupervised learning to detect recurring molecular fragments, known as "Mass2Motifs, " across spectra [117, 118]. Bioactivity-Based Molecular Networking integrates chemometric analysis to distinguish active from inactive compounds in complex mixtures, although it does not provide structural details [119].
In addition, multivariate statistical analyses then correlate elicitor identity with metabolomics shifts, flagging high-priority "hits" for follow-up [120]. Recently, interactive visualization platforms such as MetEx, generates interactive multidimensional and 2D plots, enabling global metabolome visualization, cryptic metabolite prioritization, dereplication, elicitor structure–activity relationship analysis, and ranked lead selection [121].
In addition, advanced HiTES variants integrated analytical or imaging modalities boost throughput and resolution. MALDI-HiTES couples MALDI-TOF mass spectrometry with HiTES for rapid metabolite prioritization [122]. Bioactivity-HiTES incorporates assay workflows for direct activity screening [123], HiTES-IMS leverages imaging mass spectrometry [124], and FRET-HiTES uses fluorescent resonance energy transfer sensors to report induction events [125]. For example, Zhang and Seyedsayamdost [122] applied MALDI-HiTES to Streptomyces ghanaensis, rapidly prioritizing and identifying the cryptic non-ribosomal peptide cinnapeptin.
2.4 Chemical elicitors
In diverse ecological niches, microorganisms use NPs as chemical signals to communicate within and between species, coordinate resource use, and trigger metabolite production [126]. In the lab, a wide array of chemical elicitors, including rare elements, heavy metals, hormones, signaling molecules, sulfo-compounds, organic solvents, histone inhibitors, polysaccharides, nanoparticles, metal sequestering agents, and sub-lethal antibiotics, have shown to enhance diverse NP biosynthesis, improving NPD efficiency and yield [82]. Table 2 summarizes several proprietary chemical elicitors and the concentrations (millimolar to nanomolar) at which they activate or repress genes associated with silent BGCs such as metallosensor gene [127], although their precise molecular mechanisms often remain elusive [128]. Nonetheless, discovering new chemical elicitors and optimizing HTS from large libraries remain critical challenges for targeted BGC activation in microbial systems. To overcome this, many groups now combine computational prioritization with miniaturized bioassays (e.g. microplate or droplet-based) to triage hundreds to thousands of compounds in a single run. This integrated pipeline narrows down candidates by predicting which small molecules are most likely to bind regulatory elements or trigger reporter signals, and then validates hits in rapid fluorescence, mass-spec, or bioactivity readouts (see Sect. 2.3). For example, Han et al. [125] used a fluorescence-based DNA-cleavage assay on a 400-compound library and pinpointed five steroidal elicitors that rapidly induced cryptic enediyne production in S. clavuligerus.
Chemical elicitors and selected natural products (sources)
Beyond chemical triggers, abiotic stress conditions like UV/vis irradiation and heat shock (Fig. 3A) also influence discovery and expansion of NP diversity [150]. Engineering the physical culture environment can mimic natural habitats: growing sponge-associated Pseudoalteromonas on cotton balls significantly increased levels of thiomarinol A, violacein, and bromo-alterochromide analogues [162], while adding inorganic talc microparticles accelerated morphological development in actinobacteria and improved oxygen diffusion in Aspergillus terreus mycelia, enhance lovastatin yields [163, 164].
All the approaches mentioned above are low-tech, straightforward methods used to activate silent BGCs or enhance the biosynthesis level to detectable range and are widely employed in NPs research laboratories. However, how to select the effective elicitors, growth parameters, and co-culture partners are central questions for all natural products scientists. Indeed, at present, selecting chemical elicitors or growth parameters or co culture partner(s) to activate BGCs involves a hit-and-trial. Despite their convenience and accessibility, these methods are always time-consuming and inefficient. This knowledge gap exemplifies the utmost need for the development of novel tools/strategies to predict appropriate elicitors or growth parameters for decoding the silent BGCs in vivo rather than complicated systematic trial and error approaches. Additionally, Table 3 outlines the advantages and disadvantages associated with each method.
Highlights and limitations of natural product discovery strategies
3 Advanced culturing techniques and untapped resources or poorly cultivated organisms exploration
It is well-established facts that only 0.1–1% of natural microorganisms can be cultivated in standard lab conditions, with roughly 75% of bacterial phyla lacking cultured representatives [165, 166]. Many microbes "the uncultivated microbial majority" remain uncultivable due to unknown nutrient needs, specific environments, and symbiotic dependencies. Some grow slowly, rely on other species, enter dormancy, or thrive in extreme habitats beyond standard lab conditions [167].
High-throughput culturing techniques (HTCT) have recently begun to improve the cultivation of slow-growing and metabolically talented yet uncultivated microbes, surpassing classical methods limitations (see Sect. 2 and Table 3). HTCT such as micro well and microfluidic device [168], GALT prospector [169] and QPix platform [170] streamline cultivation by miniaturing and automating isolation. When coupled with hyphenated techniques such as IMS [171] and dereplication pipelines [121], these platforms can track both microbial growth and NPs production. Although, the GALT Prospector and QPix excel at handling difficult-to-culture microbes, such as rare human gut bacteria, their use in NPs-focused metabolomics remains underdeveloped [169, 170]. In contrast, miniaturized micro bioreactor systems (e.g. MATRIX 24-well microreactor format) have become increasingly common in NP discovery efforts, enabling scalable cultivation. Using this miniature fermenters, the Capon group uncovered several rare and structurally novel scaffolds, including the 2, 6-diketopiperazine derivatives noonazines A-C, the azaphilone noonaphilone A from Aspergillus noonimiae CMB-M033980 [172], as well as anthelmintic polyketides goondapyrones A–J from Streptomyces sp. S4S-00196A1081 [173]. In addition, high-throughput dilution-to-extinction cultivation and behaviour-based models that mimic specific habitats have been employed to isolate a broad range of rare and underrepresented microbial taxa[174, 175]. These strategies have significantly advanced our understanding of microbial ecological niches. A notable example is the discovery of proteorhodopsin and its presence in Pelagibacter ubique, which underscores the value of accessing and studying culturable microbial isolates[176, 177]. Figure 3D provides an overview of additional modern cultivation techniques alongside representative microbial examples. At the simplest level, miniaturized micro-well culture systems and diffusion chambers allow parallel testing of growth conditions. Microfluidic devices further shrink volumes and increase throughput, while in situ cultivation platforms such as Ichip, C-chip, iTip, and SlipChip, replicate natural ecological niches by permitting environmental nutrients and signaling molecules to diffuse into isolated micro chambers [178].
Single-cell isolation techniques, including fluorescence-activated cell sorting [179], Raman-activated cell ejection [180], harness fluorescence intensity of unique indicators like metabolic activity, resistance profiles, and Raman spectral signatures to target individual microbes from complex consortia, respectively. Notably, the iChip was instrumental in the discovery of teixobactin, a novel antibiotic produced by Eleftheria terrae, a soil bacterium that had previously eluded cultivation [181].
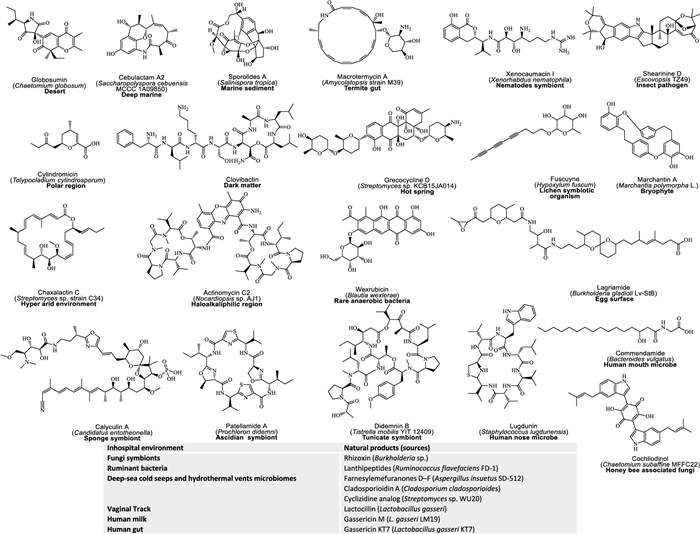
Despite outstanding track records in innovative culturing techniques and NGS, approx. 80% of microbial sources remain underexplored [166]. To wrestle this difficulty, recent efforts have begun to chip away at this "microbial dark matter" by applying high-throughput culturing improvements such as droplet micro reactors, membrane-separated co-cultures, and hyphenated analytics, to activate silent biosynthetic gene clusters in neglected taxa and in-hospital niches [182]. These strategies have already yielded complex polyketides, nonribosomal peptides, polycyclic terpenes, rearranged steroids, and hybrid metabolites from once-inaccessible strains [183–185]. Additionally, Fig. 4 highlights several classical NPs that isolated from rare or extremophile microorganisms thriving in unusual or extreme environments, each paired with its unique environmental source and bioactive scaffold.
Representative examples of natural product from untapped and exotic environments. Each chemical structure is accompanied by its name, and source organism (provided in parentheses), along with the organism's exotic habitat, which are highlighted in bold
Beyond, culture-based methods, breakthroughs in culture-independent workflow (Fig. 5A) have enabled rare bioactive natural products characterization, while multiomic strategies have expanded access to untapped microbial resources, presenting an exciting frontier NPD [186] (Fig. 4). Despite these outstanding achievements in microbial culturing systems, many of these innovative techniques remain underutilized in NPD. The modernization of culturing methods has led to impressive strides in detecting and producing diverse NPs, yet, accessibility issues and implementation complexities hinder their full potential. Many natural product researcher find these emerging technologies out of reach, often due to limitation in resources, infrastructure, and technical expertise.
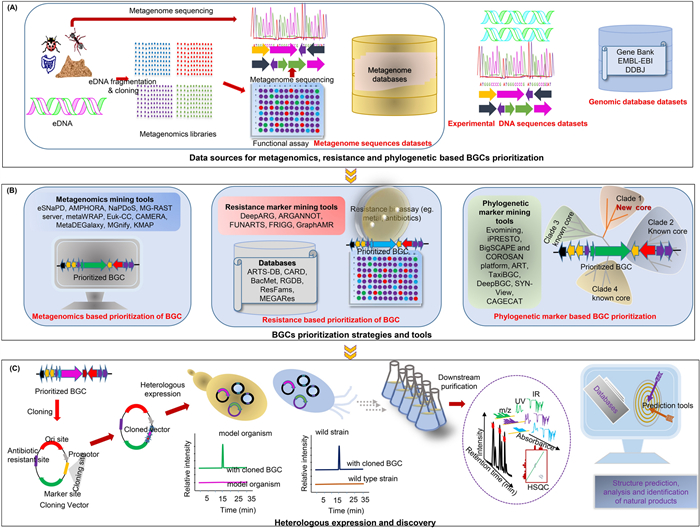
Schematic workflow and tools for genomic strategies in natural product discovery. Phylogenetic based mining, metagenomics based mining, and resistance gene based mining A Data sources; B Mining tools and databases and C Cloning, heterologous expression and identification of metabolites)
4 Genomic and metagenomics innovations driving natural product discovery
NPs exhibit immense chemical diversity, yet their biosynthetic machinery is often highly conserved. Biosynthetic core enzymes typically share significant amino acid sequence similarity, enabling researchers to screen genomic data for specific biosynthetic genes that encode key enzymatic functions using genome mining tools [187]. Unlike bioactivity-guided isolation strategy, genome-based approaches are highly specific but do not provide immediate insight into a compound biological activity. In particular, in genome inspired discovery, researchers first identify candidate BGCs, benchmark enzyme domains against known pathways, and then experimentally probe the function and chemical output of these clusters [188]. Consequently, this strategy also led to the characterization of biosynthetic enzymes from uncultured microorganisms and cryptic BGCs that catalyze novel and exceptional chemistry, potentially linked to metabolites that are still poorly understood or entirely unknown [189].
Additionally, technological upgrade in sequencing have fueled this genomics driven NPD renaissance. Long read platforms (e.g. PacBio SMRT, Oxford Nanopore MinION), and high-throughput shorts reads (e.g. Illumina) have enabled rapid, high-quality, and cost-effective whole-genome and metagenomics assemblies [190]. At the same time, genomic studies have uncovered that bacteria, fungi, and even complex organisms possess a far greater biosynthesis capacity than lab experiments typically indicate [18], likely due to issues such as gene silencing and low metabolite yields. Unlocking these hidden compounds requires targeted- strategies or stimuli to activate silent or weakly expressed BGCs [191]. Nonetheless, key challenges such as identifying and prioritizing promising BGCs, effectively switching them on, and linking each BGC to its metabolite exists. To overcome these bottlenecks, a range of bioinformatics pipelines, genome-mining algorithms, integrative databases and online resources has emerged (Table 1). These resources enable researchers to identify BGCs, predict their chemical outputs, and rapidly dereplicate known compounds. Tools such as antiSMASH, and BiG-SCAPE facilitate the annotation, clustering, and prioritization of BGCs, while platforms like MIBiG and NPAtlas provide reference datasets for comparative analysis. When integrated with metabolomics workflows and interactive dashboards as detailed in Table 1, these platforms streamline NP discovery by reducing redundancy, boosting novelty detection, and directly correlating genomic predictions with LC–MS data [192].
Chronologically, evolution-based phenotypic characterization (1940–1970s), knowledge-based approaches (1970–1990s), computational-based approaches (1990s to early 2000s), and genome-AI integrated approaches (mid-2000s onward) indicate the trend and progress of genome mining [193]. Despite its remarkable progress, connecting genetic information to the enzymatic and structural characterization of the encoded NPs remains a significant bottleneck (roughly 90% of Actinobacterial BGCs are still uncharacterized) [18]. To bridge this gap, large-scale data-driven bioinformatics platforms now multiplexed with hyphenated analytical datasets (e.g. iSNAP) [194], microcrystal electron diffraction techniques (MicroED) [195], and machine- and deep learning architectures [39]. This enables the characterization of novel molecules, chemical building blocks, biosynthetic signatures, tailoring enzymes, and biosynthetic pathways [43], by targeting chemical features, families, entire BGCs or only domains while minimizing labor-intensive efforts and rediscovery, and maximizing NP diversification [196]. For example, Kim et al. (2021) employed genome mining to pinpoint the icc BGC in Penicillium variabile, leading to the discovery of a novel 2-pyridone natural product, Py-469 and solved the structure via MicroED, demonstrating the synergy between genomic prediction and structural elucidation [195]. Likewise, Li et al. [195] identified a previously cryptic csp cluster in the anaerobe Clostridium roseum; through gene knockouts and heterologous expression, they confirmed its role in biosynthesizing the new clostyrylpyrones. Moreover, Liu et al. (2025) developed NegMDF, a workflow integrating mass defect filtering and bioinformatics to link BGCs with metabolite ions under negative ionization. Applied to Streptomyces cattleya NRRL 8057, it rapidly identified 22 polyketides, including rare tetronate-containing cattleyatetronates. Collectively, these case studies illustrate how coupling advanced bioinformatics, structural techniques, classical genetics, and metabologenomic can unlock the vast chemical potential encoded in silent BGCs.
Metagenomics, first coined in the late 1990s, enables culture-independent exploration of microbial genomes, including those from individual cells, across a wide range of extreme and untapped environments such as Polar Regions, deep-sea ecosystems, and the gut microbiome. Over the past two decades, its robustness has powered discovery of novel BGCs and pathways in uncultivable microbiomes, often by directly cloning environmental DNA libraries into heterologous hosts for expression [186, 197, 198]. Standard metagenomics protocols (Fig. 5A employ functional screening of eDNA or shotgun sequencing followed by BGC prediction. Indeed, a crucial pillar in NPD, this approach faces challenges such as labor-intensive procedures for constructing and screening large libraries, dealing with non-genomic DNA during metagenomics bin assembly, and masking low-abundance microorganism DNA, making it difficult to accurately link metabolites or genes to phylogenetic knowledge [199, 200].
To combat these issues, targeted phylogenetic metagenomics that focuses on capturing large BGC fragments by PCR amplification of conserved biosynthetic domains (e.g. AT, KT, ACP) directly from the eDNA [201]. And, parallel strategies screen for entirely novel biocatalysts in metagenomes using activity-based assays [202] and leverage computational tools for in silico prioritization. More recently, modern consortiums and web-based annotation platforms such as MetaSU for urban environments [203] and MetaHIT for the human gut [204] now map AMR markers and discover new BGCs global scale.
Additionally, single-cell metagenomics adds another dimensions by sorting individual cells from environmental samples via FACS [205] or microfluidic droplet platforms [206] before whole-genome amplification and sequencing, unlike traditional metagenomics, which sequences pooled DNA from entire communities. This approach enables precise bioinformatics assignment of genes to specific microbes, making it especially useful for studying uncultivable organisms. By linking metabolites and genetic pathways to phylogenetic data, it offers deeper insights into microbial diversity and biosynthetic potential serving as a powerful complement to community-level sequencing methods [207].
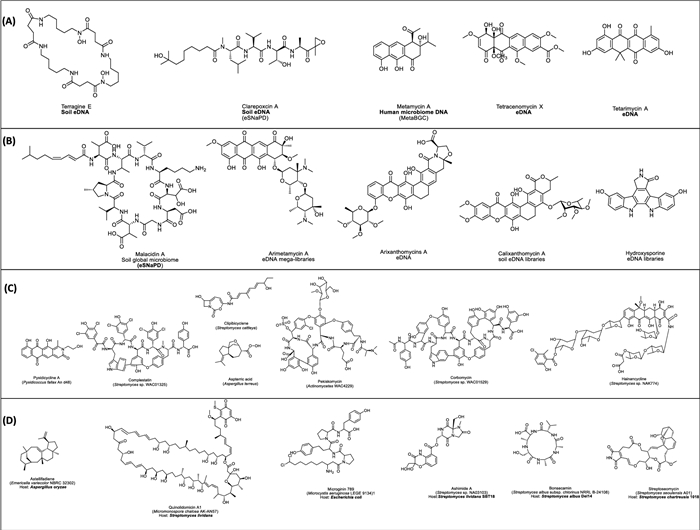
Traditional and modern single-cell metagenomics techniques are powerful tools for exploring uncultured microorganisms in environmental samples or eDNA. These approaches have enabled the discovery of diverse NPs like apratoxin A (e.g. konbamides, nazumamide A, keramamides, cyclotheonamide), misakinolide A and theonellamides [200]. Additionally, Fig. 6A–B highlights additional examples, discovered through metagenomics approaches, particularly from uncultured or symbiotic microorganisms. These discoveries underscore the power of both traditional and single-cell metagenomics in accessing cryptic BGCs and expanding the chemical diversity.
Examples of representative examples of natural products (NPs) discovered with different genomic mining approaches. A Examples of NPs discovered via metagenome mining B Examples of NPs isoalted by resistance gene mining, C Examples of NPs discovered via phylogeny-inspired mining and D heterologous expression based discovered NPs. Each chemical structure is provide with its name, sources or tools/expression system applied
5 Large-scale genome mining in modern genomics: exploring resistance-gene or phylogenetic relationship
5.1 Resistance gene based genome mining
Prioritizing BGCs for targeted production of NPs with desired bioactivities remains a complex challenge in genome mining. Beyond core synthesis enzymes, BGCs often include genes for transport, modification, and self-resistance. Self-resistance clusters have evolved to neutralize environmental antibiotics via detoxification, efflux pumps, target binding or modification, and even horizontal gene transfer [208]. Exploiting these resistance genes as "molecular beacons" provides a powerful genome-mining strategy: by flagging clusters that harbor self-resistance determinants, researchers can predict NP modes of action and narrow down BGCs for further study [64, 209]. For example, Panter et al. [210] screened myxobacterial genomes for pentapeptide-repeat proteins homologous to the cystobactamid resistance mechanism and discovered the pyxidicyclines, a new class of type Ⅱ polyketides with a nitrogen-containing tetracene core. These compounds inhibit bacterial topoisomerase Ⅳ (IC50 1.6–6.25 μg/mL) and human topoisomerase I, and exhibit potent cytotoxicity against HCT-116 cells (MIC 0.06 μg/mL).
Additionally, modern bioinformatics platforms such as ARTS-DB [64], CARD [65], DeepARG [66] automate the identification, annotation, and functional analysis of resistance genes from genomic and metagenomics data, accelerating resistance-guided NP discovery. As listed in Table 1F, multiple databases and tools support this workflow; and Fig. 6B highlights notable NPs uncovered through resistance-gene directed mining.
