b Laboratory for Marine Drugs and Bioproducts of Qingdao National Laboratory for Marine Science and Technology, Qingdao 266237, China
Cancer is predicted to be the most important barrier to increasing life expectancy in the 21st century [1, 2]. Almost onesixth of deaths resulted from cancer, accounting for 8.8 million globally [3]. Lung cancer is the leading cause of cancer death among males, followed by liver and stomach cancer [1]. Current treatments for cancer include chemotherapy, radiotherapy, surgery, palliative care and immunotherapy, however, the overall survival rate of cancer patients still remains low [4, 5]. Therefore, it is urgent to discover more effective agents to improve the overall survival rate of cancer patients.
Recently, α-helical peptides have emerged as a compelling new therapeutic modality to treat cancer [6]. Venoms of hymenopteran insects are one of the important sources of a-helical peptides. A new hexadecapeptide sequence (GIMSSLMKKLAAHIAK) named HYL has been discovered in the venom of the solitary bee Hylaeus signatus (Hymenoptera: Colletidae), which belongs to α-helical peptides [7]. Nesuta et al. prepared a set of HYL analogues to increase its antimicrobial activity by lysine (Lys) and hydrophobic amino acid substitution strategies, and several analogues exhibited significantly greater antimicrobial activity compared to HYL [7]. The helical wheel projection of HYL is showed in Fig. 1 [8]. We synthesized the peptide HYL by a solid-phase synthesis method and it exhibited significant antitumor activity against human nonsmall cell lung adenocarcinoma A549 (10.01 μmol/L), human hepatic adenocarcinoma HepG2 (14.71 μmol/L) and human colorectal adenocarcinoma HCT116 (22.89 μmol/L) in vitro. However, the natural peptides have poor proteolytic stability and easily lose their biologically relevant conformation when crossing membranes and entering the cellular environment [9]. Therefore, we expect to further improve the antitumor activity, conformational rigidity and proteolytic stability of HYL by structural modification.

|
Download:
|
| Fig. 1. The helical wheel presentation of HYL. The yellow points to the hydrophobic residues, and the blue points to the positively charged hydrophilic residues. The purple points to non-charged polar residues, the light blue points to the histidine (His) and the gray represents other residues. | |
Stapled α-helical peptides have experienced exponential growth in recent years, mostly due to their potential as drug candidates and biological tools [10-12]. The helical conformation of stapled peptides can be stabilized by a side-chain to side-chain covalent linker [13, 14]. Generally, stabilization of helix has been fulfilled through side-chain constraints by covalent connection of two side-chain residues with Ru-catalyzed olefin metathesis, lactam-bridging, disulfides, lactams, triazoles and others [15-24]. Among them, the all-hydrocarbon stapled peptide strategy developed by Verdine et al. has been considered as a promising stapling strategy to improve the activity and property of peptides [25, 26]. Our group improved the antitumor activity and proteolytic stability of host-defense peptide hymenochirin-1B effectively by using the hydrocarbon stapling strategy [27, 28].
Here, we report the synthesis of HYL by a solid-phase synthesis method and its structural modification by alanine scanning and hydrocarbon stapling strategies to improve its antitumor activity and proteolytic stability.
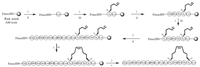
Firstly, we synthesized HYL by using a solid-phase synthesis method and found it had significant antitumor activity (Fig. S1 in Supporting information). Then, we further synthesized HYL-1 to HYL-13 (Table 1) by alanine scanning strategy to determine the key residues of HYL. To improve the antitumor activity and proteolytic stability of HYL, we also synthesized its analogues HYL-14~HYL-18 by using hydrocarbon stapling strategy without destroying the key residues. As shown in Scheme 1, the all-hydrocarbon stapled peptides were prepared using an Fmoc solid-phase peptide synthesis procedure with Rink Amide MBHA resin as the solid support [28]. 5-Chloro-1-[bis(dimethylamino)methylene]-1H-benzotriazolium 3-oxide hexafluorophosphate (HCTU) was used as a coupling reagent. 2-(7-Azabenzotriazol-1-yl)-N, N, N', N'-tetramethyluronium hexafluorophosphate (HATU), 1-hydroxy-7-azabenzotriazole (HOAT) and N, N-diisopropylethylamine (DIEA) were used for coupling of (S)-N-Fmoc-2-(4-pentenyl)alanine (S5). After the linear peptides assembly was completed, the olefincontaining peptides were stapled using Grubbs' first-generation catalyst. The peptides were cleaved off from the resin and globally deprotected with reagent K (TFA/H2O/EDT/thioanisole/phenol = 82.5:5:2.5:5:5).
