b College of Pharmaceutical Sciences, Zhejiang University, Hangzhou 310058, China;
c School of Medicine, Zhejiang University, Hangzhou 310058, China
In spite of recent advances in cancer therapy, breast cancer remains to be one of most threatening diseases to the females [1-3]. Over the past few decades, the traditional chemotherapeutic drugs, such as paclitaxel (PTX) and doxorubicin [4], have been the first-line treatment for breast cancer in clinical practice; however, the serve side effects and poor pharmacological effects have significantly comprised the therapeutic effects in vivo [5, 6]. Furthermore, breast cancer is also highly invasive and metastatic [7], which poses additional barriers for its therapy. To overcome these issues, some recent studies reveal that the combination of chemotherapy and RNA interference (RNAi) is a promising strategy to improve the efficacy of current chemotherapies [8-12]. By silencing unwanted gene expression, siRNA-based therapy is proven to be an exceptionally appealing strategy to inhibit tumor growth and metastasis in combination of chemotherapeutic drugs [13]. As a negatively charged nucleic acid, naked siRNA is difficult to induce effective gene silencing, primarily owing to the poor cellular uptake caused by the electrostatic repulsion against the negatively charged cellular membranes [14-16]. Furthermore, the major challenge for the combination of chemotherapy and RNAibased therapy is the requirement of safe and efficient deliver carriers that can simultaneously co-deliver chemotherapeutic drugs and nucleic acids [17, 18]. Therefore, rational design of codelivery vectors is of paramount importance in achieving effective combination cancer therapy.
Chondroitin sulfate (CS) is one of glycosaminoglycans and plays an important role in many essential biological processes, such as cell proliferation, migration and development [19-21]. Recently, CS has been found to interact with overexpressed membrane receptors on the surface of breast cancer cells [22]. As a well-known biomarker of breast cancer cells, CD44 receptors are documented to strongly interact with CS, and are therefore believed to be an ideal target for breast cancer therapy. In addition, CS also acts as a substrate for P-selectins, which facilitate the inhibition of tumor metastasis [23]. In fact, CS has been widely used as a building block for constructing drug delivery carrier, owing to outstanding biocompatibility, good biodegradability, and extremely low toxicity [24-27]. Therefore, these characteristics of CS make it as a potential candidate for constructing functional nanoparticle on drug delivery system.
In the current study, we developed a new delivery system that can target CD44 receptors for the co-deliver chemotherapeutics and siRNA to improve the anticancer effect and inhibit the metastasis for breast cancer treatment [28, 29]. We combine PTX and siRNAtargetingCD146 for thecombination treatmentof breast cancer. CD146 is known as a potential marker for the prognosis of cancer metastasis [30]; however, the recent study provided evidence that CD146 is a downstream signal of CD44, and the high expression of CD146 promote breast cancer growth and metastasis [30]. The small interferingRNA (siRNA) targetingCD146 have been proved to be effective inhibit CD146 expression, and significantly inhibit the metastasis of breast cancer cells to the treatment of chemotherapeutics [31]. This suggests the delivery of siRNA targeting CD146 is likely to enhance the efficacy of breast cancer chemotherapy.
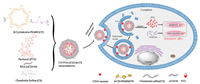
As shown in Scheme 1, this nanosystem is composed of supramolecular complex β-CyD-PEI600 (CP), a cationic polymer that has been widely explored to deliver nucleic acids (including both DNA and siRNA) in our previous studies [32-36]. Encapsulate PTX and siRNA (siCD146), in brief, a chemotherapeutics PTX as an anticancer drug to inhibit the survival of cancer cells, and siCD146 as an inhibitor to realize cancer anti-metastasis. In order to improve the anti-metastatic effect and reduce side effects, the CS is coated on the surface of this nanosystem and serve as a targeting group to recognize CD44-overexpressed breast cancer cells. This CD44-targeted nanosystem of anticancer drug and metastasis inhibitor for combination therapy display great promises to effectively inhibit cancer cell growth and metastasis.