5.2 Phylogenetic relationship based genome mining
Complex evolutionary and metabolic processes such as insertions, deletions, duplications, rearrangements, and both vertical and lateral gene transfer shape multigene BGCs, which encode proteins for core molecule synthesis, diversification, regulation, and transport [211]. Conserved enzyme domains such as ketosynthase alpha (KSα) and beta (KSβ) often evolve through concerted mechanisms, originating from within BGCs, from other clusters, or even from central metabolism [212, 213]. By building phylogenies around these domains, researchers can pinpoint divergent or novel sequences and then mine representative genomes for the underlying chemistry. For example, Mullins et al. (2021) reconstructed a phylogeny of alkyne and polyyne biosynthetic cassettes, allowing them to identify distinct phylogenetic clades of interest. By mining representative genomes from these clades, they discovered previously uncharacterized polyyne BGCs. Notably, within the Gammaproteobacteria clade; led to the discovery of a novel polyyne "protegencin" produced by Pseudomonas protegens (formerly P. fluorescens) strains Pf-5 and CHA0. Similarly, Deng et al. (2025) discovered mandimycin, a polyene antifungal antibiotic, using a phylogeny-guided platform based on conserved mycosamine-transferring glycosyltransferases, a modification enzyme from S. netropsis DSM 40259. Mandimycin showed potent, broad-spectrum activity against multidrug-resistant fungal pathogens, with MICs ranging from 0.125 to 2 μg/mL [214]. Additionally, Table 1 summarizes automated bioinformatics tools that automate phylogeny-based NPD and outlines their specific functions and features, and Fig. 6C presents additional landmark phylogeny-guided discoveries.
Recently, beyond single-cluster screens, large-scale pan-genomic mining of closely related entire taxa [215], genus [216], special niche microbiome (e.g. Ocean, acid mine drainage, soil) [217, 218] and specialized strain collections [219] has revealed thousands of BGCs and unraveled their evolution relationships. To date, diverse microbial sources including Shark Bay microbial mats, Virgibacillus, Cyanobacteria, the swine gut microbiome, entomopathogenic nematode-symbiotic bacteria, Streptomyces, Saccharomonospora, Burkholderia, marine prokaryotes, turtle ant gut-symbionts, Penicillium, and archaea have been mined for NPs. These mining efforts systematically investigate thousands of BGCs, revealing complex evolutionary relationships and mechanisms [220]. In particular, setting biosynthetic genes or specific functional domains within the BGCs into a phylogenetic relationship with known sequences to track the proximity and outliers to the known sequences [221] predicts and prioritizes substructures and full chemical structures from BGCs, often facilitated by recent automatic platforms.
Notably, automatic platforms, EvoMining for enzyme family evolution [222], DeepBGC for machine learning based BGC detection [223], TaxiBGC for taxon-guided mining [215] and BiG-SCAPE/CORASON for chemical-family clustering [36, 224] predicts and prioritizes substructures of both substructures and full chemical scaffolds. NaPDoS, for instance, a bioinformatics tool to study the phylogenetic relationship studies of PKS and NRPS domain. Researchers used the tool to analyze ketosynthase (KS) domains from the marine actinomycete genus Salinispora [225], and Cruz-Morales et al. [226] applied EvoMining to actinobacterial genomes and reconstructed the evolutionary history of 23 enzyme families, uncovering a previously unrecognized BGC in Streptomyces coelicolor and S. lividans responsible for producing arseno-organic metabolites.
Focusing on candidate BGCs associated with resistance genes offers a strategic advantage, as the resulting compounds are more likely to possess biological activity. This approach is grounded in the idea that the compound's mechanism of action mirrors the native function of the resistance determinant, thereby enabling for more accurate predictions of bioactivity. In contrast, phylogeny-based studies expand search insights by linking to habitat-specific, taxonomic (species/genus), behavioral, or evolutionary traits to novel chemistries, beyond relying solely bioactivity-driven selection alone. Together, resistance- and phylogeny-guided mining form complementary pathways that accelerate the discovery of NPs with both new structures and targeted bioactivities.
6 Downstream applications of genome mining for natural product discovery: cloning, heterologous expression and CRISPR-Cas
6.1 Cloning and heterologous expression
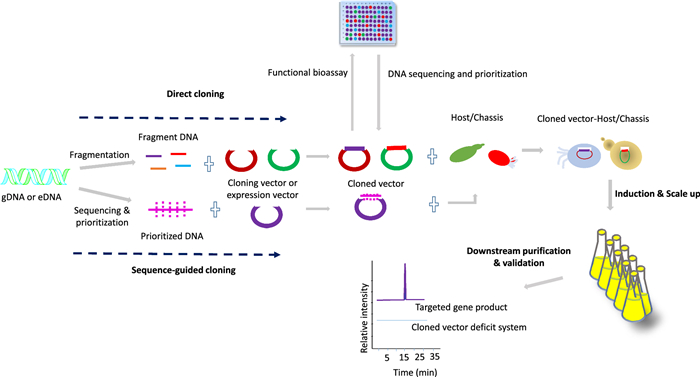
Beyond conventional methods like phenotype screening, co-culture, and elicitor-based assays, an emerging strategy involves cloning and heterologous expression of BGCs within optimized host organisms. Cloning and heterologous expression systems are indispensable and closely linked for overproduction and NPs discovery. First introduced in the 1960s, scaled up with recombinant DNA advancements in the 1990s, enabling efficient biosynthetic gene expression in model organisms to optimize titer, rate, and yield ((TRY), including NCEs discovery [76, 227]. In particular, this technique involves cloning and expressing engineered gene clusters often sourced from untapped and extreme environments [76, 228]. In addition, it enables functional expression of previously silent or uncharacterized pathways that may encode valuable bioactive compounds or NCEs [76, 226]. Figure 7 illustrates the foundational workflow involved in cloning for NPD: prioritize and prepare DNA fragments, choose suitable cloning vectors and assembly methods, then select and engineer host strains to functionally express silent or novel pathways.
Schematic workflow for cloning and heterologous expression in natural product discovery
Early cloning vector systems such as cosmids and fosmids accommodated inserts up to ~ 50 kb, but many BGCs fall in the 100–350 kb range. To handle these larger clusters, researchers developed high-capacity vectors P1-derived artificial chromosomes, bacterial artificial chromosomes, and yeast artificial chromosomes, capable of stable maintenance of large DNA inserts for complex biosynthetic pathways and expression. Detailed information on these vectors is available in specialized review article [229]. While E.coli followed by Streptomyces, Schizosaccharomyces, Saccharomyces and Aspergillus were the contemporary heterologous expression system. Nevertheless, their applications remains limited by incompatibilities in transcriptional regulation, codon usage, and lack of post-translational modifications. Moreover, other challenges include insufficient precursor/co-factor availability, product toxicity, and poor biosynthetic assembly of NCEs. Eukaryotic gene clusters present additional hurdles, requiring intron splicing and strategic insertion of promoters and terminators for functional expression [230]. Of late, to overcome these limitations, diverse engineered streptomycetes such as S. avermitilis, S. venezuelea and emerging non-Streptomyces hosts, for instance, Myxococcus xanthus, Yarrowia lipolytica, Burkholderia thailandensis strain E264, Bacillus subtilis, P. putida have been the choice of chassis for the HES for more compatible expression environments [231, 232].
Similarly, selecting an appropriate recombination method is equally critical in success of cloning-based approaches for NPD. The choice of DNA assembly method influences the efficiency, accuracy, and scalability of DNA assembly particularly when dealing with complex BGCs. Classical in vitro DNA assembly methods, such as restriction enzyme-mediated digestion and ligation, rely on specific recognition sequences, limiting design flexibility and making them inefficient for assembling large or multiple DNA fragments. These approaches often leave behind unwanted scar sequences and require labor-intensive steps like digestion, purification, and transformation making them ill-suited for high-throughput or modular synthetic biology applications [229]. In contrast, modern recombination-based strategies including Gibson Assembly, DiPaC, CATCH, and CCTL enable seamless, scarless assembly without dependence on restriction sites, while enzyme-independent approaches offer additional design versatility. Additionally, in vivo approaches such as phage recombinase-mediated homologous recombination in E. coli, TAR cloning in yeast, and site-specific integrase systems (e.g. Cre/loxP, ΦC31, ΦBT1) allow efficient and precise capture of large DNA segments directly within host cells, further expanding the toolkit for complex NPD [233, 234].
6.1.1 Direct cloning
Direct cloning (Fig. 7), where fragmented environmental or genomic DNA is immediately inserted into expression vectors without prior sequencing. These vectors are then used to construct large libraries, which are screened through in vivo or in vitro functional assays to identify bioactive compounds or novel biosynthetic pathways. This strategy is particularly valuable when genomic information from the native host is poorly characterized or unavailable. By bypassing the need of prior sequence information, sequence-independent approaches enable comprehensive exploration of a sample genetic content, often increasing the likelihood of capturing entire BGCs and uncovering structurally novel NPS.
However, direct cloning faces notable constraints. Random insertion of DNA fragments may yield incomplete or nonfunctional BGCs, and large clusters frequently exceed standard vector capacities. In addition, native regulatory elements may be incompatible with the heterologous host, leading to inefficient transcription and translation. Moreover, functional screening is labor-intensive and prone to false negatives due to the low frequency of bioactive clones.
To overcome these limitations, careful pairing of cloning systems with high-throughput screening assays is essential. In place of traditional E. coli, researchers increasingly opt for phylogenetically closer expression hosts such as S. lividans, S. albus, and P. putida. Additionally, introducing regulatory elements such as sigma factors into heterologous hosts can significantly enhance the expression of target biosynthetic genes. Figure 6 highlights several classical examples of NPs discovered through both direct cloning and the use of advanced heterologous expression systems.
6.1.2 Sequence-guided cloning
Sequencing-guided cloning begins with the sequencing of environmental or genomic DNA, followed by bioinformatics analysis to identify and prioritize BGCs with high potential for producing bioactive compounds [235]. Tools such as antiSMASH 7, PRISM 3, ARTS, and BiG-SCAPE assist in predicting BGCs functions and selecting promising candidates [236] (see Table 1B–C). Once prioritized, the targeted gene clusters are cloned into suitable expression vectors, enabling direct functional characterization and NPD. This approach bypasses traditional random library construction and broad screening by focusing efforts on specific, high-value clusters, streamlining the discovery of novel NPs.
A notable example is the discovery of cadasides A and B, acidic lipopeptides, acidic lipopeptides identified by Wu et al. [237], sequenced nonribosomal peptide synthetase (NRPS) adenylation domains to pinpoint and clone the cde BGC from calcium carbonate rich soil samples. Their analysis revealed a correlation between cadaside-like domain abundance and specific geochemical environments, enabling targeted recovery and successful expression of the cluster. Similarly, Zhao's group cloned and heterologously expressed 105 BGCs in Streptomyces lividans TK24. Their use of the Antibiotic-Resistant Target Seeker tool to prioritize clusters based on self-resistance gene markers led to eight novel bioactive compounds from five productive BGCs, with efficient cloning achieved using the CAPTURE platform [238].
Together, Cloning and HES have revolutionized NPD, enabling the scalable production of novel compounds and unlocking hidden chemical treasures (For instance, Fig. 6D). Yet, research on selecting the optimal cloning vector, heterologous host for maximum natural product yields remains a significant challenge due to the differences in metabolic scaffolding among hosts, such as sporulation, machinery tools, biofilm formation, and autolysis.
6.2 CRISPR-Cas9 system
The CRISPR-Cas9 system, originally derived from bacterial adaptive immunity and conceptually identified in the late 1980s, became a programmable genome-editing tool in the early 2010s. Its emergence revolutionized genetic engineering and has since played a crucial role in NPD, especially for targeted genetic disruption. By enabling precise modifications such as gene knockouts, knock-ins, and knockdowns, CRISPR-Cas9 allows researchers to screen specific phenotypes and manipulate biosynthetic and metabolic pathways to enhance NPs yields and unlock new compounds [239, 240].
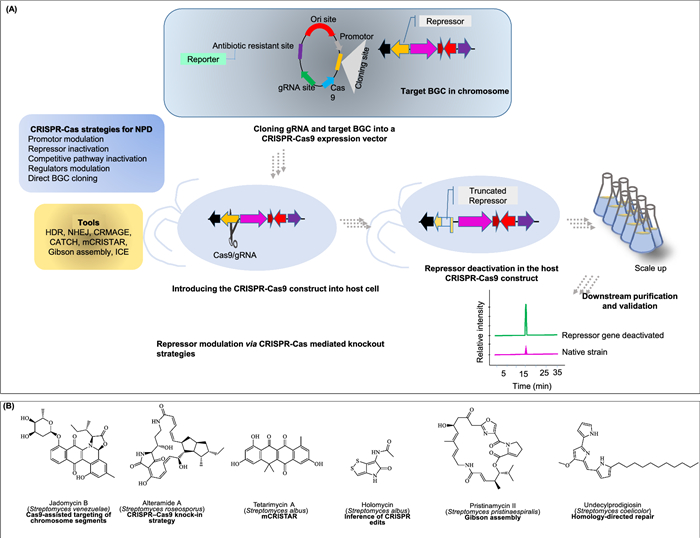
Earlier genome editing technologies such as meganuclease I-SceI [241], zinc-finger nucleases [242], and transcription activator-like effector nucleases [243] formed the basis for CRISPR-Cas9. This technology marked a significant milestone in genomic biology [244], enabling applications such as targeted gene regulation, epigenetic perturbation, chromatin manipulation, and live cell chromatin imaging paradigm [245]. Figure 8A illustrates the workflow of the CRISPR-Cas knockout strategy used in NPD.
CRiSPR-Cas for natural product discovery. A Workflow of knockout strategies of CRiSPR-cas strategies. B Examples of discovered metabolites via CRiSPR-cas strategies
Importantly, CRISPR-Cas systems are not primarily used to discover entirely new BGCs, but to investigate, optimize strain improvement and scalable production function of known ones. Through precise genome editing, CRISPR-Cas enables researchers to dissect the roles of individual genes within biosynthetic pathways, identifying key enzymes, regulatory elements, and metabolic bottlenecks. More importantly, it allows for the fine-tuning of these pathways by knocking out inhibitory genes, activating silent clusters, or engineering promoter regions, ultimately boosting the yield and efficiency of NPs biosynthesis [246, 247]. Consequently, CRISPR-Cas9-based genome editing is extensively used for genetic and metabolic manipulation of bacteria, yeast and plants to discover hidden chemical treasure troves [248].
Several hallmark studies illustrate the breadth of CRISPR-Cas applications in NPD. For example, Bushin et al. [249] identified a quorum-sensing, regulated RiPP gene cluster from Streptococcus cloned using CRISPR-assisted and expressed the cluster in S. albus, leading to the production of of streptosactin, a sactipeptide with unique crosslinks and strong antifungal activity against Candida albicans and Aspergillus fumigatus. Ameruoso et al. [250] developed CRISPR interference (CRISPRi) and activation (CRISPRa) systems in Streptomyces venezuelae to regulate transcription of silent biosynthetic gene clusters. By modulating key regulatory networks, they successfully activated the jadomycin B cluster, leading to natural product synthesis previously undetectable under standard conditions. Likewise Peng et al. [251] constructed a CRISPR-dCas9 system in Myxococcus xanthus, enhancing epothilone B production. By fusing dCas9 with a transcriptional activator and targeting promoter regions of the 56-kb biosynthetic gene cluster, they achieved a 6.8-fold yield increase. These three examples illustrate the successful application of CRISPR-Cas technology in distinct areas of natural product research: the discovery of new compounds, activation of silent gene cluster and the scalable production of valuable metabolites. Figure 8B presents examples of NP modulated using CRISPR-Cas technology.
Despite its versatility, CRISPR-Cas9 faces limitations such as off-target effects and strict PAM sequence requirements for SpCas9 from Streptococcus pyogenes, which is widely used in bacteria and archaea [248]. To overcome these challenges, improved Cas proteins such as saCas9 and fnCpf1, have been developed or engineered, offering greater specificity and broader target accessibility [252]. For instance, in a recent study by Zhou et al. [253], researchers addressed the limitations of class 2 CRISPR systems like Cas9, which often underperform in Streptomyces, by repurposing the native type I-E CRISPR system into transcriptional regulators (CRISPRi and CRISPRa). Applied across nine diverse Streptomyces strains, these tools successfully activated 13 of 21 cryptic biosynthetic gene clusters, revealing five new NPs: one polyketide, one RiPP, and three alkaloids.
7 Advances in genomic mining-based metabolic engineering for natural product discovery
Historically, metabolic engineering aimed to enhance the TRY of target molecules using cost-effective nutrients, particularly in pharmaceuticals production [254]. The biosynthesis of NPs, however, is influenced by a variety of factors, including growth conditions, carbon source availability, and complex layer of genetic regulation. These factors pose significant challenges when attempting to clone and fine-tune large BGCs, particularly because many of these clusters remain genetically intractable, exhibit poor expression under laboratory conditions, and are often silent, meaning they are not naturally expressed due to unknown regulatory mechanisms, diverse secretion systems, and intricate metabolite profiles. Modern bioinformatics tools and omics knowledge, including "metabolic engineering" and "BGC refactoring" have addressed some of these limitations of NPD exploration and overproduction [230, 255].
Over the past decade, its application has expanded to manipulate biological systems to build alternative pre-programmed systems harboring features for targeted compounds biosynthesis. A key focus has been on the prediction and reconstruction of metabolic pathways using available genomic and metabolic data. Recently, three prominent tools that have emerged to support this effort include GENREs, BiGMeC pipeline, and Galaxy-SynBioCAD portal. GENREs provides a quantitative framework that integrates genomic, biochemical, and phenotypic data to model and analyze biochemical reactions systematically across defined metabolic categories [256]. While, the BiGMeC pipeline, on the other hand, automates the reconstruction of metabolic pathways specifically associated with PKS and NRPS gene clusters, streamlining NPD from complex genomic datasets [257]. Galaxy-SynBioCAD portal facilitates the creation of strain libraries optimized for producing specific chemical compounds, encompassing the entire workflow from selecting target molecules and host strains to designing and engineering complete metabolic pathways [258] (Table 1D–E). Additionally, recent efforts have focused on constructing metabolic pathways by leveraging time-series multi-omics data to capture metabolic dynamics [259], alongside analyzing flux distributions [260] is discussed elsewhere.
Basically, metabolic engineering in NPD aims at, (i) enriching the target compounds TRY [261], (ii) modifying NPs scaffolds for improved bioactivity [262], (iii) engineering and expressing the BGCs from diverse sources (e.g. marine microorganism [263], endosymbionts [264]), and (iv) dissecting unsolved biosynthesis pathways [262]. Although, the term "metabolic engineering" is relatively generic, its core principles have been consistently recognized across the literature and are summarized in Table 4.
Major metabolic engineering strategy
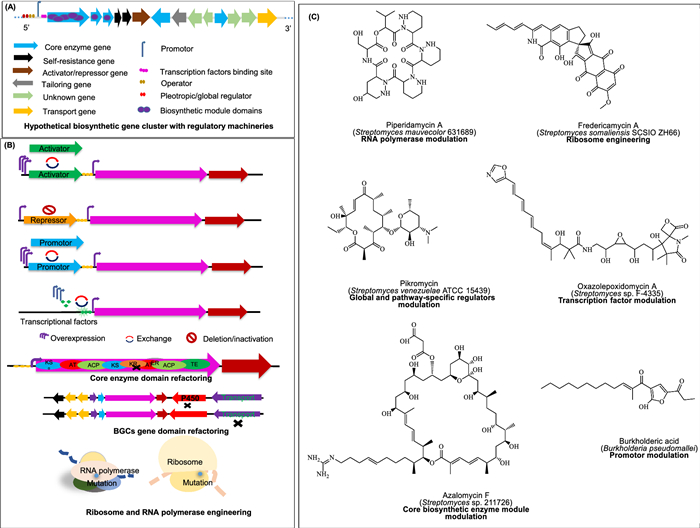
Undoubtedly, the various metabolic engineering strategies outlined in Table 4 work synergistically to support either NP enhancement or NPD. Among them, BGC refactoring enables activation of silent pathways and diversification of NPs. It involves core biosynthetic genes modulation such as modular domain- deletion [271], addition or sequence alteration [272], shuffling [273]. For instance, in a recent study, the Calcott research group introduced over 1, 000 unique metagenomics domains into a pyoverdine NRPS system, resulting in the production of 16 distinct pyoverdines as major products [274]. In contrast to single-module modifications, Song et al. [275] developed RedEx, a technique that enables the precise insertion or deletion of large DNA fragments within extensive polyketide synthase (PKS) clusters. Using this approach, they successfully replaced the C-21 ethyl group of the macrolide insecticide spinosad with a butenyl group, creating a new analog with modified structure. A complementary strategy focuses on engineering non-core biosynthetic elements such as promoters; regulatory genes (see Sect. 7.1–7.3).
Despite numerous innovative protocols advancing NP discovery and production, the formation of toxic intermediates remains a persistent challenge. These compounds can disrupt biosynthetic pathways and trigger feedback inhibition, posing significant barriers to pathway stability and productivity. The mechanisms underlying their generation and impact are still not fully elucidated. To address this issue in metabolic engineering, several strategies have been developed. These include enhancing host organism tolerance to toxic metabolites, integrating continuous flow bioreactors to facilitate real-time toxic metabolites removal, and fine-tuning regulatory elements such as promoters, ribosome binding sites, and terminators (Fig. 9A–C), to ensure balanced pathway expression and minimize accumulation of harmful intermediates.
