Cucurbitane triterpenoids from Hemsleya penxianensis
Abstract
Two new cucurbitacins, jinfushanencins A(1) and B(2), seven new cucurbitane glycosides, jinfushanosides E-K(3-9), along with nine known analogues, were obtained from the tubers of Hemsleya penxianensis. Their structures were elucidated on the basis of extensive spectroscopic and chemical methods. Selected isolates were tested their anti-HIV-1 activities, and compound 5 showed weak anti-HIV-1 in C8166 cell(EC50=5.9 μg/mL) with a selectivity index of 13.5.Keywords
cucurbitane triterpenoid jinfushanencin jinfushanoside Hemsleya penxianensisIntroduction
The genus Hemsleya(Cucurbitaceae), consisting of more than twenty species, are mainly distributed in Yunnan and Sichuan Provinces.1 The tubers of Hemsleya plants are known as famous herbal medicine in China, used in the treatment of bacillary dysentery, bronchitis, and tuberculosis etc.1 The Hemsleya species distributed at Jinfu Mountain was initially recorded as H. jinfushanensis, and then as H. penxianensis var. jinfushanensis by Zhang W. J. and Shen L. T., but was ascribed to H. penxianensis by Li D. Z.2, 3 Previously, the phytochemical and bioactive studies of this plant have been reported.4, 5 In continuation of these study, nine new cucurbitane triterpenoids and nine known analogues were obtained. Herein, we describe the isolation and structural elucidation of the new ones.
|
Results and Discussion
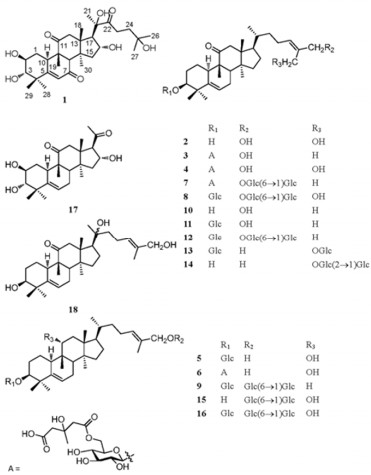
Jinfushanencin A (1) was obtained as colorless powder with the empirical molecular formula of C30H46O8, in agreement with the negative ion HRESIMS (m/z 533.3145 [M – H]-, calcd for C30H45O8, 533.3114) and 13C NMR spectroscopic data. The IR spectrum revealed absorptions at 3436, 1696, and 1648 cm–1, suggestive of hydroxyl and conjugated carbonyl groups. Obvious signals observed in the 1H NMR spectrum were eight methyl singlets at δH 1.24 (3H, s), 1.27 (3H, s), 1.36 (3H×2, s), 1.40 (3H, s), 1.48 (3H, s), 1.57 (3H, s), and 1.63 (3H, s), an olefinic proton singlet at δH 6.50 (1H, s), as well as two doublets at δH 3.54 (1H, d, J=9.1 Hz) and 2.92 (1H, d, J=7.0 Hz). The 13C NMR and DEPT spectra revealed 30 carbon signals due to eight methyls, five methylenes, seven methines, and ten quaternary carbons (including three carbonyl carbons and one olefinic carbon), which were assigned to a triterpene skeleton. Considering the fact that the tetracyclic triterpenoids isolated thus far from the genus Hemsleya are cucurbitane-type compounds, in combination with four characteristic quaternary carbons (δC 44.4 (C-4), 49.2 (C-9), 48.5 (C-13), and 50.7 (C-14)) at high field, compound 1 was tentatively proposed to be a cucurbitacin.6 Further, comparison of the NMR data of 1 with those of 2β, 3β, 16α, 20(R), 25-pentahydroxy-9-methyl-19-norlanost-5-en-7, 11, 22-trione indicated that these two compounds were structurally almost identical with exception of the configuration of the hydroxyl group at C-3.7 Detailed comparison of the NMR data of these two compounds disclosed that two proton signals at δH 4.22 (1H, m) and 3.54 (1H, d, J=9.1 Hz), ascribed to H-2 and H-3 by 2D NMR, in 1, instead of the signals at δH 4.61 (1H, br. d, J=10.9 Hz, H-2) and 3.98 (1H, br. s, H-3) in the known compound. All evidences mentioned above suggested that compound 1 had a 2β, 3α-diol moiety.