|
|
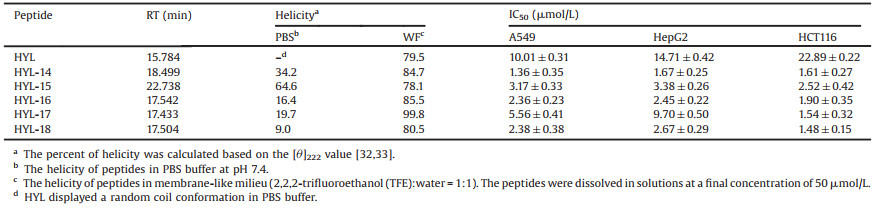
Table 1 Amino acid sequences, MS data and physical properties of HYL and its analogues.a |
|
Download:
|
| Scheme 1. Solid-phase synthesis of stapled peptide HYL-14. Conditions: (ⅰ) 20% piperidine in DMF 5 min (2 times), 35 ℃; (ⅱ) Fmoc-AA-OH (4 equiv.)/HCTU (4 equiv.)/DIEA (4 equiv.), 1 h, 35 ℃; (ⅲ) Fmoc-S5-OH (4 equiv.)/HATU (4 equiv.)/HOAT (4 equiv.)/DIEA (4 equiv.), 2 h, 35 ℃; (ⅳ) 6 mmol/L 1st Grubbs', catalyst, DCE, 2 h, 35 ℃; (ⅴ) reagent K (TFA/ H2O/EDT/thioanisole/phenol = 82.5:5:2.5:5:5), 3 h, 35 ℃. | |
Precooled diethyl ether was added to precipitate crude peptides, and then peptides were purified by semi-preparative RP-HPLC. Matrix-assisted laser desorption/ionization time-of-fight mass spectroscopy (MALDI-TOF-MS) was employed to confirm the molecular weight of peptide, and the measured molecular weight of each peptide was consistent with the theoretically calculated value.
The growth-inhibitory activities of peptides were evaluated by using the 3-(4, 5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) assay against three cancer cells, including A549, HCT116 and HepG2 [29]. As shown in Table 2, we can determine the key residues, namely, isoleucine (Ile), leucine (Leu) and methionine (Met), which significantly affect the antitumor activity of HYL by comparing with the antitumor activity of the parent peptide HYL. They are all hydrophobic amino acids, suggesting that hydrophobicity has an important effect on the anti-tumor activity of HYL. Moreover, the retention time (RT) of HYL-1 to HYL-13 also proved the importance of hydrophobicity to antitumor activity of HYL, because of the longer the RT, the more hydrophobic the effect peptide is. We also found that the charge did not have a significant effect on the anti-tumor activity of HYL, however, the substitution of lysine (Lys) and histidine (His) enhanced the anti-tumor activity of HYL (HYL-9, HYL-11). This result may be due to the increase in hydrophobicity of HYL-9 and HYL-11. Replacement of Lys at different positions had different effects on the anti-tumor activity and RT of HYL. The antitumor activity and the RT of HYL, HYL-8 and HYL-13 showed no significant change, while HYL-9 showed improvement in both anti-tumor activity and RT. This indicates that Lys at different positions plays different roles in the whole peptide.
|
|
Table 2 The antitumor activity of HYL and its alanine scanning sequences. |
To improve the antitumor activity and proteolytic stability of HYL, we further designed a series of stapled peptides without destroying the key residues. We substituted pairs of S5 at the i and i +4 positions that accommodate one turn of the a-helix to obtain HYL-14~HYL-18 [30, 31]. All the stapled peptides showed growthinhibitory activity to 0.5–15.3 folds than that of the parent peptide HYL (Table 3) [32, 33]. By comparing the RT of stapled peptides HYL-14 to HYL-18, we find that hydrophobicity is an important reason for the increase of stapled peptides activity. By comparing RT and sequences of the stapled peptides HYL-14 and HYL-15, we find that amphiphilicity may be another important factor in determining anti-tumor activity.