Metabolic engineering strategies for fine-tuning biosynthetic gene cluster domains A A schematic diagram for hypothetical BGC. B Different metabolic strategies applied to activate BGCs. C Examples of NPs discovered viz-metabolic modulation strategies. Each metabolite is provided with its name, source and strategy applied
7.1 Post- and transcriptional regulatory features modulation
Modern bioinformatics analysis uncovers that over half of BGCs harbor regulatory genes, but the networks controlling BGC expression remain complex and not fully understood [276]. These regulatory proteins bind to DNA elements influencing transcription levels [277] and can be classified as global (e.g. N-utilizing factor G, A-factor-dependent protein A, CodY, LaeA, MilR3), epigenetic (Mehat, NnaB, rpdA) and pathway-specific (e.g. samR0484, gbnA/gbnB, tnmR1/tnmR3/tnmR7, ACE1, chal, SlnR, MonH/MonRI/MonRII) [278, 279]. For a deeper understanding of individual regulators roles in NPD, readers are referred to literatures on epigenetic-[280], fungal-[278] regulatory system. Over the past decade, engineering these regulators has markedly improved silent BGC activation and NP yields in Bacillus [281], Streptomyces [282], Penicillium [283] and Aspergillus [284]. Briefly, strategies for gene expression modulation include disrupting negative regulator genes [285], altering mRNA processing [286], inserting synthetic regulatory systems [287], and modifying epigenetic processes [280]. Although RNA interference based approaches for BGC expression modulation show promise, their application in NPD remains in its infancy and has so far enabled the discovery of only a few NPs (e.g. platensimycin and platencin [288], phomallenic acids A-C [288]).
Advanced high-throughput biosensor enabled screening systems have revolutionized the way we modulate regulatory networks in microbial hosts by directly coupling intracellular metabolite levels to easily measured signals. For example, transcription factor-based biosensors such as VanR-VanO vanillate sensor [289] and FRET-based and NADH/NAD + based ratio metric biosensor [290] allow dynamic monitoring and control of pathway intermediates and redox states in real time. When combined with automated liquid-handling platforms and microfluidic cultivation, these biosensors enable systematic optimization of host chassis, genetic constructs, and culture conditions, while also supporting high-throughput screening of vast regulator libraries to pinpoint variants that enhance target molecule production [291, 292]. Complementing these experimental advances, the Ligify software mines enzyme-reaction databases to predict transcription factors likely responsive to user-defined chemicals, thereby accelerating the design of bespoke biosensors for novel NPs [293].
7.2 Promoter modulation
Promoters, small DNA segments located upstream of the 5′-ends of structural genes, enable RNA polymerase recognition and binding, thus controlling gene expression. These elements can be identified in silico using tools such as iProEP [294] and PromGER [295], and validated in vivo through techniques like ChIP-on-chip [296, 297] or electrophoretic mobility shift assays [298]. Because native promoters are often quiescent under laboratory conditions, they are prime candidates for modification by random- or site-directed mutagenesis, hybridization, error-prone PCR, and sequence randomization. Engineering or substituting these dormant promoters with well-characterized constitutive, host-specific, or inducible promoters has proven highly effective for activating cryptic BGCs and enhancing compound titers [299–301], including in actinomycetes, E. coli, cyanobacteria, and fungi [302]. For instance, Lin et al. [303] introduced the inducible promoters alcA and aldA, which are activated by alcohols, aldehydes, and ketones, to drive expression of the sartorypyrone BGC (spy) from A. fumigatus Af293 in the heterologous host A. nidulans, yielding twelve sartorypyrones (five known and seven novel).
Orthogonal synthetic circuits such as the Q-system from Neurospora crassa [304] or synthetic Tet-On/Off in Aspergillus[305] repurpose non‐native regulators to drive BGC transcription without perturbing host networks. For instance, harnessing the modular Q-system from Neurospora crassa, Lalwani et al. [304] engineered optogenetic circuits in Saccharomyces cerevisiae to achieve precise light-responsive gene regulation. These include two complementary platforms: OptoQ-INVRT circuits, which activate transcription in darkness, and OptoQ-AMP circuits, which trigger robust expression under blue light, yielding up to a 39-fold increase in gene activity for geraniol and linalool terpenoids production.
While diverse promoter libraries accelerate pathway regulation, they struggle to coordinate multi-gene expression in complex networks. To address this, researchers now pair promoter tuning with metabolic flux analysis [306], real time intermediate biosensor [307], mathematical model simulation [308], and AI-driven construct design [309]. For instance, in a recent study, Liu's team enhanced GlcNAc production in B. subtilis by integrating promoter tuning with a real-time GlcN6P biosensor and ADC-based feedback circuits. This dynamic regulation increased GlcNAc titers in a 15 L fed-batch bioreactor from 59.9 to 97.1 g/L (with acetoin) and from 81.7 to 131.6 g/L (without acetoin), demonstrating the robustness and scalability of promotor integrated approaches [310].
Recent advances in AI-based promoter modeling have harnessed deep learning and generative frameworks to predict and design synthetic promoters with precise strength, specificity, and regulatory features for diverse microbial hosts [309]. At the same time, CRISPR-Cas9 mediated promoter replacement enables exact, in situ swapping of native regulatory sequences with engineered or inducible promoters directly within biosynthetic gene clusters, dramatically improving transcriptional tuning [311]. In addition, tools like Easy Promoter Activated Compound Identification [312] uses in situ promoter exchange to selectively activate BGCs encoding NRPS, PKS, NRPS-PKS hybrids, or other BGC classes, yielding targeted NPs. Applied to Xenorhabdus mutants, this approach uncovered antiprotozoal metabolites including fabclavines, xenocoumacins, xenorhabdins, and PAX peptides [313]. Together, these integrated approaches deliver efficient, dynamic transcriptional control and underscore the transformative power of promoter engineering in NP biosynthesis.
7.3 Editing ribosome binding sites (RBSs) and terminators
Transcription regulation of BGCs is complex and influenced by various internal and external factors [314]. At the transcriptional level, synthetic transcription factor decoys can be tuned via copy number or decoy‐site sequence to control native and heterologous gene expression, driving a 16-fold increase in arginine production in E. coli [315]. At the translational level, the ribosome-binding site's translation initiation rate (TIR) is critical for balancing multi‐gene operons; engineering RBS nucleotides through design and HTS of synthetic [316] or pre-characterized libraries [317] fine-tunes TIR to optimize biosynthetic output. For example, targeted mutations in the RBSs of vioB, vioC, vioD, and vioE relieved bottlenecks in violacein biosynthesis, yielding a 2.41-fold titer increase in E. coli [315]. Similarly, constructing a 5′-UTR library by random base insertions on truncated upstream sequences boosted riboflavin titers 2.09-fold in B. subtilis [318].
Further improvements in transcript stability and pathway efficiency come from terminator engineering [319], sigma factors modulation [320] including ribosome and RNA polymerase engineering [321], can also be crucial for maximizing NPs output enhancing BGC expression. Finally, RNA-based post-transcriptional controls such as riboswitch [322] and riboregulators [323] offer additional tuning layers but remain underexploited for BGC regulation.
8 Strategies in natural product analogues discovery
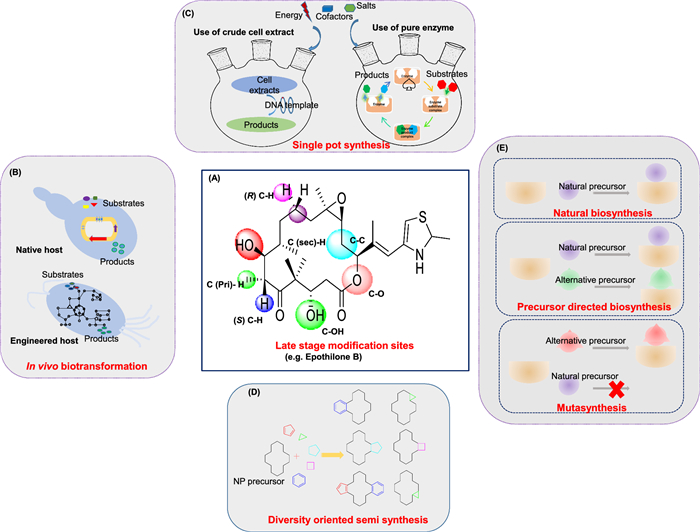
The abundance of stereo centers and unusual carbon skeletons (e.g. multiple- rings system, functional groups), along with the challenges of synthesis involving harsh reactions and critical (de) protection steps, pose significant challenges for NPs synthetic chemists to synthesize complex NPs scaffolds [324]. Discovery of novel catalysts, reagents, selective reactions, and strain selection, often combined with computational approaches, have enhanced NP diversification [325], even molecules not found in the nature. Efficient chemical modifications, like systematic ring-distortion and functional group additions (e.g. halogenation, oxidation, epoxidation), create NP-like small molecules with improved biological activities [324, 326]. Figure 10A highlight possible functionalizing of NP Epothilone B. Additionally, advances in large-scale metagenomics sequencing have uncovered novel biosynthetic enzymes with the ability to catalyze a wide array of chemical transformations [202], paving the way for microbial biotransformation and cell-free extract-based synthesis of valuable compounds or unprecedented chemical scaffolds.
Schematic illustration of workflow of late stage diversification. A Late-stage functionalization in Epothilone B is illustrated through its molecular structure, where colored spheres highlight the specific bonds and functional groups modified. B In vivo biotransformation viz model organism and wild strain. C Single pot synthesis via cell free extract utilization and enzymatic recycle platform. D Diversity oriented semi synthesis. E Precursors directed and mutasynthesis
8.1 Late-stage diversification
Late-stage NP diversification through microbial biotransformation and cell-free approaches has gained interest for optimizing pharmacological properties and investigating structure–activity relationships [324].
8.1.1 Microbial biotransformation
Microbial biotransformation (Fig. 10B) employ either genetically engineered or wild strains to biosynthesize NP derivatives under mild conditions, avoiding complex purification steps low yields, and need for pure enzymes [327, 328] associated with chemical and biocatalyst [329]. Nevertheless, optimizing culture parameters (e.g. nutrient, pH, temperature) and selecting appropriate host strains [330] are utmost for maximizing productivity. These transformations effectively synthesize flavonoids [331], terpenes [332], glycosides, and vitamins analogues. For example, Huang et al. [331] employed recombinant E. coli whole cells to convert rutin into isoquercitrin, an antioxidant flavonoid glycoside. The process took place in a polyvinylidene fluoride membrane reactor that integrated reaction and separation, achieving over 80% conversion under mild conditions with efficient recyclability. Recent advances in microbial whole-cell biotransformation platforms, often integrated with semi-synthetic chemistry, feature multi-layered systems utilizing diverse wild type and engineered chassis strains. These strategies include co-cultivation of distinct microbial entities in both in vivo and in vitro setups to enhance biocatalytic efficiency [333]. Moreover, to streamline and improve scalability and productivity of NP analogues, attempts have also been made to implement continuous flow culture [334] or fed-batch culture [335], integrated of in vivo and in vitro platform [336], or micro droplets assisted synthesis [337].
In a recent study by Zhang et al. [333], developed hyper-porous hydrogel blocks for scalable cell encapsulation, ensuring nutrient access and limiting E. coli growth while supporting protein production. This enabled stable co-cultivation with Streptomyces, where encapsulated E. coli expressing RadH halogenase effectively halogenated genistein, unlike unencapsulated controls. Nevertheless, whole-cell biotransformation still faces key challenges, such as byproduct competition, toxicity of substrates, intermediates, and end product, and complex nutritional demands. The use of foreign enzymes can impose a metabolic burden, disrupting native pathways and affecting cell viability. Additionally, diverse nutritional needs and environmental conditions make co-culture systems challenging to maintain.
8.1.2 Cell-free platforms
As an alternative approach, cell-free platforms help to overcome the limitations of microbial biotransformation by enabling the NPs functionalization in a cell-free environment. Among various strategies within this framework, in vitro enzymatic methods [330] and diversity-oriented synthesis [338] have emerged as particularly effective, offering scalable and rapid NPs diversification. These methods leverage enzymes for stereo selective synthesis in a single pot, ideal for remote settings [339]. They handle toxic molecules and biosynthesize complex NPs like NRPS peptides, RIPPs peptides, cannabinoid, and limonene [340].
8.1.2.1 In-vitro enzymatic workflow
The NPs diversification using in vitro enzymatic cell-free workflows can be achieved through the use of purified enzymes [341], recombinant enzymes or crude cell extracts [342], alongside necessary cofactors, energy sources, and reactants, all integrated into a single "one-pot" reaction system as depicted in Fig. 10C. This strategy enables the biosynthesis of scaffold-specific NPs with high regio- and stereoselectivity. In particular, in vitro enzymatic cell-free workflows commonly employ redox tailoring enzymes, specialized biocatalysts that introduce oxidative modifications. Tailoring enzyme such as cytochrome P450 enzyme [343], α-ketoglutarate-dependent dioxygenases [344], flavin adenine dinucleotide or flavin mononucleotide- dependent oxygenases [345], play a pivotal role in diversifying primary and secondary metabolites through oxidative modifications, both in-vitro and in-vivo. For instance, the in vitro structural complexification of andiconin D was achieved using two α-ketoglutarate-dependent dioxygenases, SptF and SptN, which catalyzed oxidative transformations leading to the formation of the emervaridiones and emeridones[344]. Landscape of NP diversified through these methods have been described elsewhere [346, 347].
Nonetheless, it is not limited to tailoring enzymes alone; NP diversification also involves a wide array of other enzyme. For instance, Ditzel et al. [348] introduced a cell-free protein synthesis approach for constructing the NP caffeine. Their study highlighted how SAM-dependent methyltransferase reactions could be utilized within an in vitro framework to achieve partial biosynthesis. Specifically, the enzyme tea caffeine synthase was employed to catalyze caffeine production in this cell-free setup. A separate study reported the in vitro synthesis of indigoidine and rhabdopeptides using multidomain NRPSs, BpsA from S.lavendulae and KJ12ABC from Xenorhabdus KJ12.1, respectively [349].
8.1.2.2 Diversity oriented synthesis
Introduced in the early 2000s, diversity-oriented synthesis (DOS) emerged as a powerful approach for constructing structurally diverse molecular libraries for high-throughput screening. Specifically, it serves as a synthetic strategy for NP diversification, focusing on high structural complexity and molecular diversity while placing less emphasis on regio- and stereoselectivity [350, 351]. This approach enables chemists to retain the core NP scaffold while systematically modifying peripheral functionalities, stereochemical elements, or ring systems as illustrated in Fig. 10D. Remarkably, two key strategies are employed in this framework: the reagent-based approach, which modifies reagents or reaction conditions while keeping the substrate constant. For instance, use of amino acetophenones as building blocks for the synthesis of NP analogs such as 5- and 7-aminoflavones, azaflavones, 3-aryl-2-quinolones, epoxychalcones, azaaurones [352]. Other is substrate-based approach, which alters the starting substrate while keeping the reagents and conditions constant [353]. In particular, metals mediated reactions (e.g. Pd, Ni, Au, and Cu) [324] are commonly employed for selective NPs modification. Illustrative representative example of organocatlysis based diversity-oriented syntheses of 51 macrocycles with 48 unique scaffolds. By merging organocatalytic transformations with alkene metathesis, often in a one-pot setup, researchers achieved drug-like macrocycles with natural-product-like shape diversity [354].
Among several other strategic approaches, one is DOS libraries based on biosynthesis-inspired, as seen with PPAP analogues. These compounds are believed to derive from a desoxyhumulone core and two prenyl cation equivalents, which assemble the bicyclo [3.3.1] nonane scaffold via dearomative and alkene-intercepted prenylation. Leveraging this biosynthetic logic, a wide array of analogues with key bioactive structural features was synthesized [355]. In other cases, NPs-based hybrid molecules chemosynthesis, inspired by the unusual spiro-linked scaffold of indole–isatin hybrids, a DOS strategy was employed to synthesize 11 compounds, including dihydro- and tetrahydro-β-carbolines, piperidine- and pyrrolidine-fused β-carbolines, and spiropyrrolooxoindoles. Among them, two 1-aryltetrahydro-β-carbolines exhibited notable antimalarial activity [356]. Other multiple well-established DOS strategies for NP analogues discovery have been documented in the literature include diverted total synthesis, function-oriented synthesis, biology-oriented synthesis, and complexity-to-diversity. These approaches enhance chemical diversity and bioactivity, and are comprehensively reviewed elsewhere [357].
8.2 Precursor-directed biosynthesis and mutasynthesis
Precursor-directed biosynthesis and mutasynthesis are versatile strategies combining biological and chemical methods to modify NPs. Precursor-directed biosynthesis uses altered or chemically synthesized precursors (Fig. 10E) in wild or modified strains to produce NP analogues. Examples include producing alkynyl- and alkenyl-substituted erythromycin A analogues in E. coli [358] and using isotope-labeled precursors for genomic studies, such as isolating novel polyketides from P. fluorescens [359]. This approach also aids in monitoring metabolic products using techniques like stable isotope probing Raman [360] and nano secondary ion MS (e.g. Nano SIMS) [361]. In a recent study by Zhang et al. [362] used single-pot, two-stage precursor-directed biosynthesis of diverse talaroenamines by Penicillium malacosphaerulum HPU-J01. In this approach, p-methylaniline initially served as a carrier to trap the biologically synthesized cyclohexanedione, forming talaroenamine F. Subsequently, various aniline derivatives were introduced to replace the p-methylaniline moiety, yielding the final products.
Challenges like competition between synthetic and natural precursors and poor intermediate incorporation rates can complicate purification and yields. These issues can be mitigated by blocking natural precursor synthesis through mutasynthesis [363], which involves mutating or inactivating key genes or adding specific enzyme inhibitors [358]. Improvement in omics, enzymology, bioinformatics, and tools like CRISPR-Cas have improved targeted gene knockouts, enhancing mutasynthesis. In particular, modern chassis strains like Corynebacterium glutamicum, Dictyostelium discoideum, P. putida, and M. xanthus are used for mutasynthesis-driven structural diversification. Successful story of this strategy include biosynthesis of halogenated actinomycin [364], pyrrole spiroketal [365], bipyridyl collismycin A [366], amychelin siderophores [367], and isopropylstilbene [368].
9 Artificial intelligence in natural product discovery: innovations in screening, prediction, and metabolomics mining
This review has provided a concise discussion of NP strategies, yet a brief exploration of AI tools-driven discoveries in NPD would enhance its completeness. AI algorithms rapidly identify, categorize, and dereplicate compounds from complex mixtures, accelerating the discovery of novel NP, often with AI-powered web tools. The simplest variation among these tools relay on the choice of algorithms, quality of datasets for model generation, their accuracy of prediction or annotation accuracy, and computational efficiency [44]. AI has been applied across multiple facets areas of NP research, enhancing screening, pharmacological and molecular property prediction, and NP-inspired drug design. It also aids in NP target identification, deorphaning, genome and metabolomics mining, bio-/synthesis planning, classification, and structural characterization [369–371]. Among the current applications in NPD, most computational tools rely on building language models from simple rule-based systems to complex neural architectures that extract individual metabolite structures, chemical classes, or biosynthetic genes from experimental or archived datasets, such as genomic sequences or mass spectral profiles. Yet, given the structural and biosynthetic diversity of natural products, uniquely tailored models may be required to simultaneously predict multiple metabolite classes or reconstruct full biosynthetic pathways with higher fidelity [369]. NP-BGC identification largely relies on rule-based methods like AntiSMASH [29], PRISM [372] and CO-OCCUR [373], which effectively detect known BGCs but lacks in prediction of novel or unclustered pathways. Recent advances in machine learning and deep learning (DL) approaches have improved precision by recognizing unique BGC features. Tools like NPlinker [374], EFI-CGFP[375], and MAGI [376] aid biochemical enzyme or pathway predictions, while feature-based models such as BAGEL4 [377], SANDPUMA [378], and MetaMiner [379] classify metabolite types. ML-driven platforms like DeepBGC [223], GECCO [380], and SanntiS [381], outperform traditional methods in pattern recognition and scalability. Large-scale genome mining tools, BGC-Prophet [382], BiG-SLICE [39], BiG-SCAPE and CORASON [36]- enhance classification and phylogenetic analysis. Tools like decRiPPter [383], NeuRiPP [384] and DeepRiPP [385] have successfully predicted novel biosynthetic gene clusters, accelerating NP discovery. The BiG-FAM database, powered by ML, enhances gene cluster visualization and comparison [43]. ML-driven molecular networking classifier NPOmix [386] and DeepSAT [387] further refines novel NP discovery by refining MS and NMR analysis, respectively, addressing spectral reconstruction and molecular annotation challenges. Modern DL models (e.g. BiGCARP) [388], capable of predicting biosynthetic routes from NP structures, offer a foundation for matching NP to their respective BGCs. Currently landscape studies have focused on AI-guided NPD enabling the NP discovery. We emphasized these reviews to provide the reader with a deeper understanding of the subject [44, 369–371]. Additional, AI tools employed in NPD are presented in Table 1B–C, along with their respective descriptions. Furthermore, Fig. 11 presents a schematic workflow for the design of AI-inspired prediction tools in NPD, while Fig. 12 showcases the unique metabolite structures that have been mined from natural sources through AI-driven approaches.