|
Download:
|
| Scheme 1. Schematic overview of the co-delivery nanosystem for breast cancer cell apoptosis and metastasis. | |
The β-CyD has been widely exploited as a drug delivery carrier to load hydrophobic drugs via the host-guest interaction. As shown in Fig. S1 (Supporting information), cationic CP was first synthesized according to our previous methods [14, 35], β-CyD and polyethyleneimine (PEI, 600 Da) were linked by carbamate bond through CDI-mediate coupling reaction to obtain CP. Due to the host-guest interaction of the β-CyD and hydrophobic drug PTX, the supramolecular complexation between CP and PTX was obtained by continuous stirring for 24 h in a mixed solvent (MeOH/H2O, 1:1). The chemical structures of CP and CP-PTX were characterized by 1H NMR. As shown in Fig. S2 (Supporting information), the 1H NMR spectrum indicated that the molar ratio of PEI to β-CyD was approximately close to 1:1, which is calculated based on the integration of the broad peaks at δ 2.3–3.0 (-CH2-CH2-NH-) in PEI and the integration of a single peak at δ 4.9 (-CH) of methyl group from β-CyD. In Fig. S3 (Supporting information), the proton signals at δ 7.2–8.4 appeared in the 1H NMR spectra of CP-PTX indicated the supramolecular formation of CP-PTX complexes. The CP-PTX-RNA-CS nanocomplexes were prepared via two-step methods. The CP-PTX was first incubated with siRNA to form CP-PTX-siRNA complexes, followed by the selfassembly between positively charged CP-PTX-RNA and negatively charged CS. For CP-PTX-RNA-CS nanocomplexes, the loading efficiency of PTX was calculated as 15.7%.
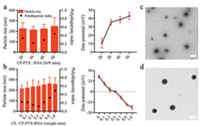
In order to investigate the whether siRNA was effectively loaded, the agarose gel electrophoresis was performed to check the electrostatic binding ability of CP-PTX with siRNA. Fig. S4 (Supporting information) showed that CP-PTX was able to bind siCD146 at the N/P ratios above 4, which revealed that the CP-PTX could efficiently load the siRNA to form CP-PTX-RNA complexes. In addition, the size distribution, zeta potential and morphology of CP-PTX-RNA nanocomplex were examined by using DLS and TEM, respectively. As shown in Fig. 1a, with the increase of N/P ratio from 20 to 50, the average diameter of CP-PTX-RNA slightly ranged from 210 nm to 250 nm. Note that the size distribution was slightly polydispersed, as indicated by polydispersity index (PDI) at different N/P ratios. In addition, the zeta potential of nanocomplexes increased from 11 mV to 42 mV with the increased N/P ratio. Owing to the low PDI (0.21) and suitable surface charge (35.6 mV), we decided to choose N/P ratio of 30 for CS coating. As shown in Fig. 1b, whereas the size of complexes slightly increased at different weight ratios (CS to CP-PTX-RNA), the zeta potential of CP-PTX-siRNA nearly achieved neutral at the weight ratio of 0.5. As shown in Figs. 1c and d, the TEM demonstrated that the morphology of CP-PTX-RNA and CP-PTX-RNA-CS were the regular spherical structure and the size was about 200 nm that similar with the data by DLS.