|
|
Table 3 The α-helicity (%) and antitumor activity of HYL and stapled peptides. |
Circular dichroism (CD) spectroscopy was used to investigate the secondary structure and structural changes of HYL and the stapled peptides in PBS buffer and membrane-like milieu, respectively. The CD spectroscopy provides a typical signature for a-helices with a maximum near 190 nm and double minima at 208 nm and 222 nm, indicating that HYL displayed a random coil conformation in PBS buffer (Fig. 2A) [34]. It is worth mentioning that the stapled peptides HYL-14 to HYL-18 exhibited 9.0%–64.6% helicity in PBS buffer (Table 3), indicating that the hydrocarbon stapling strategy can effectively enhance the helicity of HYL in PBS buffer. In addition, there are significant differences in the helicity of stapled peptides, suggesting that stapling position is an important factor affecting the helicity of peptides in PBS buffer. By comparing the helicity of HYL and stapled peptides in PBS buffer and membrane-like milieu (Table 3 and Fig. 2), we find that different solution environment has significant influence on the helicity of HYL and its analogues, however, the helicity of the stapled peptides HYL-14 to HYL-18 showed no significant change in membrane-like milieu. In addition, there is no significant correlation between the antitumor activity and helicity by comparing the antitumor activity and helicity of HYL and the stapled peptides.

|
Download:
|
| Fig. 2. CD spectroscopy analysis of 50 μmol/L peptide at 37 ℃ in PBS buffer at pH 7.4 (A) and membrane-like milieu (B). | |
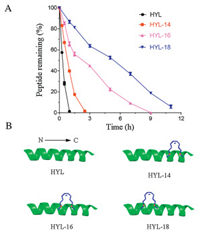
In order to further understand the role of stapling strategy, we compared the proteolytic stability of HYL and its analogues HYL-14, HYL-16 and HYL-18 by measuring their susceptibility toward chymotrypsin degradation at room temperature in PBS buffer (pH 7.4) with monitored by HPLC [35, 36]. In Fig. 3A, we can see that the half-life (t1/2) of HYL is 0.36 h. As expected, hydrocarbon-stapled peptides possessed higher protease resistance and the t1/2 of HYL- 14, HYL-16 and HYL-18 is 0.77 h, 2.23 h and 5.27 h, respectively. These results demonstrated the obvious superiority of the hydrocarbon-stapled peptides over the linear peptides with respect to protease resistance. Meanwhile, we also found that the stapling position near the N-terminal was more conducive to the improvement of the protease resistance than that of the Cterminal by comparing the protease resistance of the stapled peptides HYL-14 to HYL-18 (Fig. 3 and Fig. S2 in Supporting information).
|
Download:
|
| Fig. 3. (A) The proteolytic stability of HYL, HYL-14, HYL-16 and HYL-18 incubated in chymotrypsin solution (0.5 ng/mL in 50 mmol/L PBS buffer, pH 7.4) at a final concentration of 0.1 mmol/L. Date points are displayed as the mean value SEM of three independent experiments. The percent of residual peptide was monitored by analytic HPLC. (B) Structures of the linear template peptide HYL and stapled peptides HYL-14, HLY-16 and HYL-18. The green represents peptide chain, the blue points to the hydrophobic carbon chains. | |
To further research the selectivity of these peptides on cancer cells, we evaluated the growth-inhibitory activities of HYL, HYL-14, HYL-16 and HYL-18 against two normal cells, namely, human umbilical vein endothelial cell (HUVEC) and human normal kidney cell 293T (Table S2 in Supporting information) [29]. The experiments procedure was performed just same as the growth inhibition of peptides against tumor cells. The results showed that the stapled peptides increased in normal cells toxicity, but they still remained the selectivity towards cancer cells over normal cells. The increased toxicity might be due to the high hydrophobicity produced by six additional methylene and two olefinic carbons to increase their toxicity to human normal cells. However, this situation may be resolved by addition of polar character or glycosylation into the sequence as our previous work [28]. The modification of stapled peptides is ongoing, and the present study mainly focused on the improvement of anti-tumor activity of parent peptide HYL.
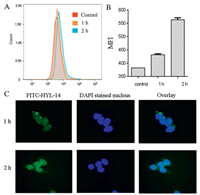
To explore the action mechanism of stapled peptide HYL-14, we used fluorescein isothiocyanate (FITC)-labeled HYL-14 to study its localization in HCT116 cells [37]. As shown in Figs. 4A and B, the flow cytometry assay suggests that the stapled peptide HYL-14 continued to enter the cell within two hours. Also, FITC-HYL-14 was significantly concentrated in the cell nucleus after 2 h, indicating that HYL-14 can enter into the nucleus quickly (Fig. 4C). However, the action mechanism of HYL-14 needs further study.