Overview of AI-Inspired advances in natural product discovery. The schematic illustrates the core stages involved in developing AI-driven automated tools for natural product research, utilizing either experimental spectroscopic data or curated datasets
Molecular structures of selected drug candidates discovered using AI-powered techniques are shown, each labeled with its name, biological source (in parentheses), and the AI method applied (highlighted in bold)
10 Conclusions
NPs offer holistic benefits for human health and sustainability, with advantages over synthetic compounds such as diverse chemical scaffolds, strong biocompatibility, and eco-friendliness. However, their low titers under standard conditions often fall below levels required for isolation and characterization, making scalable production challenging. Recent advances in analytical tools, dereplication, and genomic techniques have improved NPs targeting, discovery and identification, while BGCs refactoring and metabolic engineering have enabled limited success in boosting and diversifying NPs yields. Nevertheless, discovery remains slow, labor-intensive, and prone to rediscovery of known compounds. Downstream processing complexity, high costs, and infrastructure demands further limit the adoption of advanced methods, leaving many labs reliant on traditional approaches. Therefore, to enable characterization-scale production of novel and complex NPs, more rational and efficient strategies for improving titers are urgently needed. Although modern sequencing and bioinformatics reveal rich biosynthetic potential in microbial genomes, less than 3% of BGCs have been experimentally characterized [389]. Unlocking silent or poorly expressed BGCs remains a major challenge due to the inherent complexities of NP research, as summarized in Table 3.
To overcome this bottleneck, next-generation automatic tools, including brain-computer interfaces and robotics, are advisable, and should focus on high-resolution chromatographic techniques, integrated omics approaches, and automated metabolomics platforms including, (1) Developing tools or protocols to activate majority of silent biosynthetic gene clusters and selecting efficient elicitors or optimization of culture conditions. (2) Developing integrated methods to isolate, sequence, and sort specific microbial strains from complex environmental samples. (3) Designing fully automated metabolomics platforms to streamline extraction, purification, structural elucidation, bioassays, and prediction of putative biological targets. (4) Creating efficient and affordable biosynthetic systems. (5) Implementing self-optimizing techniques for NPs functionalization and characterization. (6) Developing tools that correlate chemical spectral data with previously uncharacterized BGCs. Lastly, serendipity, continuous efforts, and patience remain crucial for groundbreaking discoveries and advancements in NP research.
Notes
Acknowledgements
The authors gratefully acknowledge the support from the National Key Research and Development Programs (nos. 2022YFC2804104 and 2022YFC2804700, China), the Fundamental Research Funds for the Provincial Universities of Zhejiang (no. RF-A2022013, China), the programs of the National Natural Science Foundation of China (no. 42276137, China), and Zhejiang International Sci‐Tech Cooperation Base for the Exploitation and Utilization of Nature Product.
Author contributions
Conceptualisation H. W., W.B and B. B. B.; collecting literature B. B., R. B. and Z-Y. Z.; writing – original draft, review and editing B. B. B.; visualisation R. B., Z-Y. B. and B. B. B.; supervision H. W. and W. B.; funding acquisition H. W. and W.B.
Data availability
No primary research results, software or codes have been included and no new data were generated or analysed as part of this review and examples of suitable statements you can use.
Declarations
Competing interests
There are no conflicts to declare.
References
-
1.Kuhlisch C, Pohnert G. Metabolomics in chemical ecology. Nat Prod Rep 2015;32(7): 937-55. CrossRef PubMed Google Scholar
-
2.Walsh CT, Tang Y. Natural product biosynthesis [Internet]. The Royal Society of Chemistry; 2022 [cited 2024 Jan 24]. https://books.rsc.org/books/book/2042/Natural-Product-Biosynthesis PubMed Google Scholar
-
3.Crawford JM, Tang GL, Herzon SB. Natural products: an era of discovery in organic chemistry. J Org Chem 2021;86(16): 10943-5. CrossRef PubMed Google Scholar
-
4.Kim HS, Hassan AHE, Moon K, Sim J. Natural products targeting the metabolism of amino acids: from discovery to synthetic development. Nat Prod Rep [Internet]. 2025 [cited 2025 July 21]; https://xlink.rsc.org/?DOI=D5NP00039D PubMed Google Scholar
-
5.Newman DJ, Cragg GM. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J Nat Prod 2020;83(3): 770-803. CrossRef PubMed Google Scholar
-
6.Hanzlik PJ. 125th Anniversary of the Discovery of Morphine by Sertürner**Read before the Medical History Society of the University of Oregon Medical School, Portland, Oregon, January 11, 1929. J Am Pharm Assoc (1912) 1929;18(4): 375-84. PubMed Google Scholar
-
7.Mann CC, Plummer ML. The aspirin wars: money, medicine, and 100 years of rampant competition. 1st ed. New York: Knopf; 1991. p. 420. PubMed Google Scholar
-
8.Hou X, Sun M, Bao T, Xie X, Wei F, Wang S. Recent advances in screening active components from natural products based on bioaffinity techniques. Acta Pharmaceutica Sinica B 2020;10(10): 1800-13. CrossRef PubMed Google Scholar
-
9.Clardy J, Fischbach MA, Walsh CT. New antibiotics from bacterial natural products. Nat Biotechnol 2006;24(12): 1541-50. CrossRef PubMed Google Scholar
-
10.Tong Y, Deng Z. An aurora of natural products-based drug discovery is coming. Synth Syst Biotechnol 2020;5(2): 92-6. CrossRef PubMed Google Scholar
-
11.Hutchings MI, Truman AW, Wilkinson B. Antibiotics: past, present and future. Curr Opin Microbiol 2019;51: 72-80. CrossRef PubMed Google Scholar
-
12.Ramachandraiah C, Subramaniam N, Tancer M. The story of antipsychotics: past and present. Indian J Psychiatry 2009;51(4): 324. CrossRef PubMed Google Scholar
-
13.Gach-Janczak K, Drogosz-Stachowicz J, Janecka A, Wtorek K, Mirowski M. Historical perspective and current trends in anticancer drug development. Cancers 2024;16(10): 1878. CrossRef PubMed Google Scholar
-
14.Wright GD. Opportunities for natural products in 21st century antibiotic discovery. Nat Prod Rep 2017;34(7): 694-701. CrossRef PubMed Google Scholar
-
15.Bajorath J. Integration of virtual and high-throughput screening. Nat Rev Drug Discov 2002;1(11): 882-94. CrossRef PubMed Google Scholar
-
16.Macarron R, Banks MN, Bojanic D, Burns DJ, Cirovic DA, Garyantes T, et al. Impact of high-throughput screening in biomedical research. Nat Rev Drug Discov 2011;10(3): 188-95. CrossRef PubMed Google Scholar
-
17.Gaudêncio SP, Bayram E, Lukić Bilela L, Cueto M, Díaz-Marrero AR, Haznedaroglu BZ, et al. Advanced methods for natural products discovery: bioactivity screening, dereplication, metabolomics profiling, genomic sequencing, databases and informatic tools, and structure elucidation. Mar Drugs 2023;21(5): 308. CrossRef PubMed Google Scholar
-
18.Shen B. A new golden age of natural products drug discovery. Cell 2015;163(6): 1297-300. CrossRef PubMed Google Scholar
-
19.Zhang X, Hindra EMA. Unlocking the trove of metabolic treasures: activating silent biosynthetic gene clusters in bacteria and fungi. Curr Opin Microbiol 2019;51: 9-15. CrossRef PubMed Google Scholar
-
20.Lyu C, Chen T, Qiang B, Liu N, Wang H, Zhang L, et al. CMNPD: a comprehensive marine natural products database towards facilitating drug discovery from the ocean. Nucl Acids Res 2021;49(D1): D509-15. CrossRef PubMed Google Scholar
-
21.Sorokina M, Merseburger P, Rajan K, Yirik MA, Steinbeck C. COCONUT online: collection of open natural products database. J Cheminformatics 2021;13(1): 2. CrossRef PubMed Google Scholar
-
22.van Santen JA, Jacob G, Singh AL, Aniebok V, Balunas MJ, Bunsko D, et al. The natural products atlas: an open access knowledge base for microbial natural products discovery. ACS Cent Sci 2019;5(11): 1824-33. CrossRef PubMed Google Scholar
-
23.Rutz A, Sorokina M, Galgonek J, Mietchen D, Willighagen E, Gaudry A, et al. The LOTUS initiative for open knowledge management in natural products research. Elife 2022;11: e70780. CrossRef PubMed Google Scholar
-
24.Gómez-García A, Medina-Franco JL. Progress and impact of latin american natural product databases. Biomolecules 2022;12(9): 1202. CrossRef PubMed Google Scholar
-
25.Zhao H, Yang Y, Wang S, Yang X, Zhou K, Xu C, et al. NPASS database update 2023: quantitative natural product activity and species source database for biomedical research. Nucl Acids Res 2023;51(D1): D621-8. CrossRef PubMed Google Scholar
-
26.Wishart DS, Guo A, Oler E, Wang F, Anjum A, Peters H, et al. HMDB 5.0: the human metabolome database for 2022. Nucl Acids Res 2022;50(D1): D622-31. CrossRef PubMed Google Scholar
-
27.Panagiotopoulos AA, Kalyvianaki K, Notas G, Pirintsos SA, Castanas E, Kampa M. New antagonists of the membrane androgen receptor OXER1 from the ZINC natural product database. ACS Omega 2021;6(44): 29664-74. CrossRef PubMed Google Scholar
-
28.Khaldi N, Seifuddin FT, Turner G, Haft D, Nierman WC, Wolfe KH, et al. SMURF: genomic mapping of fungal secondary metabolite clusters. Fungal Genetics Biol 2010;47(9): 736-41. CrossRef PubMed Google Scholar
-
29.Blin K, Shaw S, Augustijn HE, Reitz ZL, Biermann F, Alanjary M, et al. antiSMASH 7.0: new and improved predictions for detection, regulation, chemical structures and visualisation. Nucl Acids Res 2023;51(W1): W46-50. CrossRef PubMed Google Scholar
-
30.Weber T, Rausch C, Lopez P, Hoof I, Gaykova V, Huson DH, et al. CLUSEAN: A computer-based framework for the automated analysis of bacterial secondary metabolite biosynthetic gene clusters. J Biotechnol 2009;140(1): 13-7. CrossRef PubMed Google Scholar
-
31.Starcevic A, Zucko J, Simunkovic J, Long PF, Cullum J, Hranueli D. ClustScan : an integrated program package for the semi-automatic annotation of modular biosynthetic gene clusters and in silico prediction of novel chemical structures. Nucl Acids Res 2008;36(21): 6882-92. CrossRef PubMed Google Scholar
-
32.Anker AS, Friis-Jensen U, Johansen FL, Billinge SJL, Jensen KMØ. ClusterFinder : a fast tool to find cluster structures from pair distribution function data. Acta Crystallogr A Found Adv 2024;80(2): 213-20. CrossRef PubMed Google Scholar
-
33.Li MH, Ung PM, Zajkowski J, Garneau-Tsodikova S, Sherman DH. Automated genome mining for natural products. BMC Bioinform 2009;10(1): 185. CrossRef PubMed Google Scholar
-
34.Zierep PF, Ceci AT, Dobrusin I, Rockwell-Kollmann SC, Günther S. SeMPI 2.0—A Web Server for PKS and NRPS predictions combined with metabolite screening in natural product databases. Metabolites 2020;11(1): 13. CrossRef PubMed Google Scholar
-
35.Anand S, Prasad MVR, Yadav G, Kumar N, Shehara J, Ansari MZ, et al. SBSPKS: structure based sequence analysis of polyketide synthases. Nucl Acids Res 2010;38(2): W487-96. CrossRef PubMed Google Scholar
-
36.Navarro-Muñoz JC, Selem-Mojica N, Mullowney MW, Kautsar SA, Tryon JH, Parkinson EI, et al. A computational framework to explore large-scale biosynthetic diversity. Nat Chem Biol 2020;16(1): 60-8. CrossRef PubMed Google Scholar
-
37.Sadybekov AV, Katritch V. Computational approaches streamlining drug discovery. Nature 2023;616(7958): 673-85. CrossRef PubMed Google Scholar
-
38.Medema MH, Fischbach MA. Computational approaches to natural product discovery. Nat Chem Biol 2015;11(9): 639-48. CrossRef PubMed Google Scholar
-
39.Kautsar SA, van der Hooft JJJ, de Ridder D, Medema MH. BiG-SLiCE: a highly scalable tool maps the diversity of 1.2 million biosynthetic gene clusters. GigaScience 2021;10(1): giaa154. CrossRef PubMed Google Scholar
-
40.Zhong Z, He B, Li J, Li YX. Challenges and advances in genome mining of ribosomally synthesized and post-translationally modified peptides (RiPPs). Synth Syst Biotechnol 2020;5(3): 155-72. CrossRef PubMed Google Scholar
-
41.Atanasov AG, Zotchev SB, Dirsch VM, Supuran CT. Natural products in drug discovery: advances and opportunities. Nat Rev Drug Discov 2021;20(3): 200-16. CrossRef PubMed Google Scholar
-
42.Xu X, Liu Y, Du G, Ledesma-Amaro R, Liu L. Microbial chassis development for natural product biosynthesis. Trends Biotechnol 2020;38(7): 779-96. CrossRef PubMed Google Scholar
-
43.Kautsar SA, Blin K, Shaw S, Weber T, Medema MH. BiG-FAM: the biosynthetic gene cluster families database. Nucleic Acids Res 2021;49(D1): D490-7. CrossRef PubMed Google Scholar
-
44.Mullowney MW, Duncan KR, Elsayed SS, Garg N, Van Der Hooft JJJ, Martin NI, et al. Artificial intelligence for natural product drug discovery. Nat Rev Drug Discov 2023;22(11): 895-916. CrossRef PubMed Google Scholar
-
45.Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucl Acids Res 2016;44(D1): D457-62. CrossRef PubMed Google Scholar
-
46.Labena AA, Gao YZ, Dong C, Hua H, Guo FB. Metabolic pathway databases and model repositories. Quant Biol 2018;6(1): 30-9. CrossRef PubMed Google Scholar
-
47.Jin Z, Sato Y, Kawashima M, Kanehisa M. KEGG tools for classification and analysis of viral proteins. Protein Sci 2023;32(12): e4820. CrossRef PubMed Google Scholar
-
48.Seaver SMD, Liu F, Zhang Q, Jeffryes J. The ModelSEED Biochemistry Database for the integration of metabolic annotations and the reconstruction, comparison and analysis of metabolic models for plants, fungi and microbes. Nucl Acids Res 2020;49: D575-88. CrossRef PubMed Google Scholar
-
49.Lim PK, Julca I, Mutwil M. Redesigning plant specialized metabolism with supervised machine learning using publicly available reactome data. Comput Struct Biotechnol J 2023;1(21): 1639-50. CrossRef PubMed Google Scholar
-
50.Yuan Y, Shi C, Zhao H. Machine learning-enabled genome mining and bioactivity prediction of natural products. ACS Synthetic Biol 2023;12(9): 2650. CrossRef PubMed Google Scholar
-
51.Cunha E, Sousa V, Geada P, Teixeira JA, Vicente AA, Dias O. Systems biology's role in leveraging microalgal biomass potential: current status and future perspectives. Algal Res 2023;1(69): 102963. CrossRef PubMed Google Scholar
-
52.Prešern U, Goličnik M. Enzyme databases in the era of omics and artificial intelligence. Int J Mol Sci 2023;24(23): 16918. CrossRef PubMed Google Scholar
-
53.Karp PD, Paley S, Caspi R, Kothari A, Krummenacker M, Midford PE, et al. The EcoCyc Database (2023). EcoSal Plus 2023;11(1): eesp-0002-2023. CrossRef PubMed Google Scholar
-
54.Kumar A, Suthers PF, Maranas CD. MetRxn: a knowledgebase of metabolites and reactions spanning metabolic models and databases. BMC Bioinformatics 2012;13(1): 6. CrossRef PubMed Google Scholar
-
55.Petrovsky DV, Malsagova KA, Rudnev VR, Kulikova LI, Pustovoyt VI, Balakin EI, et al. Bioinformatics methods for constructing metabolic networks. Processes 2023;11(12): 3430. CrossRef PubMed Google Scholar
-
56.Olivier B. SystemsBioinformatics/cbmpy-metadraft: MetaDraft is now available [Internet]. [object Object]; 2018 [cited 2024 Mar 19]. https://zenodo.org/record/2398336 PubMed Google Scholar
-
57.Mendoza SN, Olivier BG, Molenaar D, Teusink B. A systematic assessment of current genome-scale metabolic reconstruction tools. Genome Biol 2019;20(1): 1-20. CrossRef PubMed Google Scholar
-
58.Kuwahara H, Alazmi M, Cui X, Gao X. MRE: a web tool to suggest foreign enzymes for the biosynthesis pathway design with competing endogenous reactions in mind. Nucleic Acids Res 2016;44: W217-25. CrossRef PubMed Google Scholar
-
59.Kanehisa M, Sato Y, Morishima K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J Mol Biol 2016;428(4): 726-31. CrossRef PubMed Google Scholar
-
60.Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, et al. The RAST server: rapid annotations using subsystems technology. BMC Genomics 2008;9(1): 75. CrossRef PubMed Google Scholar
-
61.Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucl Acids Res 2007;35(2): W182-5. CrossRef PubMed Google Scholar
-
62.Wang H, Marcišauskas S, Sánchez BJ, Domenzain I, Hermansson D, Agren R, et al. RAVEN 2.0: a versatile toolbox for metabolic network reconstruction and a case study on Streptomyces coelicolor. PLOS Comput Biol 2018;14(10): e1006541. CrossRef PubMed Google Scholar
-
63.Karp PD, Midford PE, Billington R, Kothari A, Krummenacker M, Latendresse M, et al. Pathway tools version 23.0 update: software for pathway/genome informatics and systems biology. Brief Bioinform 2021;22(1): 109-26. CrossRef PubMed Google Scholar
-
64.Mungan MD, Blin K, Ziemert N. ARTS-DB: a database for antibiotic resistant targets. Nucl Acids Res 2022;50(D1): D736-40. CrossRef PubMed Google Scholar
-
65.Alcock BP, Huynh W, Chalil R, Smith KW, Raphenya AR, Wlodarski MA, et al. CARD 2023: expanded curation, support for machine learning, and resistome prediction at the comprehensive antibiotic resistance database. Nucleic Acids Res 2023;51(D1): D690-9. CrossRef PubMed Google Scholar
-
66.Arango-Argoty G, Garner E, Pruden A, Heath LS, Vikesland P, Zhang L. DeepARG: a deep learning approach for predicting antibiotic resistance genes from metagenomic data. Microbiome 2018;6(1): 23. CrossRef PubMed Google Scholar
-
67.Gupta SK, Padmanabhan BR, Diene SM, Lopez-Rojas R, Kempf M, Landraud L, et al. ARG-ANNOT, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob Agents Chemother 2014;58(1): 212-20. CrossRef PubMed Google Scholar
-
68.Yılmaz TM, Mungan MD, Berasategui A, Ziemert N. FunARTS, the Fungal bioActive compound Resistant Target Seeker, an exploration engine for target-directed genome mining in fungi. Nucl Acids Res 2023;51(W1): W191-7. CrossRef PubMed Google Scholar
-
69.Kjærbølling I, Vesth T, Andersen MR. Resistance gene-directed genome mining of 50 Aspergillus species. mSystems 2019;4(4): e00085-e119. CrossRef PubMed Google Scholar
-