|
Download:
|
| Fig. 1. Characterization of the CP-PTX-RNA and CP-PTX-RNA-CS. Particle size, polydispersity index (left) and zeta potential (right) of CP-PTX-RNA (a) and CP-PTX-RNA-CS (b). (c, d) Visualization of CP-PTX-RNA (c) and CP-PTX-RNA-CS (d) by transmission electron microscopy (TEM). The scale bar of TEM represents 200 nm. | |
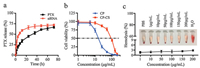
The release behavior of CP-PTX-RNA-CS was first investigated under physiological conditions (PBS, pH 7.4, 37 ℃). The release rate of PTX was monitored by UV spectroscopy. As shown in Fig. 2a, the release rate of PTX gradually increased with time and cumulative release reached 73% in 72 h, suggesting the disassembly of PTX from β-CyD moieties of CP. Furthermore, the release of siRNA (labeled with Cy5) was continuously monitored by fluorescence spectroscopy. The release kinetics exhibited a burst release pattern in the first 10 h, followed by sustained release. The cumulative release of siRNA reached 68% in 72 h. These results demonstrate that both siRNA and PTX can release from the CP-PTX-RNA-CS.

|
Download:
|
| Fig. 2. (a) PTX and siRNA release from CP-PTX-RNA-CS in PBS (pH 7.4). (b) Cytotoxicity of CP and blank CP-CS against MDA-MB-231 cells. The cytotoxicity was evaluated by MTT assay. Data represent mean ± SD (n = 6). (c) Blood hemolytic effect of CP-PTX-RNA-CS, the deionized water was used as a positive control, and PBS buffer solution was used as a negative control. Inset: RBCs treated with CP-PTX-RNA-CS nanocomplex of different concentrations, compared to the water and PBS groups. | |
An ideal co-delivery vector should not only be of high efficiency, but also possess low toxicity [32]. In order to evaluate the cytotoxicity of CP and CP-CS, MTT assays were first carried out in the breast cancer cell line (MDA-MB-231). As shown in Fig. 2b, both CP and CP-CS showed dose-dependent cytotoxicity, and the calculated half inhibitory concentration (IC50) value of CP-CS (91.2 μg/mL) was much higher than that of CP (19.7 μg/mL), indicating the introduction of CS could significantly improve the biocompatibility of CP. We also evaluated the cytotoxicity in human embryonic kidney cells (HEK293), and found that the CP-CS exhibited much higher IC50 value (104.3 μg/mL), in comparison with those in cancer cells (Fig. S5 in Supporting information). Therefore, the above results suggested that the CS coating layer significantly improved the biocompatibility of CP. The hemolysis rate represents the percentage of the destroyed membrane of red blood cells (RBCs) by a material at given concentration, which reflects the hemocompatibility under physiological conditions. In Fig. 2c, the hemolytic activity of CP-PTX-RNA-CS was only less than 3% up to 200 μg/mL of CP-PTX-RNA-CS in RBCs solution, indicating its good hemocompatibility.
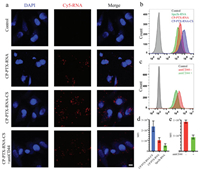
We next investigated the cellular uptake of CP-PTX-RNA-CS in MDA-MB-231 cells that overexpress CD44 by using confocal laser scanning microscopy. As shown in Fig. 3a, after CP-PTX-siRNA-CS nanocomplexes were incubated with MDA-MB-231 for 4 h, strong red fluorescence could be observed in the cytoplasm, indicating the successful intracellular delivery of CP-PTX-RNA-CS nanocomplexes. As a control, the nanocomplex CP-PTX-siRNA without targeting group CS exhibited relatively weaker intracellular fluorescence, in comparison with that of CP-PTX-RNA-CS group. In addition, we also quantitatively analyze the intracellular fluorescence intensity by flow cytometry. Figs. 3b and d showed that the level of cellular uptake of CP-PTX-RNA-CS was about 2.4- fold higher than that of CP-PTX-RNA. To further confirm whether high cellular uptake stem from overexpressed CD44 receptors, the MDA-MB-231 cells were first pre-treated with antiCD44 antibody to suppress the expression of CD44, and were then incubated with CP-PTX-RNA-CS complexes. The flow cytometry analysis indicated that the suppression of the CD44 receptors significantly lower intracellular accumulation of CP-PTX-RNA-CS (Figs. 3c and e), the same results in confocal was showed in Fig. 3a. Moreover, the MCF- 7 cells, a breast cancer cell line which do not overexpress CD44 receptors, was also used to assess the cellular uptake. Obviously, the red fluorescence from CP-PTX-RNA-CS was also weaker under the similar experimental conditions (Fig. S6 in Supporting information). The above results clearly indicate that the CS can improve the cellular uptake of nanocomplex due to its targeting ability towards cancer cells overexpressing CD44 receptors.