|
Download:
|
| Fig. 4. (A) Peptide uptake by the cancer cells HCT116. The peptide FITC-HYL-14 was incubated with the cells for 1 h, 2 h at 37 ℃. The data are expressed as the mean ± SD of three experiments. (B) Mean fluorescence intensity (MFI) of FITC positive cells. (C) The localization of HYL-14 in HCT116 cells. HCT116 cells were treated for 1 h or 2 h with 0.1 μmol/L FITC-labeled HYL-14. Images from left to right include FITC channel, DAPI stained nucleus, and FITC/DAPI overlay. | |
We reported the antitumor activity of antibacterial peptide HYL for the first time, and identified its key residues on antitumor activity by alanine scanning strategy. Based on these findings, we designed and synthesized 18 peptides and found that the hydrocarbon stapling strategy effectively improved the antitumor activity, helicity and proteolytic stability of HYL. The stapled peptides HYL-14, HYL-16 and HYL-18 show a promising prospect for novel anti-tumor drug development.
AcknowledgmentsThis research was supported by NSFC-Shandong Joint Fund (No. U1606403) and Innovation Project of Qingdao National Laboratory for Marine Science and Technology (No. 2015ASKJ02). We are grateful to the Instrumental Analysis Center of Ocean University of China for NMR spectroscopic and mass spectrometric analysis.
Appendix A. Supplementary dataSupplementary material related to this article can be found, in the online version, at doi:https://doi.org/10.1016/j.cclet.2019.10.013.
| [1] |
F. Bray, J. Ferlay, I. Soerjomataram, et al., CA Cancer J. Clin. 68 (2018) 394-424. DOI:10.3322/caac.21492 |
| [2] |
C.E. DeSantis, C.C. Lin, A.B. Mariotto, et al., CA Cancer J. Clin. 64 (2014) 252-271. DOI:10.3322/caac.21235 |
| [3] |
R.L. Siegel, K.D. Miller, A. Jemal, CA Cancer J. Clin. 68 (2018) 7-30. DOI:10.3322/caac.21442 |
| [4] |
R. Shashidharamurthy, E.N. Bozeman, J. Patel, et al., Med. Res. Rev. 32 (2012) 1197-1219. DOI:10.1002/med.20237 |
| [5] |
M.M. Wei, Y.S. Wang, X.S. Ye, Med. Res. Rev. 38 (2018) 1003-1026. DOI:10.1002/med.21493 |
| [6] |
T.K. Sawyer, A.W. Partridge, H.Y.K. Kaan, et al., Bioorg. Med. Chem. 26 (2018) 2807-2815. DOI:10.1016/j.bmc.2018.03.008 |
| [7] |
O. Nesuta, R. Hexnerova, M. Budesinsky, et al., J. Nat. Prod. 79 (2016) 1073-1083. DOI:10.1021/acs.jnatprod.5b01129 |
| [8] |
R. Gautier, D. Douguet, B. Antonny, G. Drin, Bioinformatics 24 (2008) 2101-2102. DOI:10.1093/bioinformatics/btn392 |
| [9] |
B. Villavicencio, R. Ligabue-Braun, H. Verli, J. Chem. Inf. Model. 58 (2018) 2015-2023. DOI:10.1021/acs.jcim.8b00404 |
| [10] |
A.A. Vinogradov, Y.Z. Yin, H. Suga, J. Am. Chem. Soc. 141 (2019) 4167-4181. DOI:10.1021/jacs.8b13178 |
| [11] |
X. Qin, H. Zhao, Y.H. Jiang, et al., Chin. Chem. Lett. 29 (2018) 1160-1162. DOI:10.1016/j.cclet.2018.04.004 |
| [12] |
Y. Guo, L.L. Fu, X.W. Fan, X.L. Shi, Chin. Chem. Lett. 29 (2018) 1167-1170. DOI:10.1016/j.cclet.2018.03.024 |
| [13] |