70.Pal C, Bengtsson-Palme J, Rensing C, Kristiansson E, Larsson DGJ. BacMet: antibacterial biocide and metal resistance genes database. Nucl Acids Res 2014;42(D1): D737-43. CrossRef PubMed Google Scholar
-
71.Dong H, Ming D. A comprehensive self-resistance gene database for natural-product discovery with an application to marine bacterial genome mining. IJMS 2023;24(15): 12446. CrossRef PubMed Google Scholar
-
72.Florensa AF, Kaas RS, Clausen PTLC, Aytan-Aktug D, Aarestrup FM. ResFinder–an open online resource for identification of antimicrobial resistance genes in next-generation sequencing data and prediction of phenotypes from genotypes. Microb Genom. 2022. https://doi.org/10.1099/mgen.0.000748. PubMed Google Scholar
-
73.Shafranskaya D, Chori A, Korobeynikov A. Graph-based approaches significantly improve the recovery of antibiotic resistance genes from complex metagenomic datasets. Front Microbiol 2021;6(12): 714836. CrossRef PubMed Google Scholar
-
74.Gibson MK, Forsberg KJ, Dantas G. Improved annotation of antibiotic resistance determinants reveals microbial resistomes cluster by ecology. ISME J 2015;9(1): 207-16. CrossRef PubMed Google Scholar
-
75.Doster E, Lakin SM, Dean CJ, Wolfe C, Young JG, Boucher C, et al. MEGARes 2.0: a database for classification of antimicrobial drug, biocide and metal resistance determinants in metagenomic sequence data. Nucl Acids Res 2020;48(D1): D561-9. CrossRef PubMed Google Scholar
-
76.Huo L, Hug JJ, Fu C, Bian X, Zhang Y, Müller R. Heterologous expression of bacterial natural product biosynthetic pathways. Nat Prod Rep 2019;36(10): 1412-36. CrossRef PubMed Google Scholar
-
77.Shi YM, Hirschmann M, Shi YN, Ahmed S, Abebew D, Tobias NJ, et al. Global analysis of biosynthetic gene clusters reveals conserved and unique natural products in entomopathogenic nematode-symbiotic bacteria. Nat Chem 2022;14(6): 701-12. CrossRef PubMed Google Scholar
-
78.Tay DWP, Yeo NZX, Adaikkappan K, Lim YH, Ang SJ. 67 million natural product-like compound database generated via molecular language processing. Sci Data 2023;10(1): 296. CrossRef PubMed Google Scholar
-
79.Gorgulla C, Boeszoermenyi A, Wang ZF, Fischer PD, Coote PW, Padmanabha Das KM, et al. An open-source drug discovery platform enables ultra-large virtual screens. Nature 2020;580(7805): 663-8. CrossRef PubMed Google Scholar
-
80.Zampaloni C, Mattei P, Bleicher K, Winther L, Thäte C, Bucher C, et al. A novel antibiotic class targeting the lipopolysaccharide transporter. Nature 2024;625(7995): 566-71. CrossRef PubMed Google Scholar
-
81.Subramanian A, Narayan R, Corsello SM, Peck DD, Natoli TE, Lu X, et al. A next generation connectivity map: L1000 platform and the first 1, 000, 000 profiles. Cell 2017;171(6): 1437-1452.e17. CrossRef PubMed Google Scholar
-
82.Bode HB, Bethe B, Höfs R, Zeeck A. Big effects from small changes: possible ways to explore nature's chemical diversity. ChemBioChem 2002;3(7): 619. CrossRef PubMed Google Scholar
-
83.Seyedsayamdost MR. High-throughput platform for the discovery of elicitors of silent bacterial gene clusters. Proc Natl Acad Sci USA 2014;111(20): 7266-71. CrossRef PubMed Google Scholar
-
84.Romano S, Jackson S, Patry S, Dobson A. Extending the "One Strain Many Compounds" (OSMAC) principle to marine microorganisms. Mar Drugs 2018;16(7): 244. CrossRef PubMed Google Scholar
-
85.Hebra T, Pollet N, Touboul D, Eparvier V. Combining OSMAC, metabolomic and genomic methods for the production and annotation of halogenated azaphilones and ilicicolins in termite symbiotic fungi. Sci Rep 2022;12(1): 17310. CrossRef PubMed Google Scholar
-
86.Wang QX, Bao L, Yang XL, Guo H, Ren B, Guo LD, et al. Tricycloalternarenes F-H: Three new mixed terpenoids produced by an endolichenic fungus Ulocladium sp. using OSMAC method. Fitoterapia 2013;85: 8-13. CrossRef PubMed Google Scholar
-
87.Armin R, Zühlke S, Mahnkopp-Dirks F, Winkelmann T, Kusari S. Evaluation of apple root-associated endophytic streptomyces pulveraceus strain ES16 by an OSMAC-assisted metabolomics approach. Front Sustain Food Syst 2021;23(5): 643225. CrossRef PubMed Google Scholar
-
88.Zhang Y, Feng L, Hemu X, Tan NH, Wang Z. OSMAC strategy: a promising way to explore microbial cyclic peptides. Eur J Med Chem 2024;268: 116175. CrossRef PubMed Google Scholar
-
89.Zhu W, Zhang H. Enhancing chemical diversity of fungal secondary metabolite by OSMAC strategy. In: Deshmukh SK, Takahashi JA, Saxena S, editors. Fungi bioactive metabolites. Singapore: Springer Nature Singapore; 2024. p. 567–604. https://doi.org/10.1007/978-981-99-5696-8_18. PubMed Google Scholar
-
90.Wei X, Chan TK, Kong CTD, Matsuda Y. Biosynthetic characterization, heterologous production, and genomics-guided discovery of GABA-containing fungal heptapeptides. J Nat Prod 2023;86(2): 416-22. CrossRef PubMed Google Scholar
-
91.Zang Y, Gong Y, Gong J, Liu J, Chen C, Gu L, et al. Fungal polyketides with three distinctive ring skeletons from the fungus Penicillium canescens uncovered by OSMAC and molecular networking strategies. J Org Chem 2020;85(7): 4973-80. CrossRef PubMed Google Scholar
-
92.Palma Esposito F, Giugliano R, Della Sala G, Vitale GA, Buonocore C, Ausuri J, et al. Combining OSMAC approach and untargeted metabolomics for the identification of new glycolipids with potent antiviral activity produced by a marine rhodococcus. IJMS 2021;22(16): 9055. CrossRef PubMed Google Scholar
-
93.Wei Q, Bai J, Yan D, Bao X, Li W, Liu B, et al. Genome mining combined metabolic shunting and OSMAC strategy of an endophytic fungus leads to the production of diverse natural products. Acta Pharmaceutica Sinica B 2021;11(2): 572-87. CrossRef PubMed Google Scholar
-
94.Wu J, Wang Y, Wang Y, Li X, Li Y, Zhang M, et al. A combination of genome mining with OSMAC strategy facilitates the discovery of bioactive metabolites produced from termite-associated Streptomyces tanashiensis BYF -112. Pest Manag Sci 2025;81(4): 2364-78. CrossRef PubMed Google Scholar
-
95.Mittermeier F, Bäumler M, Arulrajah P, García Lima JDJ, Hauke S, Stock A, et al. Artificial microbial consortia for bioproduction processes. Eng Life Sci 2023;23(1): e2100152. CrossRef PubMed Google Scholar
-
96.Bader J, Mast-Gerlach E, Popović MK, Bajpai R, Stahl U. Relevance of microbial coculture fermentations in biotechnology. J Appl Microbiol 2010;109(2): 371-87. CrossRef PubMed Google Scholar
-
97.Li YZ, Zhang WQ, Hu PF, Yang QQ, Molnár I, Xu P, et al. Harnessing microbial co-culture to increase the production of known secondary metabolites. Nat Prod Rep 2025;42(3): 623-37. CrossRef PubMed Google Scholar
-
98.Peng XY, Wu JT, Shao CL, Li ZY, Chen M, Wang CY. Co-culture: stimulate the metabolic potential and explore the molecular diversity of natural products from microorganisms. Mar Life Sci Technol 2021;3(3): 363-74. CrossRef PubMed Google Scholar
-
99.Guo X, Li Z, Wang X, Wang J, Chala J, Lu Y, et al. De novo phenol bioproduction from glucose using biosensor-assisted microbial coculture engineering. Biotech Bioeng 2019;116(12): 3349-59. CrossRef PubMed Google Scholar
-
100.Lalwani MA, Kawabe H, Mays RL, Hoffman SM, Avalos JL. Optogenetic control of microbial consortia populations for chemical production. ACS Synth Biol 2021;10(8): 2015-29. CrossRef PubMed Google Scholar
-
101.Johnston TG, Yuan SF, Wagner JM, Yi X, Saha A, Smith P, et al. Compartmentalized microbes and co-cultures in hydrogels for on-demand bioproduction and preservation. Nat Commun 2020;11(1): 563. CrossRef PubMed Google Scholar
-
102.Gonzalez DJ, Haste NM, Hollands A, Fleming TC, Hamby M, Pogliano K, et al. Microbial competition between Bacillus subtilis and Staphylococcus aureus monitored by imaging mass spectrometry. Microbiology 2011;157(9): 2485-92. CrossRef PubMed Google Scholar
-
103.Moon K, Xu F, Seyedsayamdost MR. Cebulantin, a cryptic lanthipeptide antibiotic uncovered using bioactivity-coupled HiTES. Angew Chem Int Ed 2019;58(18): 5973-7. CrossRef PubMed Google Scholar
-
104.Chen T, Zhou Y, Lu Y, Zhang H. Advances in heterologous biosynthesis of plant and fungal natural products by modular co-culture engineering. Biotechnol Lett 2019;41(1): 27-34. CrossRef PubMed Google Scholar
-
105.Wang R, Zhao S, Wang Z, Koffas MA. Recent advances in modular co-culture engineering for synthesis of natural products. Curr Opin Biotechnol 2020;62: 65-71. CrossRef PubMed Google Scholar
-
106.Marsafari M, Azi F, Dou S, Xu P. Modular co-culture engineering of Yarrowia lipolytica for amorphadiene biosynthesis. Microb Cell Fact. 2022. https://doi.org/10.1186/s12934-022-02010-0. PubMed Google Scholar
-
107.Okada BK, Wu Y, Mao D, Bushin LB, Seyedsayamdost MR. Mapping the trimethoprim-induced secondary metabolome of Burkholderia thailandensis. ACS Chem Biol 2016;11(8): 2124-30. CrossRef PubMed Google Scholar
-
108.Xu F, Nazari B, Moon K, Bushin LB, Seyedsayamdost MR. Discovery of a cryptic antifungal compound from Streptomyces albus J1074 using high-throughput elicitor screens. J Am Chem Soc 2017;139(27): 9203-12. CrossRef PubMed Google Scholar
-
109.Aron AT, Gentry EC, McPhail KL, Nothias LF, Nothias-Esposito M, Bouslimani A, et al. Reproducible molecular networking of untargeted mass spectrometry data using GNPS. Nat Protoc 2020;15(6): 1954-91. CrossRef PubMed Google Scholar
-
110.Dührkop K, Fleischauer M, Ludwig M, Aksenov AA, Melnik AV, Meusel M, et al. SIRIUS 4: a rapid tool for turning tandem mass spectra into metabolite structure information. Nat Methods 2019;16(4): 299-302. CrossRef PubMed Google Scholar
-
111.Wang M, Carver JJ, Phelan VV, Sanchez LM, Garg N, Peng Y, et al. Sharing and community curation of mass spectrometry data with global natural products social molecular networking. Nat Biotechnol 2016;34(8): 828-37. CrossRef PubMed Google Scholar
-
112.Fox Ramos AE, Evanno L, Poupon E, Champy P, Beniddir MA. Natural products targeting strategies involving molecular networking: different manners, one goal. Nat Prod Rep 2019;36(7): 960-80. CrossRef PubMed Google Scholar
-
113.Qin GF, Zhang X, Zhu F, Huo ZQ, Yao QQ, Feng Q, et al. MS/MS-based molecular networking: an efficient approach for natural products dereplication. Molecules 2022;28(1): 157. CrossRef PubMed Google Scholar
-
114.Nothias LF, Petras D, Schmid R, Dührkop K, Rainer J, Sarvepalli A, et al. Feature-based molecular networking in the GNPS analysis environment. Nat Methods 2020;17(9): 905-8. CrossRef PubMed Google Scholar
-
115.Schmid R, Petras D, Nothias LF, Wang M, Aron AT, Jagels A, et al. Ion identity molecular networking for mass spectrometry-based metabolomics in the GNPS environment. Nat Commun 2021;12(1): 3832. CrossRef PubMed Google Scholar
-
116.He Q, Wu Z, Li L, Sun W, Wang G, Jiang R, et al. Discovery of Neuritogenic securinega alkaloids from Flueggea suffruticosa by a building blocks-based molecular network strategy. Angew Chem Int Ed 2021;60(36): 19609-13. CrossRef PubMed Google Scholar
-
117.Van Der Hooft JJJ, Wandy J, Barrett MP, Burgess KEV, Rogers S. Topic modeling for untargeted substructure exploration in metabolomics. Proc Natl Acad Sci USA 2016;113(48): 13738-43. CrossRef PubMed Google Scholar
-
118.Van Der Hooft JJJ, Wandy J, Young F, Padmanabhan S, Gerasimidis K, Burgess KEV, et al. Unsupervised discovery and comparison of structural families across multiple samples in untargeted metabolomics. Anal Chem 2017;89(14): 7569-77. CrossRef PubMed Google Scholar
-
119.Nothias LF, Nothias-Esposito M, Da Silva R, Wang M, Protsyuk I, Zhang Z, et al. Bioactivity-based molecular networking for the discovery of drug leads in natural product bioassay-guided fractionation. J Nat Prod 2018;81(4): 758-67. CrossRef PubMed Google Scholar
-
120.Chanana S, Thomas CS, Zhang F, Rajski SR, Bugni TS. hcapca: automated hierarchical clustering and principal component analysis of large metabolomic datasets in R. Metabolites 2020;10(7): 297. CrossRef PubMed Google Scholar
-
121.Covington BC, Seyedsayamdost MR. MetEx, a metabolomics explorer application for natural product discovery. ACS Chem Biol 2021;16(12): 2825-33. CrossRef PubMed Google Scholar
-
122.Zhang C, Seyedsayamdost MR. Discovery of a cryptic depsipeptide from Streptomyces ghanaensis via MALDI-MS-guided high-throughput elicitor screening. Angew Chem Int Ed 2020;59(51): 23005-9. CrossRef PubMed Google Scholar
-
123.Moon K, Xu F, Zhang C, Seyedsayamdost MR. Bioactivity-HiTES unveils cryptic antibiotics encoded in actinomycete bacteria. ACS Chem Biol 2019;14(4): 767-74. CrossRef PubMed Google Scholar
-
124.Xu F, Wu Y, Zhang C, Davis KM, Moon K, Bushin LB, et al. A genetics-free method for high-throughput discovery of cryptic microbial metabolites. Nat Chem Biol 2019;15(2): 161-8. CrossRef PubMed Google Scholar
-
125.Han EJ, Lee SR, Townsend CA, Seyedsayamdost MR. Targeted discovery of cryptic enediyne natural products via FRET-coupled high-throughput elicitor screening. ACS Chem Biol 2023;18(8): 1854-62. CrossRef PubMed Google Scholar
-
126.Khalid S, Keller NP. Chemical signals driving bacterial–fungal interactions. Environ Microbiol 2021;23(3): 1334-47. CrossRef PubMed Google Scholar
-
127.Weinberg ED. Roles of trace metals in transcriptional control of microbial secondary metabolism. Biol Metals 1990;2(4): 191-6. CrossRef PubMed Google Scholar
-
128.Giri CC, Zaheer M. Chemical elicitors versus secondary metabolite production in vitro using plant cell, tissue and organ cultures: recent trends and a sky eye view appraisal. Plant Cell Tiss Organ Cult 2016;126(1): 1-18. CrossRef PubMed Google Scholar
-
129.Tanaka Y, Hosaka T, Ochi K. Rare earth elements activate the secondary metabolite–biosynthetic gene clusters in Streptomyces coelicolor A3(2). J Antibiot 2010;63(8): 477-81. CrossRef PubMed Google Scholar
-
130.Shentu XP, Cao ZY, Xiao Y, Tang G, Ochi K, Yu XP. Substantial improvement of toyocamycin production in Streptomyces diastatochromogenes by cumulative drug-resistance mutations. Virolle MJ, editor. PLoS ONE. 2018 Aug 30;13(8):e0203006. PubMed Google Scholar
-
131.Xu D, Han L, Li C, Cao Q, Zhu D, Barrett NH, et al. Bioprospecting deep-sea actinobacteria for novel anti-infective natural products. Front Microbiol 2018;30(9): 787. CrossRef PubMed Google Scholar
-
132.Paranagama PA, Wijeratne EMK, Gunatilaka AAL. Uncovering biosynthetic potential of plant-associated fungi: effect of culture conditions on metabolite production by Paraphaeosphaeria quadriseptata and Chaetomium chiversii. J Nat Prod 2007;70(12): 1939-45. CrossRef PubMed Google Scholar
-
133.Hassan SSU, Muhammad I, Abbas SQ, Hassan M, Majid M, Jin HZ, et al. Stress driven discovery of natural products from actinobacteria with anti-oxidant and cytotoxic activities including docking and ADMET properties. IJMS 2021;22(21): 11432. CrossRef PubMed Google Scholar
-
134.Wang C, Huang D, Liang S. Identification and metabolomic analysis of chemical elicitors for tacrolimus accumulation in Streptomyces tsukubaensis. Appl Microbiol Biotechnol 2018;102(17): 7541-53. CrossRef PubMed Google Scholar
-
135.Akhter N, Liu Y, Auckloo B, Shi Y, Wang K, Chen J, et al. Stress-driven discovery of new angucycline-type antibiotics from a marine streptomyces pratensis NA-ZhouS1. Mar Drugs 2018;16(9): 331. CrossRef PubMed Google Scholar
-
136.Wang L, Li Y, Li Y. Metal ions driven production, characterization and bioactivity of extracellular melanin from Streptomyces sp. ZL-24. Int J Biol Macromol 2019;123: 521-30. CrossRef PubMed Google Scholar
-
137.Krishnan N, Singh PK, Devadasan V, Mariappanadar V, Gopinath SCB, Chinni SV, et al. Enhanced production of actinidine and glaziovine alkaloids from Nardostachys jatamansi (D. Don) DC. through cell suspension culture with elicitors treatment. Process Biochem. 2024;138:139–49. PubMed Google Scholar
-
138.Buraphaka H, Putalun W. Stimulation of health-promoting triterpenoids accumulation in Centella asiatica (L.) Urban leaves triggered by postharvest application of methyl jasmonate and salicylic acid elicitors. Ind Crops Prod 2020;146: 112171. CrossRef PubMed Google Scholar
-
139.Daniel-Ivad M, Pimentel-Elardo S, Nodwell JR. Control of specialized metabolism by signaling and transcriptional regulation: opportunities for new platforms for drug discovery. Annu Rev Microbiol 2018;72(1): 25-48. CrossRef PubMed Google Scholar
-
140.Chen G, Wang GYS, Li X, Waters B, Davies J. Enhanced production of microbial metabolites in the presence of dimethyl sulfoxide. J Antibiot 2000;53(10): 1145-53. CrossRef PubMed Google Scholar
-
141.Sekurova ON, Zhang J, Kristiansen KA, Zotchev SB. Activation of chloramphenicol biosynthesis in Streptomyces venezuelae ATCC 10712 by ethanol shock: insights from the promoter fusion studies. Microb Cell Fact 2016;15(1): 85. CrossRef PubMed Google Scholar
-
142.Zhou WW, Ma B, Tang YJ, Zhong JJ, Zheng X. Enhancement of validamycin A production by addition of ethanol in fermentation of Streptomyces hygroscopicus 5008. Biores Technol 2012;114: 616-21. CrossRef PubMed Google Scholar
-
143.Wei ZH, Bai L, Deng Z, Zhong JJ. Enhanced production of validamycin A by H2O2-induced reactive oxygen species in fermentation of Streptomyces hygroscopicus 5008. Biores Technol 2011;102(2): 1783-7. CrossRef PubMed Google Scholar
-
144.Xu J, Bao JW, Su XF, Zhang HJ, Zeng X, Tang L, et al. Effect of propionic acid on citric acid fermentation in an integrated citric acid–methane fermentation process. Bioprocess Biosyst Eng 2016;39(3): 391-400. CrossRef PubMed Google Scholar
-
145.Bhosale P. Environmental and cultural stimulants in the production of carotenoids from microorganisms. Appl Microbiol Biotechnol 2004;63(4): 351-61. CrossRef PubMed Google Scholar
-
146.Van Arnam EB, Ruzzini AC, Sit CS, Horn H, Pinto-Tomás AA, Currie CR, et al. Selvamicin, an atypical antifungal polyene from two alternative genomic contexts. Proc Natl Acad Sci USA 2016;113(46): 12940-5. CrossRef PubMed Google Scholar
-
147.Moore JM, Bradshaw E, Seipke RF, Hutchings MI, McArthur M. Use and discovery of chemical elicitors that stimulate biosynthetic gene clusters in streptomyces bacteria. In: Methods in Enzymology. Elsevier; 2012. p. 367–85. PubMed Google Scholar