1H NMR (500 MHz) spectroscopic data of compounds 1 and 2, and the aglycones of 3-9 (in C5D5N, δ in ppm, J in Hz)
The ROESY correlation observed between δH 1.40 (3H, s, H-28) and δH 3.24 (1H, m, H-10) established an α configuration for C-28, together with the ROESY correlations between δH 4.22 (1H, m, H-2) and H-10, and between δH 1.48 (3H, s, H-29) and δH 3.54 (1H, d, J=9.1 Hz, H-3), confirmed the 2β and 3α substituents. The rings BD and side chain of 1 were identical to those of 2β, 3β, 16α, 20(R), 25-pentahydroxy-9-methyl-19-norlanost-5-en-7, 11, 22-trione by detailed analysis of the 13C, 1H, HMBC, HSQC, and 1H-1H COSY spectra of 1.
The molecular formula of jinfushanencin B (2) was determined to be C30H48O4 by negative ion HRESIMS (found [M - H]-471.3487, calcd for 471.3474). From the 13C NMR data in combination with DEPT spectrum, a ketone carbon, two olefinic methines, two olefinic quaternary carbons, one oxygenated methine, two oxygenated methylenes, four methines, eight methylenes, four quaternary carbons, and six methyls were observed. The NMR data of 2 were very similar with those of carnosiflogenin A (10).8 The difference was that the methyl signal for C-27 in 10 was replaced by an oxygenated methylene (δC 58.5) in 2, in accordance with the olefinic quaternary carbon resonance at δC136.1 (C-25) in 10 being shifted downfield to δC 140.9 in 2. These changes suggested that the methyl at C-27 in 10 was oxygenated to be a hydroxymethyl in 2. The HMBC correlations from δH 4.73 (s, H-27) to δC 65.4 (t, C-26), 140.9 (s, C-25), and 127.5 (d, C-24), were also supporting evidences for the above deduction.
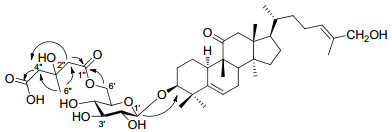
Jinfushanoside E (3) had a molecular formula of C42H66O12 determined by HRESIMS, 13C NMR, and DEPT experiments. The molecular weight of 3 was 144 mass units more than that of delavanoside D (11).9The 13C NMR and DEPT data (Tables 1-5) of 3 were very similar with those of 11 exception of six additional resonances observed at δC 171.8 (s, C-1″), 46.4 (t, C-2″), 70.2 (s, C-3″), 46.8 (t, C-4″), 174.9 (s, C-5″), and 28.8 (q, C-6″). In the HMBC spectrum, correlations revealed from δH 3.14 (H-2″) to C-1″, 3″, 4″, 6″ and from δH 3.21 (2H, br. d, J=14.2 Hz, H-4″) to C-2″, 3″, 5″, 6″ (Figure 1) manifested the additional structural feature in 3 being a hydroxymethyl glutaryl (HMG) group.10 Additionally, HMBC correlation140 J. C. CHEN et al. Nat. Prod. Bioprospect. 2012, 2, 138–144 observed from δH 5.03 (d, J=11.1 Hz, Ha-6′) and δH 4.71 (m, Hb-6′) to δC 171.8 (s, C-1″) revealed that the HMG group was linked to C-6′ of the sugar unit. After alkaline hydrolysis of 3 with 5 M NaOH solvent, 11 was detected in reaction mixture by TLC. The rings A-D and side chain of 3 were identical to those of 11 by detailed analysis of the 13C, 1H, HMBC, HSQC, and 1H-1H COSY spectra of 3.