|
Download:
|
| Fig. 3. (a) Visualization of cellular uptake of naked Cy5-RNA (control), CP-PTX-RNA and CP-PTX-RNA-CS and CP-PTX-RNA-CS+antiCD44 by confocal laser scanning microscopy (CLSM) after 4 h of incubation. siRNA is labeled with Cy5 (red). Nucleus is stained with DAPI (blue). The bar represents 10 μm. (b, d) Distribution of fluorescence intensity and mean fluorescence intensity (MFI) of MDA-MB-231 cells after 4 h incubation with naked Cy5-RNA, CP-PTX-RNA, CP-PTX-RNA-CS or lipo2k-RNA by flow cytometry. (c, e) Distribution of fluorescence intensity and MFI of CP-PTX-RNA-CS cellular uptake competition inhibit by antiCD44 to MDA-MB-231 cells after 2 h incubation. | |
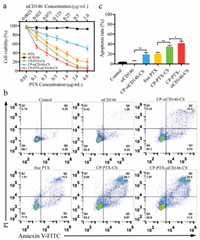
The combined therapeutic effects of PTX and siRNA towards cancer cells were assessed with typical MTT assay and flow cytometry analysis. As shown in Fig. 4a, the naked siRNA targeting CD146 (siCD146) displayed no obvious cytotoxicity against MDAMB-231 cells, while the nanocomplex CP-siCD146-CS (without PTX) exhibited a moderate cytotoxicity, causing a cell death rate of 51.2% at 1.0 μg/mL siCD146 concentration. The result suggests that CP-CS can deliver siCD146 into the breast cancer cells and inhibit the cell growth effectively. PTX is widely used to treat the breast cancer in clinical practice, and its underlying mechanism of action is to disrupt the microtubule normal function during cell proliferation [37]. By varying the PTX concentration (from 0.01 μg/mL to 4.0 μg/mL), CP-PTX-CS exhibited higher inhibition rate over free PTX after the same incubation period. The combined treatment with both PTX and siCD146 resulted in remarkable inhibition of cell growth. The IC50 value CP-PTX-siCD146-CS was 0.16 μg/mL, which was 3.6 fold lower than that of free PTX and 2 fold lower than that of CP-PTX-CS. All these results indicated that co-delivery of PTX and siCD146 mediated by CP-CS have the synergistic effects for growth inhibition of breast cancer cells. Moreover, the flow cytometry analysis indicated that the CPsiCD146-CS or siCD146 alone only led to weak apoptosis (early and late apoptosis, 6.7%–19.3%) (Figs. 4b and c). The apoptosis rate of free PTX was 21.2%, whereas CP-CS-mediated combined treatment with PTX and siCD146 could induce remarkable apoptosis, achieving an apoptosis rate of 43.0% over MDA-MB-231 cells. The above results are in agreement with the MTT results, where combined treatment exhibited higher cytotoxicity over siCD146 or PTX-mediated monotratment in MDA-MB-231 cells.