-
148.Zutz C, Gacek A, Sulyok M, Wagner M, Strauss J, Rychli K. Small chemical chromatin effectors alter secondary metabolite production in Aspergillus clavatus. Toxins 2013;5(10): 1723-41. CrossRef PubMed Google Scholar
-
149.Asai T, Morita S, Taniguchi T, Monde K, Oshima Y. Epigenetic stimulation of polyketide production in Chaetomium cancroideum by an NAD + -dependent HDAC inhibitor. Org Biomol Chem 2016;14(2): 646-51. CrossRef PubMed Google Scholar
-
150.Pettit RK. Small-molecule elicitation of microbial secondary metabolites. Microb Biotechnol 2011;4(4): 471-8. CrossRef PubMed Google Scholar
-
151.Liang CX, Li YB, Xu JW, Wang JL, Miao XL, Tang YJ, et al. Enhanced biosynthetic gene expressions and production of ganoderic acids in static liquid culture of Ganoderma lucidum under phenobarbital induction. Appl Microbiol Biotechnol 2010;86(5): 1367-74. CrossRef PubMed Google Scholar
-
152.Bode HB, Zeeck A. Sphaerolone and dihydrosphaerolone, two bisnaphthyl-pigments from the fungus Sphaeropsidales sp. F-24′707. Phytochemistry 2000;54(6): 597-601. CrossRef PubMed Google Scholar
-
153.Williams RB, Henrikson JC, Hoover AR, Lee AE, Cichewicz RH. Epigenetic remodeling of the fungal secondary metabolome. Org Biomol Chem 2008;6(11): 1895. CrossRef PubMed Google Scholar
-
154.Tawfike A, Attia EZ, Desoukey SY, Hajjar D, Makki AA, Schupp PJ, et al. New bioactive metabolites from the elicited marine sponge-derived bacterium Actinokineospora spheciospongiae sp. nov. AMB Expr 2019;9(1): 12. CrossRef PubMed Google Scholar
-
155.Kahromi S, Khara J. Chitosan stimulates secondary metabolite production and nutrient uptake in medicinal plant Dracocephalum kotschyi. J Sci Food Agric 2021;101(9): 3898-907. CrossRef PubMed Google Scholar
-
156.Liu X, Tang J, Wang L, Liu R. Mechanism of CuO nano-particles on stimulating production of actinorhodin in Streptomyces coelicolor by transcriptional analysis. Sci Rep 2019;9(1): 11253. CrossRef PubMed Google Scholar
-
157.De Felício R, Ballone P, Bazzano CF, Alves LFG, Sigrist R, Infante GP, et al. Chemical elicitors induce rare bioactive secondary metabolites in deep-sea bacteria under laboratory conditions. Metabolites 2021;11(2): 107. CrossRef PubMed Google Scholar
-
158.Lee N, Kim W, Chung J, Lee Y, Cho S, Jang KS, et al. Iron competition triggers antibiotic biosynthesis in Streptomyces coelicolor during coculture with Myxococcus xanthus. ISME J 2020;14(5): 1111-24. CrossRef PubMed Google Scholar
-
159.Tanaka Y, Izawa M, Hiraga Y, Misaki Y, Watanabe T, Ochi K. Metabolic perturbation to enhance polyketide and nonribosomal peptide antibiotic production using triclosan and ribosome-targeting drugs. Appl Microbiol Biotechnol 2017;101(11): 4417-31. CrossRef PubMed Google Scholar
-
160.Amano S, Sakurai T, Endo K, Takano H, Beppu T, Furihata K, et al. A cryptic antibiotic triggered by monensin. J Antibiot 2011;64(10): 703-703. CrossRef PubMed Google Scholar
-
161.Pimentel-Elardo SM, Gulder TAM, Hentschel U, Bringmann G. Cebulactams A1 and A2, new macrolactams isolated from Saccharopolyspora cebuensis, the first obligate marine strain of the genus Saccharopolyspora. Tetrahedron Lett 2008;49(48): 6889-92. CrossRef PubMed Google Scholar
-
162.Timmermans ML, Picott KJ, Ucciferri L, Ross AC. Culturing marine bacteria from the genus Pseudoalteromonas on a cotton scaffold alters secondary metabolite production. MicrobiologyOpen 2019;8(5): e00724. CrossRef PubMed Google Scholar
-
163.Gonciarz J, Bizukojc M. Adding talc microparticles to a spergillus terreus ATCC 20542 preculture decreases fungal pellet size and improves lovastatin production. Eng Life Sci 2014;14(2): 190-200. CrossRef PubMed Google Scholar
-
164.Kuhl M, Rückert C, Gläser L, Beganovic S, Luzhetskyy A, Kalinowski J, et al. Microparticles enhance the formation of seven major classes of natural products in native and metabolically engineered actinobacteria through accelerated morphological development. Biotech Bioeng 2021;118(8): 3076-93. CrossRef PubMed Google Scholar
-
165.Staley JT, Konopka A. Measurement of in situ activities of nonphotosynthetic microorganisms in aquatic and terrestrial habitats. Annu Rev Microbiol 1985;39(1): 321-46. CrossRef PubMed Google Scholar
-
166.Lloyd KG, Steen AD, Ladau J, Yin J, Crosby L. Phylogenetically novel uncultured microbial cells dominate earth microbiomes. mSystems 2018;3(5): e00055-e118. CrossRef PubMed Google Scholar
-
167.Metabolic Constraints and Dependencies Between "Uncultivable" Fungi and Their Hosts. In: The Mycota [Internet]. Cham: Springer International Publishing; 2024; p. 33–57. https://doi.org/10.1007/978-3-031-41648-4_2 PubMed Google Scholar
-
168.Shi H, Wang Y, Zhang Z, Yu S, Huang X, Pan D, et al. Recent advances of integrated microfluidic systems for fungal and bacterial analysis. TrAC Trends Anal Chem 2023;158: 116850. CrossRef PubMed Google Scholar
-
169.Liang J, Arellano A, Sajjadi S, Jewell T. Cultivation of diverse and rare bacteria from the human gut microbiome: rapid isolation of entire populations of microbes using the GALT prospector, an automated array-based platform. Genet Eng Biotechnol News 2020;40(7): 50-2. CrossRef PubMed Google Scholar
-
170.Geesink P, Tyc O, Küsel K, Taubert M, van de Velde C, Kumar S, et al. Growth promotion and inhibition induced by interactions of groundwater bacteria. FEMS Microbiol Ecol. 2018. https://doi.org/10.1093/femsec/fiy164/5076029. PubMed Google Scholar
-
171.Watrous JD, Dorrestein PC. Imaging mass spectrometry in microbiology. Nat Rev Microbiol 2011;9(9): 683-94. CrossRef PubMed Google Scholar
-
172.Kankanamge S, Bernhardt PV, Khalil ZG, Capon RJ. Miniaturized cultivation profiling (MATRIX)-facilitated discovery of noonazines A-C and noonaphilone A from an Australian marine-derived fungus, Aspergillus noonimiae CMB-M0339. Mar Drugs 2024;22(6): 243. CrossRef PubMed Google Scholar
-
173.Jin S, Bruhn DF, Childs CT, Burkman E, Moreno Y, Salim AA, et al. Goondapyrones A-J: polyketide α and γ pyrone anthelmintics from an australian soil-derived Streptomyces sp. Antibiotics 2024;13(10): 989. CrossRef PubMed Google Scholar
-
174.Salcher MM, Layoun P, Fernandes C, Chiriac MC, Bulzu PA, Ghai R, et al. Bringing the uncultivated microbial majority of freshwater ecosystems into culture. Nat Commun 2025;16(1): 7971. CrossRef PubMed Google Scholar
-
175.Jing J, Garbeva P, Raaijmakers JM, Medema MH. Strategies for tailoring functional microbial synthetic communities. ISME J 2024;18(1): wrae049. CrossRef PubMed Google Scholar
-
176.De La Torre JR, Christianson LM, Béjà O, Suzuki MT, Karl DM, Heidelberg J, et al. Proteorhodopsin genes are distributed among divergent marine bacterial taxa. Proc Natl Acad Sci USA 2003;100(22): 12830-5. CrossRef PubMed Google Scholar
-
177.Sabehi G, Loy A, Jung KH, Partha R, Spudich JL, Isaacson T, et al. New insights into metabolic properties of marine bacteria encoding proteorhodopsins. PLoS Biol 2005;3(8): e273. CrossRef PubMed Google Scholar
-
178.Lewis WH, Tahon G, Geesink P, Sousa DZ, Ettema TJG. Innovations to culturing the uncultured microbial majority. Nat Rev Microbiol 2021;19(4): 225-40. CrossRef PubMed Google Scholar
-
179.Chen CH, Cho SH, Chiang HI, Tsai F, Zhang K, Lo YH. Specific sorting of single bacterial cells with microfabricated fluorescence-activated cell sorting and tyramide signal amplification fluorescence in situ hybridization. Anal Chem 2011;83(19): 7269-75. CrossRef PubMed Google Scholar
-
180.Wang Y, Ji Y, Wharfe ES, Meadows RS, March P, Goodacre R, et al. Raman activated cell ejection for isolation of single cells. Anal Chem 2013;85(22): 10697-701. CrossRef PubMed Google Scholar
-
181.Ling LL, Schneider T, Peoples AJ, Spoering AL, Engels I, Conlon BP, et al. A new antibiotic kills pathogens without detectable resistance. Nature 2015;517(7535): 455-9. CrossRef PubMed Google Scholar
-
182.Hofer U. The majority is uncultured. Nat Rev Microbiol 2018;16(12): 716-7. CrossRef PubMed Google Scholar
-
183.Dror B, Wang Z, Brady SF, Jurkevitch E, Cytryn E. Elucidating the diversity and potential function of nonribosomal peptide and polyketide biosynthetic gene clusters in the root microbiome. mSystems. 2020. https://doi.org/10.1128/mSystems.00866-20. PubMed Google Scholar
-
184.Goodfellow M, Nouioui I, Sanderson R, Xie F, Bull AT. Rare taxa and dark microbial matter: novel bioactive actinobacteria abound in Atacama Desert soils. Antonie Van Leeuwenhoek 2018;111(8): 1315-32. CrossRef PubMed Google Scholar
-
185.Liu H, Xue R, Wang Y, Stirling E, Ye S, Xu J, et al. FACS-iChip: a high-efficiency iChip system for microbial 'dark matter' mining. Mar Life Sci Technol 2021;3(2): 162-8. CrossRef PubMed Google Scholar
-
186.Nayfach S, Roux S, Seshadri R, Udwary D, Varghese N, Schulz F, et al. A genomic catalog of Earth's microbiomes. Nat Biotechnol 2021;39(4): 499-509. CrossRef PubMed Google Scholar
-
187.Mukherjee A, Tikariha H, Bandla A, Pavagadhi S, Swarup S. Global analyses of biosynthetic gene clusters in phytobiomes reveal strong phylogenetic conservation of terpenes and aryl polyenes. mSystems. 2023. https://doi.org/10.1128/msystems.00387-23. PubMed Google Scholar
-
188.Lee N, Hwang S, Kim J, Cho S, Palsson B, Cho BK. Mini review: Genome mining approaches for the identification of secondary metabolite biosynthetic gene clusters in Streptomyces. Comput Struct Biotechnol J 2020;18: 1548-56. CrossRef PubMed Google Scholar
-
189.Ochi K. Insights into microbial cryptic gene activation and strain improvement: principle, application and technical aspects. J Antibiot 2017;70(1): 25-40. CrossRef PubMed Google Scholar
-
190.Satam H, Joshi K, Mangrolia U, Waghoo S, Zaidi G, Rawool S, et al. Next-generation sequencing technology: current trends and advancements. Biology 2023;12(7): 997. CrossRef PubMed Google Scholar
-
191.Scherlach K, Hertweck C. Triggering cryptic natural product biosynthesis in microorganisms. Org Biomol Chem 2009;7(9): 1753. CrossRef PubMed Google Scholar
-
192.Alam K, Islam MM, Gong K, Abbasi MN, Li R, Zhang Y, et al. In silico genome mining of potential novel biosynthetic gene clusters for drug discovery from Burkholderia bacteria. Comput Biol Med 2022;140: 105046. CrossRef PubMed Google Scholar
-
193.Ziemert N, Alanjary M, Weber T. The evolution of genome mining in microbes-a review. Nat Prod Rep 2016;33(8): 988-1005. CrossRef PubMed Google Scholar
-
194.Ibrahim A, Yang L, Johnston C, Liu X, Ma B, Magarvey NA. Dereplicating nonribosomal peptides using an informatic search algorithm for natural products (iSNAP) discovery. Proc Natl Acad Sci USA 2012;109(47): 19196-201. CrossRef PubMed Google Scholar
-
195.Kim LJ, Ohashi M, Zhang Z, Tan D, Asay M, Cascio D, et al. Prospecting for natural products by genome mining and microcrystal electron diffraction. Nat Chem Biol 2021;17(8): 872-7. CrossRef PubMed Google Scholar
-
196.Bauman KD, Butler KS, Moore BS, Chekan JR. Genome mining methods to discover bioactive natural products. Nat Prod Rep 2021;38(11): 2100-29. CrossRef PubMed Google Scholar
-
197.Du R, Xiong W, Xu L, Xu Y, Wu Q. Metagenomics reveals the habitat specificity of biosynthetic potential of secondary metabolites in global food fermentations. Microbiome 2023;11(1): 115. CrossRef PubMed Google Scholar
-
198.Zhang MM, Qiao Y, Ang EL, Zhao H. Using natural products for drug discovery: the impact of the genomics era. Expert Opin Drug Discov 2017;12(5): 475-87. CrossRef PubMed Google Scholar
-
199.Engel K, Ashby D, Brady SF, Cowan DA, Doemer J, Edwards EA, et al. Meeting report: 1st international functional metagenomics workshop May 7–8, 2012, St. Jacobs, Ontario, Canada. Stand Genomic Sci 2013;8(1): 106-11. PubMed Google Scholar
-
200.Cahn JKB, Piel J. Opening up the single-cell toolbox for microbial natural products research. Angew Chem Int Ed 2021;60(34): 18412-28. CrossRef PubMed Google Scholar
-
201.Chang FY, Ternei MA, Calle PY, Brady SF. Targeted metagenomics: finding rare tryptophan dimer natural products in the environment. J Am Chem Soc 2015;137(18): 6044-52. CrossRef PubMed Google Scholar
-
202.Robinson SL, Piel J, Sunagawa S. A roadmap for metagenomic enzyme discovery. Nat Prod Rep 2021;38(11): 1994-2023. CrossRef PubMed Google Scholar
-
203.The MetaSUB International Consortium. The Metagenomics and Metadesign of the Subways and Urban Biomes (MetaSUB) International Consortium inaugural meeting report. Microbiome 2016;4(1): 24. CrossRef PubMed Google Scholar
-
204.MetaHIT Consortium, Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 2010;464(7285): 59-65. CrossRef PubMed Google Scholar
-
205.Siegl A, Hentschel U. PKS and NRPS gene clusters from microbial symbiont cells of marine sponges by whole genome amplification. Environ Microbiol Rep 2010;2(4): 507-13. CrossRef PubMed Google Scholar
-
206.Song Y, Yin J, Huang WE, Li B, Yin H. Emerging single-cell microfluidic technology for microbiology. TrAC Trends Anal Chem 2024;170: 117444. CrossRef PubMed Google Scholar
-
207.Xu Y, Zhao F. Single-cell metagenomics: challenges and applications. Protein Cell 2018;9(5): 501-10. CrossRef PubMed Google Scholar
-
208.Larsson DGJ, Flach CF. Antibiotic resistance in the environment. Nat Rev Microbiol 2022;20(5): 257-69. CrossRef PubMed Google Scholar
-
209.Hobson C, Chan AN, Wright GD. The antibiotic resistome: a guide for the discovery of natural products as antimicrobial agents. Chem Rev 2021;121(6): 3464-94. CrossRef PubMed Google Scholar
-
210.Panter F, Krug D, Baumann S, Müller R. Self-resistance guided genome mining uncovers new topoisomerase inhibitors from myxobacteria. Chem Sci 2018;9(21): 4898-908. CrossRef PubMed Google Scholar
-
211.Chase AB, Sweeney D, Muskat MN, Guillén-Matus DG, Jensen PR. Vertical inheritance facilitates interspecies diversification in biosynthetic gene clusters and specialized metabolites. MBio. 2021. https://doi.org/10.1128/mBio.02700-21. PubMed Google Scholar
-
212.Moffitt MC, Neilan BA. Evolutionary affiliations within the superfamily of ketosynthases reflect complex pathway associations. J Mol Evol 2003;56(4): 446-57. CrossRef PubMed Google Scholar
-
213.Chevrette MG, Gutiérrez-García K, Selem-Mojica N, Aguilar-Martínez C, Yañez-Olvera A, Ramos-Aboites HE, et al. Evolutionary dynamics of natural product biosynthesis in bacteria. Nat Prod Rep 2020;37(4): 566-99. CrossRef PubMed Google Scholar
-
214.Deng Q, Li Y, He W, Chen T, Liu N, Ma L, et al. A polyene macrolide targeting phospholipids in the fungal cell membrane. Nature 2025;640(8059): 743-51. CrossRef PubMed Google Scholar
-
215.Gupta VK, Bakshi U, Chang D, Lee AR, Davis JM, Chandrasekaran S, et al. TaxiBGC: a taxonomy-guided approach for profiling experimentally characterized microbial biosynthetic gene clusters and secondary metabolite production potential in metagenomes. mSystems 2022;7(6): e00925-e1022. CrossRef PubMed Google Scholar
-
216.Belknap KC, Park CJ, Barth BM, Andam CP. Genome mining of biosynthetic and chemotherapeutic gene clusters in Streptomyces bacteria. Sci Rep 2020;10(1): 2003. CrossRef PubMed Google Scholar
-
217.Smanski MJ, Schlatter DC, Kinkel LL. Leveraging ecological theory to guide natural product discovery. J Ind Microbiol Biotechnol 2016;43(2–3): 115-28. CrossRef PubMed Google Scholar
-
218.Wei B, Hu GA, Zhou ZY, Yu WC, Du AQ, Yang CL, et al. Global analysis of the biosynthetic chemical space of marine prokaryotes. Microbiome 2023;11(1): 144. CrossRef PubMed Google Scholar
-
219.Grubbs KJ, Bleich RM, Santa Maria KC, Allen SE, Farag S, AgBiome Team, et al. Large-scale bioinformatics analysis of Bacillus genomes uncovers conserved roles of natural products in bacterial physiology. mSystems 2017;2(6): e00040-e117. CrossRef PubMed Google Scholar
-
220.Liu Z, Suarez Duran HG, Harnvanichvech Y, Stephenson MJ, Schranz ME, Nelson D, et al. Drivers of metabolic diversification: how dynamic genomic neighbourhoods generate new biosynthetic pathways in the Brassicaceae. New Phytol 2020;227(4): 1109-23. CrossRef PubMed Google Scholar
-
221.Steinke K, Mohite OS, Weber T, Kovács ÁT. Phylogenetic distribution of secondary metabolites in the bacillus subtilis species complex. mSystems 2021;6(2): e00057-e121. CrossRef PubMed Google Scholar
-
222.Chevrette MG, Selem-Mojica N, Aguilar C, Labby K, Bustos-Diaz ED, Handelsman J, et al. Evolutionary genome mining for the discovery and engineering of natural product biosynthesis. In: Skellam E (Eds) Engineering natural product biosynthesis [Internet]. New York, NY: Springer US; 2022 [cited 2023 Aug 11]. p. 129–55. (Methods in Molecular Biology; vol. 2489). https://doi.org/10.1007/978-1-0716-2273-5_8 PubMed Google Scholar
-
223.Hannigan GD, Prihoda D, Palicka A, Soukup J, Klempir O, Rampula L, et al. A deep learning genome-mining strategy for biosynthetic gene cluster prediction. Nucleic Acids Res 2019;47(18): e110-e110. CrossRef PubMed Google Scholar
-
224.Louwen JJR, Kautsar SA, Van Der Burg S, Medema MH, Van Der Hooft JJJ. iPRESTO: Automated discovery of biosynthetic sub-clusters linked to specific natural product substructures. PLoS Comput Biol 2023;19(2): e1010462. CrossRef PubMed Google Scholar
-
225.Ziemert N, Podell S, Penn K, Badger JH, Allen E, Jensen PR. The natural product domain seeker NaPDoS: a phylogeny based bioinformatic tool to classify secondary metabolite gene diversity. PLoS ONE 2012;7(3): e34064. CrossRef PubMed Google Scholar
-
226.Cruz-Morales P, Kopp JF, Martínez-Guerrero C, Yáñez-Guerra LA, Selem-Mojica N, Ramos-Aboites H, et al. Phylogenomic analysis of natural products biosynthetic gene clusters allows discovery of arseno-organic metabolites in model Streptomycetes. Genome Biol Evol 2016;8(6): 1906-16. CrossRef PubMed Google Scholar
-
227.Xu M, Wright GD. Heterologous expression-facilitated natural products' discovery in actinomycetes. J Ind Microbiol Biotechnol 2019;46(3–4): 415-31. CrossRef PubMed Google Scholar
-