Key HMBC correlations of compound 3
Jinfushanoside F (4) displayed a quasimolecular ion peak at m/z=777.4436 [M - H]-(cald for C42H65O13, 777.4425), which was consistent with the molecular formula C42H66O13. Comparison of NMR data of 4 (Tables 1-5) with those of jinfushanoside B revealed great similarity.4 The difference was that the C-6′ position of the sugar unit in 4 was ester-linked to a HMG functionality, on the basis of the HMBC correlations from δH 5.04 (d, J=11.3 Hz, Ha-6′) and δH 4.72 (m, Hb-6′) to δC 171.7 (s, C-1″), from δH 3.11 (d, J=14.0 Hz, Ha-2″) and δH 3.08 (d, J=14.0 Hz, Hb-2″) to δC 171.7 (s, C-1″), 70.2 (s, C-3″), 47.0 (t, C-4″), and 28.4 (q, C-6″), and from δH 3.17 (br. d, J=13.8 Hz, H-4″) to δC 46.7 (t, C-2″), 70.2 (s, C-3″), 175.6 (s, C-5″), and 28.4 (q, C-6″). After alkaline hydrolysis of 4 with 5 M NaOH solvent, jinfushanoside B was detected in reaction mixture by TLC. The rings A-D and side chain of 4 were identical to those of jinfushanoside B by detailed analysis of the 13C, 1H, HMBC, HSQC, and 1H-1H COSY spectra of 4.
13C NMR (125 MHz) spectroscopic data of compounds 1 and 2, and the aglycones of 3-9 (in C5D5N, δ in ppm)
1H NMR spectroscopic data of the sugar moieties of compounds 3-9
13C NMR spectroscopic data of the sugar moieties of compounds 3-9
1H and 13C NMR spectroscopic data of hydroxymethyl glutaryl (HMG) groups in compounds 3, 4, 6, and 7
The negative ion HRESIMS of jinfushanoside G (5) showed a molecular ion peak at 619.4183 [M - H]-, in according with an empirical molecular formula of C36H60O8. The molecular weight of 5 was 162 mass units more than that of 3β, 11α, 26-trihydroxycucurbita-5, 24-diene, 8 which implied an additional hexose unit in 5. Comparison of the 13C NMR spectrum of 5 with that of the known compound, an additional glucose signals at δC 107.3 (d), 75.5 (d), 78.7 (d), 71.8 (d), 78.1 (d), and 63.1 (t), as well as the signal of C-3 being shifted downfield by 11.4 ppm to δC 87.9 (d) in 5, were observed, which indicated an additional glucopyranosyl moiety being linked at C-3 in 5. This deduction was confirmed by the HMBC correlations from the anomeric proton signal at δH4.86 (d, J=7.8 Hz, H-1′) to δC 87.9 (d, C-3). The coupling value (J=7.8 Hz) of anomeric proton suggested the presence of a β-glucopyranosyl moiety, and the sugar was determined to be a D-glucose by comparison with authentic sample on GC analysis after acid hydrolysis of 5.
Jinfushanoside H (6) was determined to have the molecular formula C42H68O12 by negative ion HRESIMS (found [M - H]- 763.4648, calcd for 763.4632). Comparison of NMR data (Tables 1-5) of 6 with those of 5 displayed great similarity. The difference was that an additional HMG group was esterlinked to the C-6′ position of the sugar unit in 6, which was proved by the HMBC correlations from δH 5.03 (d, J=11.3 Hz, Ha-6') and δH 4.72 (m, Hb-6′) to δC 171.7 (s, C-1″) and from δH 3.14 (2H, m, H-2″) to δC 70.2 (s, C-3″), 46.8 (t, C-4″), and 28.3 (q, C-6″). After alkaline hydrolysis of 6 with 5 M NaOH solvent, 5 was detected in reaction mixture by TLC. The rings A-D and side chain of 6 were identical to those of 5 by detailed analysis of the 13C, 1H, HMBC, HSQC, and 1H-1H COSY spectra of 6.
Jinfushanoside I (7) had the molecular formula C54H86O22 based on HRESIMS ([M - H]-1085.5546, calcd for 1085.5532) data. The molecular weight of 7 was 144 mass units more than that of carnosifloside Ⅲ (12), 8 which implied an additional HMG group in 7. Comparison of the 13C NMR spectrum of 7 with that of the known compound, an additional HMG signals at δC 171.6 (s), 47.3 (t), 70.6 (s), 47.4 (t), 174.8 (s), and 28.3 (q), and the signal of C-6′ being shifted downfield to δC 64.6 (t) in 7, were observed, which indicated that the HMG functionality was ester-linked at C-6′ in 7. This proposal was proved by the observed HMBC correlation from δH 4.98 (m, Ha-6′) and 4.69 (m, Hb-6′) to δC 171.6 (s, C-1″). After alkaline hydrolysis of 7 with 5 M NaOH solvent, carnosifloside Ⅲ (12) was detected in reaction mixture by TLC.