|
Download:
|
| Fig. 4. (a) Cytotoxicity of free PTX, siCD146, CP-PTX-CS, CP-siCD146-CS and CP-PTXsiCD146-CS against MDA-MB-231 cells by MTT assay for 48 h. Data represent mean ± SD (n = 6). (b) Apoptosis analysis of MDA-MB-231 cells after the treatment with the different formulations for 24 h. The quantitative analysis was evaluated by means of flow cytometry. The PTX concentration is 0.5 μg/mL. (c) Relative apoptosis rate of MDA-MB-231 cells after treatment with different formulations for 24 h. Data represent mean ± SD (n = 3, Student's t-test, **P < 0.01, *P < 0.05). | |
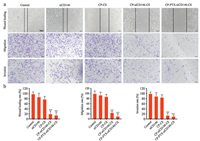
To evaluate the anti-metastatic property of CP-PTX-siCD146-CS, we first investigated the expression of CD146 in different breast cancer cells using the flow cytometry. In Fig. S7 (Supporting information), the CD146 expression of MDA-MB-231 cells was higher than that of MCF-7 cells. Therefore, the anti-metastasis effect of CP-PTX-siCD146-CS was performed in MDA-MB-231 cells by using the wound healing assays, which assessed the lateral migration ability of cancer cells. As shown in Fig. 5a, MDA-MB-231 cells without any treatment (as control group) displayed high wound healing rate, and hardly detect the initial scratching, indicating high metastatic ability of MDA-MB-231 cells. The wound healing rate of the cells pre-treated with CP-CS was less than that of control group, indicating the potential anti-metastatic property of CS. Strikingly, CP-siCD146-CS and CP-PTX-siCD146-CS complexes exhibited significant inhibitory effect of cell motility, and the wound healing rate was less than 20%. To further assess the anti-metastasis effect of CP-PTX-siCD146-CS, the migration and invasion assays were also performed to evaluate the longitudinal migration ability of the MDA-MB-231 cells (Fig. 5a). In migration and invasion assays, the number of migrated cells were counted in the lower surface of the transwell chamber. The CP-siCD146-CS and CP-PTX-siCD146-CS nanocomplexes displayed strong inhibitory effect on MDA-MB-231 cell migration and invasion, with the migration rate reduced by 74.7% and 88.7%, respectively. In the invasion assay, the treatment of CP-siCD146-CS and CP-PTXsiCD146-CS also could reduce 84.8% and 88.5% MDA-MB-231 cells invasion (Fig. 5b). The result was in agreement with the wound healing results. All above evidence indicate the CP-PTX-siCD146-CS exhibit good anti-metastasis effect for breast cancer cell.
|
Download:
|
| Fig. 5. The wound healing, migration assay and invasion assay of MDA-MB-231 cells after co-incubated with PBS, siCD146, CP-CS, CP-siCD146-CS and CP-PTX-siCD146-CS, respectively (n = 3, ***P < 0.001), scale bar 100 μm. | |
As a potent gene-silencing agent that can inhibit the CD146 over-expression in the breast cancer cells, the released siCD146 is the most important source of anti-metastasis effect. To determine the inhibition of CD146 over-expression in MDA-MB-231 cells, the immunophenotyping assay was investigated using the flow cytometry analysis. As show in Fig. 6a, the nanocomplex loading siCD146 (CP-siCD146-CS and CP-PTX-siCD146) effectively suppressed the CD146 expression (less than 3% of CD146 expression), whereas the group without siCD146 exhibited high level of CD146 expression. The similar results were also observed by performing western blotting analysis (Fig. 6b), where the MDA-MB-231 cells treated with CP-PTX-siCD146-CS or CP-siCD146-CS exhibited significant knockdown of CD146 protein. These results suggest that the siCD146 is a key element to inhibit the metastasis effect in breast cancer cells, and the efficient delivery mediated by CP-CS is critical to fulfill the inhibition function of siCD146.