228.Teijaro CN, Adhikari A, Shen B. Challenges and opportunities for natural product discovery, production, and engineering in native producers versus heterologous hosts. J Ind Microbiol Biotechnol 2019;46(3–4): 433-44. CrossRef PubMed Google Scholar
-
229.Wang W, Zheng G, Lu Y. Recent advances in strategies for the cloning of natural product biosynthetic gene clusters. Front Bioeng Biotechnol. 2021. https://doi.org/10.3389/fbioe.2021.692797/full. PubMed Google Scholar
-
230.Bai S, Luo H, Tong H, Wu Y. Application and technical challenges in design, cloning, and transfer of large DNA. Bioengineering 2023;10(12): 1425. CrossRef PubMed Google Scholar
-
231.Chi H, Wang X, Shao Y, Qin Y, Deng Z, Wang L, et al. Engineering and modification of microbial chassis for systems and synthetic biology. Synth Syst Biotechnol 2019;4(1): 25-33. CrossRef PubMed Google Scholar
-
232.Calero P, Nikel PI. Chasing bacterial chassis for metabolic engineering: a perspective review from classical to non-traditional microorganisms. Microb Biotechnol 2019;12(1): 98-124. CrossRef PubMed Google Scholar
-
233.Wan J, Ma N, Yuan H. Recent advances in the direct cloning of large natural product biosynthetic gene clusters. Eng Microbiol 2023;3(3): 100085. CrossRef PubMed Google Scholar
-
234.Abbasi MN, Fu J, Bian X, Wang H, Zhang Y, Li A. Recombineering for genetic engineering of natural product biosynthetic pathways. Trends Biotechnol 2020;38(7): 715-28. CrossRef PubMed Google Scholar
-
235.Zhang JJ, Tang X, Moore BS. Genetic platforms for heterologous expression of microbial natural products. Nat Prod Rep 2019;36(9): 1313-32. CrossRef PubMed Google Scholar
-
236.Zhu S, Xu H, Liu Y, Hong Y, Yang H, Zhou C, et al. Computational advances in biosynthetic gene cluster discovery and prediction. Biotechnol Adv 2025;79: 108532. CrossRef PubMed Google Scholar
-
237.Wu C, Shang Z, Lemetre C, Ternei MA, Brady SF. Cadasides, calcium-dependent acidic lipopeptides from the soil metagenome that are active against multidrug-resistant bacteria. J Am Chem Soc 2019;141(9): 3910-9. CrossRef PubMed Google Scholar
-
238.Yuan Y, Huang C, Singh N, Xun G, Zhao H. Self-resistance-gene-guided, high-throughput automated genome mining of bioactive natural products from Streptomyces. Cell Syst 2025;16(3): 101237. CrossRef PubMed Google Scholar
-
239.Zhang MM, Wang Y, Ang EL, Zhao H. Engineering microbial hosts for production of bacterial natural products. Nat Prod Rep 2016;33(8): 963-87. CrossRef PubMed Google Scholar
-
240.Nidhi S, Anand U, Oleksak P, Tripathi P, Lal JA, Thomas G, et al. Novel CRISPR–Cas systems: an updated review of the current achievements, applications, and future research perspectives. IJMS 2021;22(7): 3327. CrossRef PubMed Google Scholar
-
241.Stoddard BL. Homing endonucleases: from microbial genetic invaders to reagents for targeted DNA modification. Structure 2011;19(1): 7-15. CrossRef PubMed Google Scholar
-
242.Urnov FD, Rebar EJ, Holmes MC, Zhang HS, Gregory PD. Genome editing with engineered zinc finger nucleases. Nat Rev Genet 2010;11(9): 636-46. CrossRef PubMed Google Scholar
-
243.Joung JK, Sander JD. TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol 2013;14(1): 49-55. CrossRef PubMed Google Scholar
-
244.Liu G, Lin Q, Jin S, Gao C. The CRISPR-Cas toolbox and gene editing technologies. Mol Cell 2022;82(2): 333-47. CrossRef PubMed Google Scholar
-
245.Wang D, Jin S, Lu Q, Chen Y. Advances and challenges in CRISPR/Cas-based fungal genome engineering for secondary metabolite production: a review. JoF 2023;9(3): 362. CrossRef PubMed Google Scholar
-
246.Xiao S, Deng Z, Gao J. CRISPR/Cas-based strategy for unearthing hidden chemical space from microbial genomes. Trends Chem 2021;3(12): 997-1001. CrossRef PubMed Google Scholar
-
247.Heng E, Tan LL, Zhang MM, Wong FT. CRISPR-Cas strategies for natural product discovery and engineering in actinomycetes. Process Biochem 2021;102: 261-8. CrossRef PubMed Google Scholar
-
248.Yang J, Song J, Feng Z, Ma Y. Application of CRISPR-Cas9 in microbial cell factories. Biotechnol Lett. 2025. https://doi.org/10.1007/s10529-025-03592-6. PubMed Google Scholar
-
249.Bushin LB, Covington BC, Rued BE, Federle MJ, Seyedsayamdost MR. Discovery and biosynthesis of Streptosactin, a Sactipeptide with an alternative topology encoded by commensal bacteria in the human microbiome. J Am Chem Soc 2020;142(38): 16265-75. CrossRef PubMed Google Scholar
-
250.Ameruoso A, Villegas Kcam MC, Cohen KP, Chappell J. Activating natural product synthesis using CRISPR interference and activation systems in Streptomyces. Nucleic Acids Res 2022;50(13): 7751-60. CrossRef PubMed Google Scholar
-
251.Peng R, Wang Y, Feng W, Yue X, Chen J, Hu X, et al. CRISPR/dCas9-mediated transcriptional improvement of the biosynthetic gene cluster for the epothilone production in Myxococcus xanthus. Microb Cell Fact. 2018. https://doi.org/10.1186/s12934-018-0867-1. PubMed Google Scholar
-
252.Huang X, Yang D, Zhang J, Xu J, Chen YE. Recent advances in improving gene-editing specificity through CRISPR–Cas9 nuclease engineering. Cells 2022;11(14): 2186. CrossRef PubMed Google Scholar
-
253.Zhou Q, Zhao Y, Ke C, Wang H, Gao S, Li H, et al. Repurposing endogenous type I-E CRISPR-Cas systems for natural product discovery in Streptomyces. Nat Commun 2024;15(1): 9833. CrossRef PubMed Google Scholar
-
254.Katzen R, Tsao GT. A view of the history of biochemical engineering. In: Fiechter A (Eds) History of modern biotechnology Ⅱ [Internet]. Berlin, Heidelberg: Springer Berlin Heidelberg; 2000 [cited 2023 Oct 14]. p. 77–91. (Scheper T, Babel W, Blanch HW, Cooney CL, Endo I, Enfors SO, et al., editors. Advances in Biochemical Engineering/Biotechnology; vol. 70). https://doi.org/10.1007/3-540-44965-5_4 PubMed Google Scholar
-
255.Caesar LK, Montaser R, Keller NP, Kelleher NL. Metabolomics and genomics in natural products research: complementary tools for targeting new chemical entities. Nat Prod Rep 2021;38(11): 2041-65. CrossRef PubMed Google Scholar
-
256.Moutinho TJ, Neubert BC, Jenior ML, Papin JA. Quantifying cumulative phenotypic and genomic evidence for procedural generation of metabolic network reconstructions. PLoS Comput Biol 2022;18(2): e1009341. CrossRef PubMed Google Scholar
-
257.Brandherm F, Gedeon J, Abboud O, Muhlhauser M. BigMEC: scalable service migration for mobile edge computing. In: 2022 IEEE/ACM 7th symposium on edge computing (SEC) [Internet]. Seattle, WA, USA: IEEE; 2022. p. 136–48 PubMed Google Scholar
-
258.Hérisson J, Duigou T, Du Lac M, Bazi-Kabbaj K, Sabeti Azad M, Buldum G, et al. The automated galaxy-SynBioCAD pipeline for synthetic biology design and engineering. Nat Commun 2022;13(1): 5082. CrossRef PubMed Google Scholar
-
259.Roy S, Radivojevic T, Forrer M, Marti JM, Jonnalagadda V, Backman T, et al. Multiomics data collection, visualization, and utilization for guiding metabolic engineering. Front Bioeng Biotechnol 2021;9(9): 612893. CrossRef PubMed Google Scholar
-
260.Qiu S, Yang A, Zeng H. Flux balance analysis-based metabolic modeling of microbial secondary metabolism: current status and outlook. PLoS Comput Biol 2023;19(8): e1011391. CrossRef PubMed Google Scholar
-
261.Banerjee D, Eng T, Lau AK, Sasaki Y, Wang B, Chen Y, et al. Genome-scale metabolic rewiring improves titers rates and yields of the non-native product indigoidine at scale. Nat Commun 2020;11(1): 5385. CrossRef PubMed Google Scholar
-
262.Courdavault V, O'Connor SE, Jensen MK, Papon N. Metabolic engineering for plant natural products biosynthesis: new procedures, concrete achievements and remaining limits. Nat Prod Rep 2021;38(12): 2145-53. CrossRef PubMed Google Scholar
-
263.Agarwal V, Blanton JM, Podell S, Taton A, Schorn MA, Busch J, et al. Metagenomic discovery of polybrominated diphenyl ether biosynthesis by marine sponges. Nat Chem Biol 2017;13(5): 537-43. CrossRef PubMed Google Scholar
-
264.Kwan BD, Seligmann B, Nguyen TD, Franke J, Dang TTT. Leveraging synthetic biology and metabolic engineering to overcome obstacles in plant pathway elucidation. Curr Opin Plant Biol 2023;71: 102330. CrossRef PubMed Google Scholar
-
265.Mosaei H, Molodtsov V, Kepplinger B, Harbottle J, Moon CW, Jeeves RE, et al. Mode of action of Kanglemycin A, an ansamycin natural product that is active against rifampicin-resistant Mycobacterium tuberculosis. Mol Cell 2018;72(2): 263-274.e5. CrossRef PubMed Google Scholar
-
266.Xu L, Nie D, Su BM, Xu XQ, Lin J. A chemoenzymatic strategy for the efficient synthesis of amphenicol antibiotic chloramphenicol mediated by an engineered l-threonine transaldolase with high activity and stereoselectivity. Catal Sci Technol 2023;13(3): 684-93. CrossRef PubMed Google Scholar
-
267.El-Sayed ASA, George NM, Abou-Elnour A, El-Mekkawy RM, El-Demerdash MM. Production and bioprocessing of camptothecin from Aspergillus terreus, an endophyte of Cestrum parqui, restoring their biosynthetic potency by Citrus limonum peel extracts. Microb Cell Fact. 2023. https://doi.org/10.1186/s12934-022-02012-y. PubMed Google Scholar
-
268.Ro DK, Paradise EM, Ouellet M, Fisher KJ, Newman KL, Ndungu JM, et al. Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature 2006;440(7086): 940-3. CrossRef PubMed Google Scholar
-
269.Aoki R, Kumagawa E, Kamata K, Ago H, Sakai N, Hasunuma T, et al. Engineering of acyl ligase domain in non-ribosomal peptide synthetases to change fatty acid moieties of lipopeptides. Commun Chem [Internet] 2025;8(1): 17. CrossRef PubMed Google Scholar
-
270.He F, Liu X, Tang M, Wang H, Wu Y, Liang S. CRISETR: an efficient technology for multiplexed refactoring of biosynthetic gene clusters. Nucleic Acids Res 2024;52(18): 11378-93. CrossRef PubMed Google Scholar
-
271.Su L, Hôtel L, Paris C, Chepkirui C, Brachmann AO, Piel J, et al. Engineering the stambomycin modular polyketide synthase yields 37-membered mini-stambomycins. Nat Commun 2022;13(1): 515. CrossRef PubMed Google Scholar
-
272.Del Vecchio F, Petkovic H, Kendrew SG, Low L, Wilkinson B, Lill R, et al. Active-site residue, domain and module swaps in modular polyketide synthases. J Ind Microbiol Biotechnol 2003;30(8): 489-94. CrossRef PubMed Google Scholar
-
273.Englund E, Schmidt M, Nava AA, Klass S, Keiser L, Dan Q, et al. Biosensor guided polyketide synthases engineering for optimization of domain exchange boundaries. Nat Commun 2023;14(1): 4871. CrossRef PubMed Google Scholar
-
274.Messenger SR, McGuinniety EMR, Stevenson LJ, Owen JG, Challis GL, Ackerley DF, et al. Metagenomic domain substitution for the high-throughput modification of nonribosomal peptides. Nat Chem Biol 2024;20(2): 251-60. CrossRef PubMed Google Scholar
-
275.Song C, Luan J, Li R, Jiang C, Hou Y, Cui Q, et al. RedEx: a method for seamless DNA insertion and deletion in large multimodular polyketide synthase gene clusters. Nucl Acids Res 2020;48(22): e130-e130. CrossRef PubMed Google Scholar
-
276.Liu R, Liang L, Freed EF, Choudhury A, Eckert CA, Gill RT. Engineering regulatory networks for complex phenotypes in E. coli. Nat Commun. 2020;11(1):4050. PubMed Google Scholar
-
277.Wittkopp PJ, Kalay G. Cis-regulatory elements: molecular mechanisms and evolutionary processes underlying divergence. Nat Rev Genet 2012;13(1): 59-69. CrossRef PubMed Google Scholar
-
278.Lyu HN, Liu HW, Keller NP, Yin WB. Harnessing diverse transcriptional regulators for natural product discovery in fungi. Nat Prod Rep 2020;37(1): 6-16. CrossRef PubMed Google Scholar
-
279.Turnbough CL. Regulation of bacterial gene expression by transcription attenuation. Microbiol Mol Biol Rev 2019;83(3): e00019-19. CrossRef PubMed Google Scholar
-
280.Bind S, Bind S, Sharma AK, Chaturvedi P. Epigenetic modification: a key tool for secondary metabolite production in microorganisms. Front Microbiol 2022;13(13): 784109. CrossRef PubMed Google Scholar
-
281.Tolibia SEM, Pacheco AD, Balbuena SYG, Rocha J, López Y, López VE. Engineering of global transcription factors in Bacillus, a genetic tool for increasing product yields: a bioprocess overview. World J Microbiol Biotechnol 2023;39(1): 12. CrossRef PubMed Google Scholar
-
282.Romero-Rodríguez A, Robledo-Casados I, Sánchez S. An overview on transcriptional regulators in Streptomyces. Biochimica et Biophysica Acta BBA Gene Regul Mech 2015;1849(8): 1017-39. CrossRef PubMed Google Scholar
-
283.El Hajj AC, Zetina-Serrano C, Tahtah N, Khoury AE, Atoui A, Oswald IP, et al. Regulation of secondary metabolism in the Penicillium Genus. IJMS 2020;21(24): 9462. CrossRef PubMed Google Scholar
-
284.Moon H, Han KH, Yu JH. Upstream regulation of development and secondary metabolism in Aspergillus species. Cells 2022;12(1): 2. CrossRef PubMed Google Scholar
-
285.Yoo YJ, Hwang J, Shin HL, Cui H, Lee J, Yoon YJ. Characterization of negative regulatory genes for the biosynthesis of rapamycin in Streptomyces rapamycinicus and its application for improved production. J Ind Microbiol Biotechnol 2015;42(1): 125-35. CrossRef PubMed Google Scholar
-
286.Xie WH, Deng HK, Hou J, Wang LJ. Synthetic small regulatory RNAs in microbial metabolic engineering. Appl Microbiol Biotechnol 2021;105(1): 1-12. CrossRef PubMed Google Scholar
-
287.Ji CH, Kim H, Kang HS. Synthetic inducible regulatory systems optimized for the modulation of secondary metabolite production in Streptomyces. ACS Synth Biol 2019;8(3): 577-86. CrossRef PubMed Google Scholar
-
288.Ondeyka JG, Zink DL, Young K, Painter R, Kodali S, Galgoci A, et al. Discovery of bacterial fatty acid synthase inhibitors from a Phoma species as antimicrobial agents using a new antisense-based strategy. J Nat Prod 2006;69(3): 377-80. CrossRef PubMed Google Scholar
-
289.Kunjapur AM, Prather KLJ. Development of a vanillate biosensor for the vanillin biosynthesis pathway in E. coli. ACS Synth Biol. 2019;8(9):1958–67. PubMed Google Scholar
-
290.Liu Y, Landick R, Raman S. A regulatory NADH/NAD+ redox biosensor for bacteria. ACS Synth Biol 2019;8(2): 264-73. CrossRef PubMed Google Scholar
-
291.Mitchler MM, Garcia JM, Montero NE, Williams GJ. Transcription factor-based biosensors: a molecular-guided approach for natural product engineering. Curr Opin Biotechnol 2021;69: 172-81. CrossRef PubMed Google Scholar
-
292.Swank Z, Laohakunakorn N, Maerkl SJ. Cell-free gene-regulatory network engineering with synthetic transcription factors. Proc Natl Acad Sci USA 2019;116(13): 5892-901. CrossRef PubMed Google Scholar
-
293.d'Oelsnitz S, Ellington AD, Ross DJ. Ligify: automated genome mining for ligand-inducible transcription factors [Internet]. Synth Biol. 2024. https://doi.org/10.1101/2024.02.20.581298. PubMed Google Scholar
-
294.Lai HY, Zhang ZY, Su ZD, Su W, Ding H, Chen W, et al. iProEP: a computational predictor for predicting promoter. Mol Therapy Nucl Acids 2019;17: 337-46. CrossRef PubMed Google Scholar
-
295.Wang Y, Tai S, Zhang S, Sheng N, Xie X. PromGER: promoter prediction based on graph embedding and ensemble learning for eukaryotic sequence. Genes 2023;14(7): 1441. CrossRef PubMed Google Scholar
-
296.Chung D, Barker BM, Carey CC, Merriman B, Werner ER, Lechner BE, et al. ChIP-seq and in vivo transcriptome analyses of the Aspergillus fumigatus SREBP SrbA reveals a new regulator of the fungal hypoxia response and virulence. PLoS Pathog 2014;10(11): e1004487. CrossRef PubMed Google Scholar
-
297.Zheng M, Barrera LO, Ren B, Wu YN. ChIP-chip: data, model, and analysis. Biometrics 2007;63(3): 787-96. CrossRef PubMed Google Scholar
-
298.Hellman LM, Fried MG. Electrophoretic mobility shift assay (EMSA) for detecting protein–nucleic acid interactions. Nat Protoc 2007;2(8): 1849-61. CrossRef PubMed Google Scholar
-
299.Jin LQ, Jin WR, Ma ZC, Shen Q, Cai X, Liu ZQ, et al. Promoter engineering strategies for the overproduction of valuable metabolites in microbes. Appl Microbiol Biotechnol 2019;103(21–22): 8725-36. CrossRef PubMed Google Scholar
-
300.Myronovskyi M, Luzhetskyy A. Native and engineered promoters in natural product discovery. Nat Prod Rep 2016;33(8): 1006-19. CrossRef PubMed Google Scholar
-
301.Johnson AO, Gonzalez-Villanueva M, Tee KL, Wong TS. An engineered constitutive promoter set with broad activity range for Cupriavidus necator H16. ACS Synth Biol 2018;7(8): 1918-28. CrossRef PubMed Google Scholar
-
302.Ji CH, Je HW, Kim H, Kang HS. Promoter engineering of natural product biosynthetic gene clusters in actinomycetes: concepts and applications. Nat Prod Rep 2024;41(4): 672-99. CrossRef PubMed Google Scholar
-
303.Lin SY, Oakley CE, Jenkinson CB, Chiang YM, Lee CK, Jones CG, et al. A heterologous expression platform in Aspergillus nidulans for the elucidation of cryptic secondary metabolism biosynthetic gene clusters: discovery of the Aspergillus fumigatus sartorypyrone biosynthetic pathway. Chem Sci 2023;14(40): 11022-32. CrossRef PubMed Google Scholar
-
304.Lalwani MA, Zhao EM, Wegner SA, Avalos JL. The Neurospora crassa inducible Q system enables simultaneous optogenetic amplification and inversion in Saccharomyces cerevisiae for bidirectional control of gene expression. ACS Synth Biol 2021;10(8): 2060-75. CrossRef PubMed Google Scholar
-
305.Wanka F, Cairns T, Boecker S, Berens C, Happel A, Zheng X, et al. Tet-on, or Tet-off, that is the question: advanced conditional gene expression in Aspergillus. Fungal Genet Biol 2016;89: 72-83. CrossRef PubMed Google Scholar
-
306.Gupta A, Reizman IMB, Reisch CR, Prather KLJ. Dynamic regulation of metabolic flux in engineered bacteria using a pathway-independent quorum-sensing circuit. Nat Biotechnol 2017;35(3): 273-9. CrossRef PubMed Google Scholar
-
307.Feng Y, Xie Z, Jiang X, Li Z, Shen Y, Wang B, et al. The applications of promoter-gene-engineered biosensors. Sensors 2018;18(9): 2823. CrossRef PubMed Google Scholar
-
308.Gilman J, Love J. Synthetic promoter design for new microbial chassis. Biochem Soc Trans 2016;44(3): 731-7. CrossRef PubMed Google Scholar
-