G.L. Verdine, G.J. Hilinski, Methods Enzymol. 503 (2012) 3-33. DOI:10.1016/B978-0-12-396962-0.00001-X |
| [14] |
Y.H. Lau, P.D. Andrade, Y.T. Wu, D.R. Spring, Chem. Soc. Rev. 44 (2015) 91-102. DOI:10.1039/C4CS00246F |
| [15] |
G. Lautrette, F. Touti, H.G. Lee, P. Dai, B.L. Pentelute, J. Am. Chem. Soc. 138 (2016) 8340-8343. DOI:10.1021/jacs.6b03757 |
| [16] |
Y. Tian, J.X. Li, H. Zhao, et al., Chem. Sci. 7 (2016) 3325-3330. DOI:10.1039/C6SC00106H |
| [17] |
Y. Wu, Y.H. Li, W. Li, et al., Chem. Sci. 8 (2017) 7368-7373. DOI:10.1039/C7SC02420G |
| [18] |
X.D. Shi, Y.X. Jiang, D. Yang, et al., Chin. Chem. Lett. 29 (2018) 485-488. DOI:10.1016/j.cclet.2017.07.003 |
| [19] |
P. Diderich, D. Bertoldo, P. Dessen, et al., ACS Chem. Biol. 11 (2016) 1422-1427. DOI:10.1021/acschembio.5b00963 |
| [20] |
X. Li, Y. Zou, H.G. Hu, Chin. Chem. Lett. 29 (2018) 1088-1092. DOI:10.1016/j.cclet.2018.01.018 |
| [21] |
C.M. Haney, H.M. Werner, J.J. McKay, W.S. Horne, Org. Biomol. Chem. 14 (2016) 5768-5773. DOI:10.1039/C6OB00475J |
| [22] |
Y.H. Li, K.A. Clark, Z.P. Tan, Chin. Chem. Lett. 29 (2018) 1074-1078. DOI:10.1016/j.cclet.2018.05.027 |
| [23] |
X. Lu, S.J. He, W.M. Cheng, J. Shi, Chin. Chem. Lett. 29 (2018) 1001-1008. DOI:10.1016/j.cclet.2018.05.011 |
| [24] |
Y. Jiang, H.Y. Long, Y.J. Zhu, Y. Zeng, Chin. Chem. Lett. 29 (2018) 1067-1073. DOI:10.1016/j.cclet.2018.05.028 |
| [25] |
F. Bernal, A.F. Tyler, S.J. Korsmeyer, L.D. Walensky, G.L. Verdine, J. Am. Chem. Soc. 129 (2007) 2456-2457. DOI:10.1021/ja0693587 |
| [26] |
G.J. Hilinski, Y.W. Kim, J. Hong, et al., J. Am. Chem. Soc. 136 (2014) 12314-12322. DOI:10.1021/ja505141j |
| [27] |
Y.L. Li, M.H. Wu, Q. Chang, X. Zhao, RSC Adv. 8 (2018) 22268-22275. DOI:10.1039/C8RA03446J |
| [28] |
Y.L. Li, Y.H. Zhang, M.H. Wu, et al., ACS Chem. Biol. 14 (2019) 516-525. DOI:10.1021/acschembio.9b00046 |
| [29] |
B. Li, W. Gu, C. Zhang, et al., Onkologie 29 (2006) 367-371. DOI:10.1159/000094711 |
| [30] |
L.K. Henchey, A.L. Jochim, P.S. Arora, Curr. Opin. Chem. Biol. 12 (2008) 692-697. DOI:10.1016/j.cbpa.2008.08.019 |
| [31] |
K. Hojo, M.A. Hossain, J. Tailhades, et al., J. Med. Chem. 59 (2016) 7445-7456. DOI:10.1021/acs.jmedchem.6b00265 |
| [32] |
H.K. Cui, J. Qing, Y. Guo, et al., Bioorg. Med. Chem. 21 (2013) 3547-3554. DOI:10.1016/j.bmc.2013.02.011 |
| [33] |
N.E. Shepherd, H.N. Hoang, G. Abbenante, D.P. Fairlie, J. Am. Chem. Soc. 127 (2005) 2974-2983. DOI:10.1021/ja0456003 |
| [34] |
C. Wang, S. Xia, P. Zhang, et al., J. Med. Chem. 61 (2018) 2018-2026. DOI:10.1021/acs.jmedchem.7b01732 |
| [35] |
T.E. Speltz, C.G. Mayne, S.W. Fanning, et al., Org. Biomol. Chem. 16 (2018) 3702-3706. DOI:10.1039/C8OB00790J |
| [36] |
Y. Tal-Gan, M. Ivencic, G. Cornilescu, T. Yang, H.E. Blackwell, Angew. Chem. Int. Ed. 55 (2016) 8913-8917. DOI:10.1002/anie.201602974 |
| [37] |
Y.S. Chang, B. Graves, V. Guerlavais, et al., Proc. Natl. Acad. Sci. U. S. A 110 (2013) E3445-54. DOI:10.1073/pnas.1303002110 |
 2020, Vol. 31
2020, Vol. 31