Jinfushanoside J (8) displayed a quasimolecular ion peak at m/z=957.5050 [M - H]-(cald for C48H77O19, 957.5059), in accordance with the molecular formula C48H78O19. The molecular weight of 8 was 162 mass units more than that of jinfushanoside D, 4 which implied an additional hexose unit in 8. In the HMQC-TOCSY spectrum, the anomeric proton at δH 4.83 (1H, d, J=7.8 Hz, H-1′) was correlated with six carbons at δC 107.3 (d), 75.5 (d), 78.7 (d), 71.8 (d), 78.4 (d), and 63.1 (t). This glucose was attached to C-3 based on the HMBC correlation between the anomeric proton at δH 4.83 and the carbon signal at δC 87.2 (d, C-3). During the hydrolysis process of 8, jinfushanosides B-D were detected in reaction mixture by TLC. The configurations of the anomeric protons of the three glucoses were established to be β on the basis of the coupling constants of the anomeric protons, and the sugars were determined to be a D-glucose by comparison with authentic sample on GC analysis after acid hydrolysis of 8.
Jinfushanoside K (9) displayed a quasimolecular ion peak at m/z=927.5336 [M - H]-(cald for C48H77O17, 927.5317), consistent with the molecular formula C48H78O17. The molecular weight of 9 was 14 mass units less than that of carnosifloside Ⅲ (12).8 The 13C NMR data (Tables 2 and 4) of 9 was very similar with that of 12 with the exception of a methylene (βC 32.7) instead of the carbonyl functionality (βC 213.7) at C-11 in 9, in accordance with the disappearance of the AB coupling system of H-12 in 1H NMR spectrum. In HMBC spectrum, correlations observed from δH 0.85 (3H, s, H-19) to δC 44.0 (d, C-8), 38.7 (d, C-10), 35.1 (s, C-9), and 32.7 (t), also proved the presence of a methylene at C-11. The configurations of the anomeric protons of the three glucoses were established to be β on the basis of the coupling constants of the anomeric protons, and the sugars were determined to be a D-glucose by comparison with authentic sample on GC analysis after acid hydrolysis of 9. Exception of ring B, the other parts of 9 were identical to those of 12 by detailed analysis of the 13C, 1H, HMBC, HSQC, and 1H-1H COSY spectra of 9. Compound 9 was also the first occurrence of cucurbitane triterpene without carbonyl or hydroxyl functionalities at C-11 from the genus Hemsleya.
Known compounds were identified as carnosiflogenin A (10), 8 delavanoside D (11), 9 carnosifloside Ⅲ (12), 8 3β, 27-dihydroxycucurbita-5, 24-dien-11-on-3-O-β-D-glucopyranosyl-27-O-β-D-glucopyranoside (13), 8 delavanoside A (14), 9 11α, 26-dihydroxycucurbita-5, 24-dien-26-O-β-D-glucopyranosyl(1→6)-β-D-glucopyranoside (15), 8 carnosifloside Ⅵ (16), 8 hexanorcucurbitacin F (17), 12 and scandenogenin A (18)10 by comparison with literature data. Among the known compounds, 13, 15, and 18 were firstly isolated as natural products, which had been obtained by acid hydrolysis of carnosifloside Ⅳ, 8 carnosifloside Ⅳ, 8 and scandenoside R1, 11 respectively.
Compounds 2, 3, 5, 7, 8, 9, 13, and 15, were tested for in vitro inhibitory effects against HIV replication in C8166 cells. All the compounds were no or weak active (Table 6).