309.Wang Y, Wang H, Wei L, Li S, Liu L, Wang X. Synthetic promoter design in Escherichia coli based on a deep generative network. Nucl Acids Res 2020;48(12): 6403-12. CrossRef PubMed Google Scholar
-
310.Wu Y, Chen T, Liu Y, Tian R, Lv X, Li J, et al. Design of a programmable biosensor-CRISPRi genetic circuits for dynamic and autonomous dual-control of metabolic flux in Bacillus subtilis. Nucleic Acids Res 2020;48(2): 996-1009. CrossRef PubMed Google Scholar
-
311.Cobb RE, Wang Y, Zhao H. High-efficiency multiplex genome editing of Streptomyces species using an engineered CRISPR/Cas system. ACS Synth Biol 2015;4(6): 723-8. CrossRef PubMed Google Scholar
-
312.Bode E, Assmann D, Happel P, Meyer E, Münch K, Rössel N, et al. easyPACId, a simple method for induced production, isolation, identification, and testing of natural products from proteobacteria. BIO-PROTOCOL [Internet] 2023;13(13): e4709. PubMed Google Scholar
-
313.Gulsen SH, Tileklioglu E, Bode E, Cimen H, Ertabaklar H, Ulug D, et al. Antiprotozoal activity of different Xenorhabdus and Photorhabdus bacterial secondary metabolites and identification of bioactive compounds using the easyPACId approach. Sci Rep 2022;12(1): 10779. CrossRef PubMed Google Scholar
-
314.Mózsik L, Iacovelli R, Bovenberg RAL, Driessen AJM. Transcriptional activation of biosynthetic gene clusters in filamentous fungi. Front Bioeng Biotechnol 2022;15(10): 901037. CrossRef PubMed Google Scholar
-
315.Zhang Y, Chen H, Zhang Y, Yin H, Zhou C, Wang Y. Direct RBS Engineering of the biosynthetic gene cluster for efficient productivity of violaceins in E. coli. Microb Cell Fact. 2021. https://doi.org/10.1186/s12934-021-01518-1. PubMed Google Scholar
-
316.Salis HM, Mirsky EA, Voigt CA. Automated design of synthetic ribosome binding sites to control protein expression. Nat Biotechnol 2009;27(10): 946-50. CrossRef PubMed Google Scholar
-
317.Zelcbuch L, Antonovsky N, Bar-Even A, Levin-Karp A, Barenholz U, Dayagi M, et al. Spanning high-dimensional expression space using ribosome-binding site combinatorics. Nucl Acids Res 2013;41(9): e98-e98. CrossRef PubMed Google Scholar
-
318.You J, Wang Y, Wang K, Du Y, Zhang X, Zhang X, et al. Utilizing 5′ UTR engineering enables fine-tuning of multiple genes within operons to balance metabolic flux in bacillus subtilis. Biology 2024;13(4): 277. CrossRef PubMed Google Scholar
-
319.Curran KA, Karim AS, Gupta A, Alper HS. Use of expression-enhancing terminators in Saccharomyces cerevisiae to increase mRNA half-life and improve gene expression control for metabolic engineering applications. Metabol Eng 2013;19: 88-97. CrossRef PubMed Google Scholar
-
320.Sun D, Liu C, Zhu J, Liu W. Connecting metabolic pathways: sigma factors in Streptomyces spp. Front Microbiol 2017;19(8): 2546. CrossRef PubMed Google Scholar
-
321.Zhu S, Duan Y, Huang Y. The application of ribosome engineering to natural product discovery and yield improvement in Streptomyces. Antibiotics 2019;8(3): 133. CrossRef PubMed Google Scholar
-
322.Sherwood AV, Henkin TM. Riboswitch-mediated gene regulation: novel RNA architectures dictate gene expression responses. Annu Rev Microbiol 2016;70(1): 361-74. CrossRef PubMed Google Scholar
-
323.Ueno K, Tsukakoshi K, Ikebukuro K. Riboregulator elements as tools to engineer gene expression in cyanobacteria. Appl Microbiol Biotechnol 2018;102(18): 7717-23. CrossRef PubMed Google Scholar
-
324.Hong B, Luo T, Lei X. Late-stage diversification of natural products. ACS Cent Sci 2020;6(5): 622-35. CrossRef PubMed Google Scholar
-
325.Zhang P, Eun J, Elkin M, Zhao Y, Cantrell RL, Newhouse TR. A neural network model informs the total synthesis of clovane sesquiterpenoids. Nat Synth 2023;2(6): 527-34. CrossRef PubMed Google Scholar
-
326.Li Y, Cheng S, Tian Y, Zhang Y, Zhao Y. Recent ring distortion reactions for diversifying complex natural products. Nat Prod Rep 2022;39(10): 1970-92. CrossRef PubMed Google Scholar
-
327.Rudroff F, Mihovilovic MD, Gröger H, Snajdrova R, Iding H, Bornscheuer UT. Opportunities and challenges for combining chemo- and biocatalysis. Nat Catal 2018;1(1): 12-22. CrossRef PubMed Google Scholar
-
328.Yi D, Bayer T, Badenhorst CPS, Wu S, Doerr M, Höhne M, et al. Recent trends in biocatalysis. Chem Soc Rev 2021;50(14): 8003-49. CrossRef PubMed Google Scholar
-
329.Lin B, Tao Y. Whole-cell biocatalysts by design. Microb Cell Fact 2017;16(1): 106. CrossRef PubMed Google Scholar
-
330.Qin D, Dong J. Multi-level optimization and strategies in microbial biotransformation of nature products. Molecules 2023;28(6): 2619. CrossRef PubMed Google Scholar
-
331.Huang T, Zhang F, Wang B, Ye WS, Peng QM, Wu FA, et al. Flavonoid glycoside transformation catalyzed by whole-cell catalysts using a PVDF membrane reactor coupled with reaction and separation. Waste Biomass Valor 2020;11(10): 5321-32. CrossRef PubMed Google Scholar
-
332.Cen X, Liu Y, Chen B, Liu D, Chen Z. Metabolic engineering of Escherichia coli for De Novo production of 1, 5-pentanediol from glucose. ACS Synth Biol 2021;10(1): 192-203. CrossRef PubMed Google Scholar
-
333.Zhang J, Chang K, Tay J, Tiong E, Heng E, Seah T, et al. Hyper-porous encapsulation of microbes for whole cell biocatalysis and biomanufacturing. Microb Cell Fact. 2025. https://doi.org/10.1186/s12934-025-02675-3. PubMed Google Scholar
-
334.Chen Y, Emileh A, Brideau N, Britton J. Cell-free reactions in continuous manufacturing systems. Curr Opin Green Sustain Chem 2020;25: 100380. CrossRef PubMed Google Scholar
-
335.Espinel-Ríos S, Huber N, Alcalá-Orozco EA, Morabito B, Rexer TFT, Reichl U, et al. Cell-free biosynthesis meets dynamic optimization and control: a fed-batch framework. IFAC-PapersOnLine 2022;55(23): 92-7. CrossRef PubMed Google Scholar
-
336.Rasor BJ, Yi X, Brown H, Alper HS, Jewett MC. An integrated in vivo/in vitro framework to enhance cell-free biosynthesis with metabolically rewired yeast extracts. Nat Commun 2021;12(1): 5139. CrossRef PubMed Google Scholar
-
337.Huang K, Morato NM, Feng Y, Cooks RG. High-throughput diversification of complex bioactive molecules by accelerated synthesis in microdroplets. Angew Chem Int Ed 2023;62(22): e202300956. CrossRef PubMed Google Scholar
-
338.Zhao C, Ye Z, Ma Z, Wildman SA, Blaszczyk SA, Hu L, et al. A general strategy for diversifying complex natural products to polycyclic scaffolds with medium-sized rings. Nat Commun 2019;10(1): 4015. CrossRef PubMed Google Scholar
-
339.Brooks SM, Alper HS. Applications, challenges, and needs for employing synthetic biology beyond the lab. Nat Commun 2021;12(1): 1390. CrossRef PubMed Google Scholar
-
340.Bogart JW, Cabezas MD, Vögeli B, Wong DA, Karim AS, Jewett MC. Cell-free exploration of the natural product chemical space. ChemBioChem 2021;22(1): 84-91. CrossRef PubMed Google Scholar
-
341.Li J, Zhang L, Liu W. Cell-free synthetic biology for in vitro biosynthesis of pharmaceutical natural products. Synth Syst Biotechnol 2018;3(2): 83-9. CrossRef PubMed Google Scholar
-
342.Niu FX, Yan ZB, Huang YB, Liu JZ. Cell-free biosynthesis of chlorogenic acid using a mixture of chassis cell extracts and purified spy-cyclized enzymes. J Agric Food Chem 2021;69(28): 7938-47. CrossRef PubMed Google Scholar
-
343.Nguyen TD, Dang TTT. Cytochrome P450 enzymes as key drivers of alkaloid chemical diversification in plants. Front Plant Sci 2021;2(12): 682181. CrossRef PubMed Google Scholar
-
344.Bai T, Matsuda Y, Tao H, Mori T, Zhang Y, Abe I. Structural diversification of andiconin-derived natural products by α-ketoglutarate-dependent dioxygenases. Org Lett 2020;22(11): 4311-5. CrossRef PubMed Google Scholar
-
345.Toplak M, Teufel R. Three rings to rule them all: how versatile flavoenzymes orchestrate the structural diversification of natural products. Biochemistry 2022;61(2): 47-56. CrossRef PubMed Google Scholar
-
346.Deng Y, Zhou Q, Wu Y, Chen X, Zhong F. Properties and mechanisms of flavin-dependent monooxygenases and their applications in natural product synthesis. IJMS 2022;23(5): 2622. CrossRef PubMed Google Scholar
-
347.Teufel R. Flavin-catalyzed redox tailoring reactions in natural product biosynthesis. Arch Biochem Biophys 2017;632: 20-7. CrossRef PubMed Google Scholar
-
348.Ditzel A, Zhao F, Gao X, Phillips GN. Utilizing a cell-free protein synthesis platform for the biosynthesis of a natural product, caffeine. Synthetic Biol. 2023. https://doi.org/10.1093/synbio/ysad017/7491574. PubMed Google Scholar
-
349.Siebels I, Nowak S, Heil CS, Tufar P, Cortina NS, Bode HB, et al. Cell-free synthesis of natural compounds from genomic DNA of biosynthetic gene clusters. ACS Synth Biol 2020;9(9): 2418-26. CrossRef PubMed Google Scholar
-
350.Galloway WRJD, Isidro-Llobet A, Spring DR. Diversity-oriented synthesis as a tool for the discovery of novel biologically active small molecules. Nat Commun 2010;1(1): 80. CrossRef PubMed Google Scholar
-
351.Mortensen KT, Osberger TJ, King TA, Sore HF, Spring DR. Strategies for the diversity-oriented synthesis of macrocycles. Chem Rev 2019;119(17): 10288-317. CrossRef PubMed Google Scholar
-
352.Eymery M, Tran-Nguyen VK, Boumendjel A. Diversity-oriented synthesis: amino acetophenones as building blocks for the synthesis of natural product analogs. Pharmaceuticals 2021;14(11): 1127. CrossRef PubMed Google Scholar
-
353.Galloway WRJD, Isidro-Llobet A, Spring DR. Diversity-oriented synthesis as a tool for the discovery of novel biologically active small molecules. Nat Commun [Internet] 2010;1(1): 80. CrossRef PubMed Google Scholar
-
354.Grossmann A, Bartlett S, Janecek M, Hodgkinson JT, Spring DR. Diversity-oriented synthesis of drug-like macrocyclic scaffolds using an orthogonal organo- and metal catalysis strategy. Angew Chem Int Ed 2014;53(48): 13093-7. CrossRef PubMed Google Scholar
-
355.Grenning AJ, Boyce JH, Porco JA. Rapid synthesis of polyprenylated acylphloroglucinol analogs via dearomative conjunctive allylic annulation. J Am Chem Soc 2014;136(33): 11799-804. CrossRef PubMed Google Scholar
-
356.Nayak A, Saxena H, Bathula C, Kumar T, Bhattacharjee S, Sen S, et al. Diversity-oriented synthesis derived indole based spiro and fused small molecules kills artemisinin-resistant Plasmodium falciparum. Malar J. 2021. https://doi.org/10.1186/s12936-021-03632-2. PubMed Google Scholar
-
357.Maier ME. Design and synthesis of analogues of natural products. Org Biomol Chem 2015;13(19): 5302-43. CrossRef PubMed Google Scholar
-
358.Lee HY, Harvey CJB, Cane DE, Khosla C. Improved precursor-directed biosynthesis in E. coli via directed evolution. J Antibiot. 2011;64(1):59–64. PubMed Google Scholar
-
359.Gross H, Stockwell VO, Henkels MD, Nowak-Thompson B, Loper JE, Gerwick WH. The genomisotopic approach: a systematic method to isolate products of orphan biosynthetic gene clusters. Chem Biol 2007;14(1): 53-63. CrossRef PubMed Google Scholar
-
360.Pett-Ridge J, Weber PK. NanoSIP: NanoSIMS applications for microbial biology. In: Navid A, editor. Microbial systems biology. Totowa, NJ: Humana Press; 2012. p. 375–408. https://doi.org/10.1007/978-1-61779-827-6_13. PubMed Google Scholar
-
361.Foster RA, Kuypers MMM, Vagner T, Paerl RW, Musat N, Zehr JP. Nitrogen fixation and transfer in open ocean diatom–cyanobacterial symbioses. ISME J 2011;5(9): 1484-93. CrossRef PubMed Google Scholar
-
362.Zhang Z, He X, Zhang X, Li D, Wu G, Liu Z, et al. Production of multiple Talaroenamines from Penicillium malacosphaerulum via one-pot/two-stage precursor-directed biosynthesis. J Nat Prod 2022;85(9): 2168-76. CrossRef PubMed Google Scholar
-
363.Hermane J, Eichner S, Mancuso L, Schröder B, Sasse F, Zeilinger C, et al. New geldanamycin derivatives with anti Hsp properties by mutasynthesis. Org Biomol Chem 2019;17(21): 5269-78. CrossRef PubMed Google Scholar
-
364.Yao Z, Sun C, Xia Y, Wang F, Fu L, Ma J, et al. Mutasynthesis of antibacterial halogenated actinomycin analogues. J Nat Prod 2021;84(8): 2217-25. CrossRef PubMed Google Scholar
-
365.Gou L, Wu Q, Lin S, Li X, Liang J, Zhou X, et al. Mutasynthesis of pyrrole spiroketal compound using calcimycin 3-hydroxy anthranilic acid biosynthetic mutant. Appl Microbiol Biotechnol 2013;97(18): 8183-91. CrossRef PubMed Google Scholar
-
366.Sialer C, García I, González-Sabín J, Braña AF, Méndez C, Morís F, et al. Generation by mutasynthesis of potential neuroprotectant derivatives of the bipyridyl collismycin A. Bioorg Med Chem Lett 2013;23(20): 5707-9. CrossRef PubMed Google Scholar
-
367.Xie F, Dai S, Zhao Y, Huang P, Yu S, Ren B, et al. Generation of fluorinated amychelin siderophores against pseudomonas aeruginosa infections by a combination of genome mining and mutasynthesis. Cell Chem Biol 2020;27(12): 1532-1543.e6. CrossRef PubMed Google Scholar
-
368.Kronenwerth M, Brachmann AO, Kaiser M, Bode HB. Bioactive derivatives of isopropylstilbene from mutasynthesis and chemical synthesis. ChemBioChem 2014;15(18): 2689-91. CrossRef PubMed Google Scholar
-
369.Basnet BB, Zhou ZY, Wei B, Wang H. Advances in AI-based strategies and tools to facilitate natural product and drug development. Crit Rev Biotechnol 2025;30: 1-32. PubMed Google Scholar
-
370.Gangwal A, Lavecchia A. Artificial intelligence in natural product drug discovery: current applications and future perspectives. J Med Chem 2025;68(4): 3948-69. CrossRef PubMed Google Scholar
-
371.Sahayasheela VJ, Lankadasari MB, Dan VM, Dastager SG, Pandian GN, Sugiyama H. Artificial intelligence in microbial natural product drug discovery: current and emerging role. Nat Prod Rep 2022;39(12): 2215-30. CrossRef PubMed Google Scholar
-
372.Skinnider MA, Johnston CW, Gunabalasingam M, Merwin NJ, Kieliszek AM, MacLellan RJ, et al. Comprehensive prediction of secondary metabolite structure and biological activity from microbial genome sequences. Nat Commun 2020;11(1): 6058. CrossRef PubMed Google Scholar
-
373.Gluck-Thaler E, Haridas S, Binder M, Grigoriev IV, Crous PW, Spatafora JW, et al. The architecture of metabolism maximizes biosynthetic diversity in the largest class of fungi. Mol Biol Evol 2020;37(10): 2838-56. CrossRef PubMed Google Scholar
-
374.Hjörleifsson Eldjárn G, Ramsay A, Van Der Hooft JJJ, Duncan KR, Soldatou S, Rousu J, et al. Ranking microbial metabolomic and genomic links in the NPLinker framework using complementary scoring functions. PLoS Comput Biol 2021;17(5): e1008920. CrossRef PubMed Google Scholar
-
375.Zallot R, Oberg N, Gerlt JA. The EFI web resource for genomic enzymology tools: leveraging protein, genome, and metagenome databases to discover novel enzymes and metabolic pathways. Biochemistry 2019;58(41): 4169-82. CrossRef PubMed Google Scholar
-
376.Erbilgin O, Rübel O, Louie KB, Trinh M, Raad MD, Wildish T, et al. MAGI: a method for metabolite annotation and gene integration. ACS Chem Biol 2019;14(4): 704-14. CrossRef PubMed Google Scholar
-
377.van Heel AJ, de Jong A, Song C, Viel JH, Kok J, Kuipers OP. BAGEL4: a user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res 2018;46(W1): W278-81. CrossRef PubMed Google Scholar
-
378.Chevrette MG, Aicheler F, Kohlbacher O, Currie CR, Medema MH. SANDPUMA: ensemble predictions of nonribosomal peptide chemistry reveal biosynthetic diversity across Actinobacteria. Bioinformatics 2017;33(20): 3202-10. CrossRef PubMed Google Scholar
-
379.Cao L, Gurevich A, Alexander KL, Naman CB, Leão T, Glukhov E, et al. MetaMiner: a scalable peptidogenomics approach for discovery of ribosomal peptide natural products with blind modifications from microbial communities. Cell Syst 2019;9(6): 600-608.e4. CrossRef PubMed Google Scholar
-
380.Carroll LM, Larralde M, Fleck JS, Ponnudurai R, Milanese A, Cappio E, et al. Accurate de novo identification of biosynthetic gene clusters with GECCO [Internet]. Bioinformatics. 2021. https://doi.org/10.1101/2021.05.03.442509. PubMed Google Scholar
-
381.Sanchez S, Rogers JD, Rogers AB, Nassar M, McEntyre J, Welch M, et al. Expansion of novel biosynthetic gene clusters from diverse environments using SanntiS [Internet]. Bioinformatics. 2023. https://doi.org/10.1101/2023.05.23.540769. PubMed Google Scholar
-
382.Lai Q, Yao S, Zha Y, Zhang H, Zhang H, Ye Y, et al. Deciphering the biosynthetic potential of microbial genomes using a BGC language processing neural network model. Nucl Acids Res 2025;53(7): gkaf305. CrossRef PubMed Google Scholar
-
383.Kloosterman AM, Cimermancic P, Elsayed SS, Du C, Hadjithomas M, Donia MS, et al. Expansion of RiPP biosynthetic space through integration of pan-genomics and machine learning uncovers a novel class of lanthipeptides. PLoS Biol 2020;18(12): e3001026. CrossRef PubMed Google Scholar
-
384.De Los Santos ELC. NeuRiPP: neural network identification of RiPP precursor peptides. Sci Rep 2019;9(1): 13406. CrossRef PubMed Google Scholar
-
385.Merwin NJ, Mousa WK, Dejong CA, Skinnider MA, Cannon MJ, Li H, et al. DeepRiPP integrates multiomics data to automate discovery of novel ribosomally synthesized natural products. Proc Natl Acad Sci USA 2020;117(1): 371-80. CrossRef PubMed Google Scholar
-
386.Leão TF, Wang M, da Silva R, Gurevich A, Bauermeister A, Gomes PWP, et al. NPOmix: a machine learning classifier to connect mass spectrometry fragmentation data to biosynthetic gene clusters. PNAS Nexus 2022;1(5): pgac257. CrossRef PubMed Google Scholar
-
387.Kim HW, Zhang C, Reher R, Wang M, Alexander KL, Nothias LF, et al. DeepSAT: learning molecular structures from nuclear magnetic resonance data. J Cheminform 2023;15(1): 71. CrossRef PubMed Google Scholar
-
388.Rios-Martinez C, Bhattacharya N, Amini AP, Crawford L, Yang KK. Deep self-supervised learning for biosynthetic gene cluster detection and product classification. PLoS Comput Biol 2023;19(5): e1011162. CrossRef PubMed Google Scholar
-
389.Li YF, Tsai KJS, Harvey CJB, Li JJ, Ary BE, Berlew EE, et al. Comprehensive curation and analysis of fungal biosynthetic gene clusters of published natural products. Fungal Genet Biol 2016;89: 18-28. CrossRef PubMed Google Scholar
Copyright information
© The Author(s) 2026
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.