|
Download:
|
| Fig. 6. (a) Expression of CD146 in MDA-MB-231 cells after the treatment of control (a1), CP-CS (a2), CP-siCD146-CS (a3) and CP-PTX-siCD146-CS (a4) determined by immunophenotyping. (b) CD146 protein expression after the treatment of control, CP-CS, CP-siCD146-CS and CP-PTX-siCD146-CS in MDA-MB-231 cells by western blot. | |
In summary, we have developed a delivery carrier that could codeliver siRNA and PTX for the combined treatment of breast cancer. The delivery carrier, CP-CS, could simultaneously load PTX through host-guest interaction and siRNA through electrostatic interaction to form CP-PTX-siCD146-CS nanocomplexes. Owing to the exclusive interaction between CS and CD44, the nanocomplexes exhibited active targeting towards CD44-overexpresing breast cancer cells. Whereas the delivered PTX served as an active drug to inhibit cancer cells proliferation, the co-delivered siCD146 played an important role for blocking metastasis and inducing apoptosis of cancer cells. In addition, we found CP-PTX-siCD146-CS nanocomplexes exhibited higher cytotoxicity and anti-metastasis effects over PTX or siCD146 mono-treatment. This study provides a useful strategy for the simultaneous delivery of chemotherapeutic drug and RNAi therapeutics for future combined treatment of breast cancer and metastasis prevention.
AcknowledgmentThis work was supported Fundamental Research Funds for the Central Universities (No. 520002*172210381) for the financial support of this work.
Appendix A. Supplementary dataSupplementary material related to this article can be found, in the online version, at doi:https://doi.org/10.1016/j.cclet.2019.06.022.
| [1] |
Cancer Genome Atlas Network, Nature 490 (2012) 61-70. DOI:10.1038/nature11412 |
| [2] |
D. Hanahan, R.A. Weinberg, Cell 144 (2011) 574-646. |
| [3] |
W.D. Foulkes, I.E. Smith, J.S. Reis-Filho, N. Engl. J. Med. 363 (2010) 1938-1948. DOI:10.1056/NEJMra1001389 |
| [4] |
E.P. Mamounas, J. Bryant, B. Lembersky, et al., J. Clin. Oncol. 23 (2005) 3686-3696. DOI:10.1200/JCO.2005.10.517 |
| [5] |
R.A. Weinberg, C.L. Chaffer, Science 331 (2011) 1559-1564. DOI:10.1126/science.1203543 |
| [6] |
L. Carey, G. Winer, G. Viale, et al., Nat. Rev. Clin. Oncol. 7 (2010) 683-692. DOI:10.1038/nrclinonc.2010.154 |
| [7] |
L.R. Yates, S. Knappskog, D. Wedge, Cancer Cell 32 (2017) 169-184. DOI:10.1016/j.ccell.2017.07.005 |
| [8] |
M. Wu, X. Liu, W. Jin, et al., J. Control. Release 253 (2017) 110-121. DOI:10.1016/j.jconrel.2017.03.011 |
| [9] |
J. Wang, W. Xu, S. Li, et al., J. Biomed. Nanotechnol. 14 (2018) 2102-2113. DOI:10.1166/jbn.2018.2624 |
| [10] |
H. Xiao, L. Yan, E.M. Dempsey, et al., Prog. Polym. Sci. 87 (2018) 70-106. DOI:10.1016/j.progpolymsci.2018.07.004 |
| [11] |
Y. Zhang, R. Sha, L. Zhang, et al., Nat. Commun. 9 (2018) 4236. DOI:10.1038/s41467-018-06529-y |
| [12] |