Summary of cytotoxicities and anti-HIV-1 activities of the tested compounds
Experimental Section
General Experimental Procedures. Melting points were obtained on an XRC-1 apparatus and are uncorrected. Optical rotations were carried out on a Perkin-Elmer model 241 polarimeter. UV spectrum was measured in a UV 210A spectrometer. IR spectra were measured in a Bio-Rad FTS-135 spectrometer with KBr pellets. MS were recorded on a Finnigan MAT 90 instrument. 1D and 2D NMR spectra were measured on a Bruker DRX-500 instrument with TMS as internal standard. Column chromatography was performed either on silica gel (200-300 mesh; Qingdao Marine Chemical Inc., Qingdao, China), or Lichroprep RP-18 gel (40-63 μm; Merck, Darmstadt, Germany). Fractions were monitored by TLC, and spots were visualized by heating silica gel plates sprayed with 15% H2SO4 in H2O.
Plant Material. The tubers of H.penxianensis were collected in Jinfu Mountain, Congqing City, China, in 2000. A voucher specimen (No. KIB 2000-9-4) has been deposited at the State Key Laboratory of Phytochemistry and Plant Resources in West China, Kunming Institute of Botany, Chinese Academy Sciences. The plant material was identified by Prof. Wen-Jin Zhan, Penzhou Institute for Pharmaceutical Control, Sichuan.
Extraction and Isolation. Air-dried and powdered tubers (2.0 kg) of H. penxianensis were extracted with 95% ethanol under reflux (3 × 8 L), and filtered. After concentration of the combined filtrate under vacuum, 411 g residue was gotten. The residue (322 g) was absorbed on silica gel and chromatographed on silica gel column, eluting with a gradient system of CHCl3, CHCl3-Me2CO (15:1, 9:1), and CHCl3-EtOH (9:1, 7:3) to give six fractions (Frs. 1-6). Fraction 3 (2.83 g) was fractioned on silica gel, developing with a gradient system of CHCl3-MeOH (20:1, 15:1, 12:1) to furnish three parts (Fr. 3.1-3.3). Fr. 3.3 (1.1 g) was rechromatographed on silica gel, eluting with CHCl3-CH3OH (15:1, 12:1), then purified on LH-20 using MeOH as eluent to give 1(12 mg), 2 (57 mg), 10 (17 mg), 17 (32 mg), and 18 (15 mg). Fr. 4 (12.8 g) was subjected to CC (20 g) to generate four fractions (Frs. 4.1-4.4). Fr. 4.3 (8.43 g) was repeatedly subjected to silica gel column eluting with CHCl3-CH3OH(10:1) and RP18 silica gel developing with aqueous MeOH (60%→70%), to generate 3 (174 mg), 4 (19 mg), 5 (47 mg), 6 (23 mg), and 11 (86 mg). Compounds 13 (42 mg), 14 (31 mg), and 15 (19 mg) were isolated from Fr.4.4 (1.5 g) by subjected to column chromatography over silica gel (CHCl3-MeOH, from 6:1 to 9:1) and RP-18 (aqueous MeOH, from 55% to 60%), then Sephadex LH-20 (MeOH). Fr. 5 (131 g) was rechromatographed on silica gel developing with a gradient system of CHCl3-MeOH-H2O (8:1:0.1, 8:2:02, 7:3:1, 7:3:0.5) to yield Fr. 5.1-Fr. 5.5. Fr.5.3 (9 g) was further rechromatographed over RP18 silica gel developing with aqueous MeOH (40%→50%) to afford compounds 9 (39 mg), 12 (451 mg), and 16 (320 mg). Compounds 7 (150 mg) and 8 (164 mg) were isolated from Fr.5.4 (4 g) by subjected to column chromatography over RP-18 silica gel, eluting with a gradient aqueous MeOH system from 40% to 45%.
Jinfushanencin A (1): colorless powder; mp 217-219℃; [α]D20+22.0 (c 0.05, MeOH); IR (KBr)vmax: 3436, 2973, 1696, 1648, 1302 cm–1; 1H and 13C NMR data, see Tables 1 and 2; negative FABMS: m/z 533 [M - H]–; negative HRESIMS: m/z 533.3145 [M - H]–(calcd for C30H45O8, 533.3114).
Jinfushanencin B (2): colorless powder; mp 149-151℃; [α]D20+15.8(c 0.12, MeOH); IR (KBr)vmax: 3372, 2952, 2929, 2874, 1693, 1464, 1382, 1028, 1006, 979 cm–1; 1H and 13C NMR data, see Tables 1 and 2; negative FABMS: m/z 471 [M - H]–; negative HRESIMS: m/z 471.3487 [M - H]–(calcd for C30H47O4, 471.3474).
Jinfushanoside E (3): white powder; [α]D20+ 8.6(c 0.36, MeOH); IR (KBr)vmax: 3425, 2962, 2878, 1728, 1633, 1383, 1206, 1078, 978 cm–1; 1H and 13C NMR data, see Tables 1-5; negative FABMS: m/z 761 [M - H]–; negative HRESIMS: m/z 761.4491 [M - H]– (cald for C42H65O12, 761.4476).
Jinfushanoside F (4): white powder; [α]D20+9.2(c 0.24, MeOH); IR (KBr)vmax: 3424, 3369, 2962, 2877, 1726, 1692, 1383, 1207, 1077, 1038 cm–1; 1H and 13C NMR data, see Tables 1-5; negative FABMS: m/z 777 [M - H]–; negative HRESIMS: m/z777.4436 [M - H]– (cald for C42H65O13, 777.4425).
Jinfushanoside G (5): white powder; [α]D20+12.0(c 0.09, MeOH); IR (KBr) vmax: 3409, 3368, 2942, 2928, 1638, 1464, 1381, 1075, 1030 cm–1; 1H and 13C NMR data, see Tables 1-4; negative FABMS: m/z 619 [M - H]–; negative HRESIMS: m/z 619.4183 [M - H]– (cald for C36H59O8, 619.4209).
Jinfushanoside H (6):white powder; [α]D20+8.1 (c 0.09, MeOH); IR (KBr) vmax: 3431, 2928, 2874, 1722, 1453, 1076, 981 cm–1; 1H and 13C NMR data, see Tables 1-5; negative FABMS: m/z 763 [M - H]–; negative HRESIMS: m/z 763.4648 [M - H]–(cald for C42H67O12, 763.4632).
Jinfushanoside I (7): white powder; [α]D20+5.9 (c 0.09, MeOH); IR (KBr) vmax: 3431, 2945, 2875, 1729, 1579, 1458, 1076, 1021 cm–1; 1H and 13C NMR data, see Tables 1-5; negative FABMS: m/z 1085 [M - H]–; negative HRESIMS: m/z 1085.5546 [M - H]– (cald for C54H85O22, 1085.5532).
Jinfushanoside J (8): white powder; [α]D20+12.6 (c 0.11, MeOH); IR (KBr) vmax: 3452, 3283, 2926, 2877, 1690, 1467, 1075, 1020 cm–1; 1H and 13C NMR data, see Tables 1-4; negative FABMS: m/z 957 [M - H]–; negative HRESIMS: m/z 957.5050 [M - H]– (cald for C48H77O19, 957.5059).
Jinfushanoside K (9): white powder; [α]D20-5.3 (c 0.17, MeOH); IR (KBr) vmax: 3409, 2935, 2872, 1640, 1460, 1379, 1075, 1038 cm–1; 1H and 13C NMR data, see Tables 1-4; negative FABMS: m/z 927 [M - H]–; negative HRESIMS: m/z 927.5336 [M - H]-(cald for C48H77O17, 927.5317).
Acid Hydrolysis of 5, 8, and 9: Each of the compounds 5, 8, and 9 (2 mg) dissolved in 2 mL HCl-CH3OH (1 mol L–1 HCl-CH3OH, 1:1, v/v) was placed in water bath at 90℃ for 6 hrs. After the hydrolysis was finished, the water-soluble part was evaporated to dryness. The dry sugar residue and authentic sample of D-(+)-glucose (Sigma Company, G5250) were respectively diluted in 1 ml pyridine without water and treated with 0.4 mL hexamethyl disilazane and 0.2 mL trimethylchlorosilane (TMCS, Fluka) at 0℃. After that, the upper layer of the reaction mixture was analyzed by GC. (GC condition: AC-5 capillary column (30 m × Ø0.25 mm); column temperature: 180-260℃; column head pressure: 12 Pa; carrier gas: N2). GC of all samples showed same retention time, Rt (m): 11.521.