X. Feng, J. Ding, R. Gref, X. Chen, Chin. J. Polym. Sci. 35 (2017) 693-699. DOI:10.1007/s10118-017-1932-7 |
| [13] |
S. Tang, Q. Meng, H. Sun, Adv. Funct. Mater. 26 (2016) 6033-6046. DOI:10.1002/adfm.201601703 |
| [14] |
Y. Ping, C. Liu, Z. Zhang, et al., Biomaterials 32 (2011) 8328-8341. DOI:10.1016/j.biomaterials.2011.07.038 |
| [15] |
Y. Li, H. Bai, H. Wang, et al., Nanoscale 10 (2018) 203-214. DOI:10.1039/C7NR06689A |
| [16] |
C.J. Cheng, R. Bahal, I.A. Babar, et al., Nature 518 (2015) 107-110. DOI:10.1038/nature13905 |
| [17] |
J. Zhou, Y. Wu, C. Wang, et al., Angew. Chem. Int. Ed. 55 (2016) 1-6. DOI:10.1002/anie.201510990 |
| [18] |
J. Nie, B. Qiao, S. Duan, et al., Adv. Mater. 1416 (2018) 1801570. |
| [19] |
N. Afratis, C. Gialeli, D. Nikitovic, et al., FEBS J. 279 (2012) 1177-1197. DOI:10.1111/j.1742-4658.2012.08529.x |
| [20] |
R.D. Prinz, C.M. Willis, A. Viloria-Petit, M. Kluppel, Genet. Mol. Res. 14 (2011) 3901-3913. |
| [21] |
D.J. Silver, F.A. Siebzehnrubl, M.J. Schildts, et al., J. Neurosci. 33 (2013) 15603-15617. DOI:10.1523/JNEUROSCI.3004-12.2013 |
| [22] |
C.A. Cooney, F. Jousheghany, A. Yao-Borengasser, et al., Breast Cancer Res. 13 (2011) R59. DOI:10.1186/bcr2896 |
| [23] |
B. Monzavi-karbassi, J.S. Stanley, L. Hennings, et al., Int. J. Cancer 120 (2016) 1179-1191. |
| [24] |
L. Zhao, M. Liu, J. Wang, et al., Carbohydr. Polym. 133 (2015) 391-399. DOI:10.1016/j.carbpol.2015.07.063 |
| [25] |
G. Kwak, D. Kim, G. Nam, et al., ACS Nano 11 (2017) 10135-10146. DOI:10.1021/acsnano.7b04717 |
| [26] |
H. Yan, O.P. Oommen, D. Yu, et al., Adv. Funct. Mater. 25 (2015) 3907-3915. DOI:10.1002/adfm.201500695 |
| [27] |
O.P. Varghese, J. Liu, K. Sundaram, J. Hilborn, O.P. Oommen, Biomater. Sci. 4 (2016) 1310-1313. DOI:10.1039/C6BM00335D |
| [28] |
K. Liang, K.H. Bae, F. Lee, et al., J. Control. Release 226 (2016) 205-216. DOI:10.1016/j.jconrel.2016.02.004 |
| [29] |
T. Jiang, R. Mo, A. Bellotti, J. Zhou, Z. Gu, Adv. Funct. Mater. 24 (2014) 2295-2304. DOI:10.1002/adfm.201303222 |
| [30] |
Q. Zeng, W. Li, D. Lu, et al., Cell Commun. Signal 15 (2017) 45. DOI:10.1186/s12964-017-0200-3 |
| [31] |
V. Todorovic, G. Sersa, M. Cemazar, Cancer Gene Ther. 20 (2013) 208-210. DOI:10.1038/cgt.2013.3 |
| [32] |
T. Wan, D. Niu, C. Wu, et al., Mater. Today 12 (2018) 003. |
| [33] |
Z. Zhang, T. Wan, Y. Chen, et al., Macromol. Rapid Commun. (2018) 1800068. |
| [34] |
D. Li, Y. Li, H. Xing, et al., Adv. Funct. Mater. 24 (2014) 5482-5492. DOI:10.1002/adfm.201400456 |
| [35] |
Y. Ping, Q. Hu, G. Tang, et al., Biomaterials 34 (2013) 6482-6494. DOI:10.1016/j.biomaterials.2013.03.071 |
| [36] |
X. Chen, Y. Qiu, C. Owh, X.J. Loh, Y. Wu, Nanoscale 8 (2016) 18876-18881. DOI:10.1039/C6NR08055C |
| [37] |
G. Yu, Z. Yang, X. Fu, et al., Nat. Commun. 9 (2018) 766. DOI:10.1038/s41467-018-03119-w |
 2020, Vol. 31
2020, Vol. 31 