Partial Alkaline Hydrolysis of 3, 4, 6, and 7: A solution of the triterpene glycoside (each 0.5 mg dissolved in 1 ml MeOH) was added 0.5 % aq. NaOH (20 mL). After 2 h shaking at room temperature, the reaction mixture was acidified to PH 5 (5 M HCl) and was concentrated to dry. The resulting residue was dissolved in MeOH, and then was checked by using TLC. As a result, delavanoside D (11), jinfushanoside B, jinfushanoside G (5), and carnosifloside Ⅲ (12), were detected in the reaction mixture of 3, 4, 6, and 7, by TLC, respectively.
Anti-HIV-1 and Cytotoxicity Assay. The anti-HIV-1 activity was evaluated by the inhibition assay for the cytopathic effects of HIV-1 (EC50), and cytotoxicity assay against C8166 cells (IC50) was assessed using the MTT method as described in the literature.13
Notes
Acknowledgments
This work was financially supported by the Natural Science Foundation of Yunnan Province (2008CD158), the Western Doctoral Foundation of Chinese Academy of Sciences (J.C. Chen), the Foundation of State Key Laboratory of Phytochemistry and Plant Resources in West China, Kunming Institute of Botany, Chinese Academy of Sciences (P2008ZZ23, P2010ZZ14), and the Cooperative Project of Guangdong Province and CAS (2009B091300135).
References
-
1.R. L. Nie, Acta Bot. Yunnan. 16, 201-208 (1994) PubMed Google Scholar
-
2.W. J. Zhang, L. D. Shen, Acta Phytotax. Sin. 21, 182-193 (1983) PubMed Google Scholar
-
3.Li, D. Z. Systematics and Evolution of Hemsleya (Cucurbitaceae); Yunan Science and Technology Press:Kunming, 1993:pp 89. PubMed Google Scholar
-
4.J. C. Chen, X. M. Niu, Z. R. Li, M. H. Qiu, Planta Med. 71, 983-986 (2005) CrossRef PubMed Google Scholar
-
5.R. R. Tian, J. C. Chen, G. H. Zhang, M. H. Qiu, Y. H. Wang, L. Du, X. Shen, N. F. Liu, Y. T. Zheng, Chin. J. Nat. Med. 6, 214-218 (2008) CrossRef PubMed Google Scholar
-
6.J. C. Chen, M. H. Qiu, R. L. Nie, G. A. Cordell, S. X. Qiu, Nat. Prod. Rep. 22, 386-399 (2005) CrossRef PubMed Google Scholar
-
7.D. C. Chaves, J. C. C. Assunção, R. Braz-Filho, F. J. Q. Monte, Magn. Reson. Chem. 45, 389-392 (2007) CrossRef PubMed Google Scholar
-
8.R. Kasai, K. Matsumoto, R. L. Nie, T. Morita, A. Awazu, J. Zhou, O. Tanaka, Phytochemistry 26, 1371-1376 (1987) CrossRef PubMed Google Scholar
-
9.J. C. Chen, Z. Q. Zhang, M. H. Qiu, Acta Chim. Sinica 65, 1679-1684 (2007) PubMed Google Scholar
-
10.Bernardi M. De, G. Fronza, M. P. Gianotti, G. Mellerio, G. Vidari, P. Vita-Finzi, Tetrahedron Lett. 24, 1635-1638 (1983) CrossRef PubMed Google Scholar
-
11.R. Kasai, K. Matsumoto, R. L. Nie, J. Zhou, O. Tanaka, Chem. Pharm. Bull. 36, 234-243 (1988) CrossRef PubMed Google Scholar
-
12.X. Fang, C. H. Phoebe, Pezzuto J. M. Jr, H. H. Fong, N. R. Farnsworth, B. Yellin, S. M. Hecht, J. Nat. Prod. 47, 988-993 (1984) CrossRef PubMed Google Scholar
-
13.G. H. Zhang, Q. Wang, J. J. Chen, X. M. Zhang, S. C. Tam, Y. T. Zheng, Biochem. Biophys. Res. Commun. 334, 812-816 (2005) CrossRef PubMed Google Scholar
Copyright information
© The Author(s) 2012
Open Access
This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.