b Graduate School, University of Chinese Academy of Sciences, Beijing 100049, China
Gene therapy is a promising tool for the treatment of human disease that cannot be cured by rational therapies [1, 2]. Over the past few years, the clinical success of cancer gene therapy critically depends on the development of safe, efficient, and targeted gene carriers [3, 4]. Synthetic non-viral vectors have attracted considerable interest as a promising alternative to conventional viral vectors due to their lower risk of immunogenicity and larger gene delivery [5]. Among the vectors, dendrimers, a class of synthetic polymers with well-defined structure and molecular weight, have attracted considerable attention [6].
Dendrimers are highly branched macromolecules of low polydispersity that provide many opportunities for design of novel drug-carriers, gene delivery systems and imaging agents [7, 8]. They show great promise in tissue targeting applications, controlled drug release and so on [9, 10]. Dendrimers own advantages including their nanoscale spherical architecture, narrow polydispersity and the multifunctional surface offering the possibility to tailor-make their surface chemistry [11]. However, among most dendrimers that have been synthesized, despite few possible applications in medicine and pharmacy their utilization is limited, mainly because of inherent toxicity associated with them [12].
It has been reported that the main mechanism of dendrimers toxicity is based on the interaction of surface cationic charge of dendrimers with negatively charged biological membranes which results in membrane disruption and erosion [13]. Dendrimers bearing -NH2 termini display concentration and usually generation-dependent cytotoxicity [11]. Previous studies had pointed out that only dendrimers of high generations with positive charge, not negative or neutral or small cationic dendrimers, contributed to aggregation of human platelets in vitro [14]. In another study, Duncan et al. had reported that up to the concentration of 100 mg/mL tested, the cationic polyamidoamine (PAMAM) dendrimers of generation 4 was markedly cytotoxic while PAMAM of generation 1 was not toxic at all towards B16F10 cells [15]. In addition, a cytotoxicity analysis measured by MTT assay showed that cationic PAMAM of generation 2 had almost forty times higher IC50 value (the inhibitory concentration diminishing viability by 50%) than that of generation 4 [11]. Accordingly, a possible explanation for these results may be that the smaller size and less surface charge distribution of low generations dendrimers contribute their low cytotoxicity [16, 17].
In the past, a lot of work had focused on the effects of surface modification on cytotoxicity response of PAMAM, such as guanidine, carboxylate, sulfonate, phosphonate, or PEGylated surfaces [18]. To the best of our knowledge, there was little scientific information concerning the influences of structural changes on PAMAM cytotoxicity, especially caused by intracellular stimuli. In this study, we synthesized a reactive oxygen species (ROS)-responsive PAMAM dendrimer that was easily cleavable in ROS abundant conditions, which reduced the size and surface charge quantity of PAMAM. The ROS-responsive dendrimer demonstrated low toxicity than non-responsive PAMAM or polyetherimide. What is more, responding to ROS offers a parameter for manipulating nanocarrier to achieve better targeting and efficacy in complicated microenvironments [19]. The targeted delivery of siRNA to tumour cells was carried out to check the transport efficiency of nanocarrier. The results showed that the ROS-responsive dendrimer was secure and effective as a gene carrier.
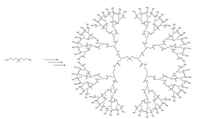
The cationic PAMAM dendrimers have primary amines groups on the surface and tertiary amine groups in their internal cavities, which become ionized at low pH values. They can effectively accept protons in acidic environments. As is known, the efficient intracellular gene delivery is mainly ascribed to strong electrostatic attraction between delivering carriers and siRNA [20, 21], therefore PAMAM is a suitable candidate for gene delivery owing to its proton buffering capacity. Moreover, various smart polymers capable of responding to intrinsic stimuli in a tumour environment, such as low extracellular pH and matrix metalloproteinases, have been developed as efficient carriers for cancer treatment [22, 23]. It was reported that, cancer cells can constantly generate high level of intracellular ROS, including H2O2, hydroxyl radial and superoxide, in comparison with normal cells [24-26]. Accordingly, we speculated that intracellular ROS in cancer cells might be utilized as an effective cancer-related stimulus to mediate intracellular gene release. It has been confirmed that the thioketal-based polymeric carrier could be cleaved by intracellular ROS and then enhance the gene delivery efficiency in cancer cells [5]. Therefore, a ROS-responsive dendrimer (G4.0), i.e., ROSPAMAM was produced from methyl acrylate and ethylenediamine using thioketal as the core (Scheme 1).
|
Download:
|
| Scheme 1. Synthetic route of ROS-PAMAM (G4.0). | |
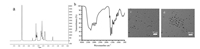
The ROS-PAMAM was characterized by 1H NMR (Fig. 1a). The appearance of alkyl protons in the 1H NMR spectrum (inset) indicates that the thioketal successfully served as the core. FTIR spectrum of ROS-PAMAM are shown in Fig. 1b. The characteristic absorption bands at 1731 cm-1 can be assigned to the ester carbonyl group of PAMAM. The molecular weights of ROS-PAMAM were calculated to be 14.35 kDa by the above molecular formula. Then, transmission electron microscopy (TEM), dynamic light scattering (DLS) and zeta potential measurements were performed to further characterize their size, morphology and surface charge properties of ROS-PAMAM and siRNA/ROS-PAMAM complex. Firstly, TEM images demonstrated that the size of siRNA/ROSPAMAM was 10.1 nm, whereas that of ROS-PAMAM was only 6.7 nm (Figs. 1c and d), indicating that siRNA effectively combined to ROS-PAMAM. Secondly, the zeta potential of ROS-PAMAM and siRNA/ROS-PAMAM were 19.3 mV and 4.07 mV (Fig. S5 in Supporting information), further implying that siRNA had linked to ROS-PAMAM. Then, DLS results showed that the hydrodynamic size of ROS-PAMAM was 29.2 nm, and that of siRNA/ROS-PAMAM increased to 45.7 nm (Fig. S5). As comparison, the hydrodynamic diameters of siRNA/PAMAM and siRNA/PEI were shown in Table S1 (Supporting information). Taken together, these results provide clear evidence for the formation of ROS-PAMAM and siRNA/ROSPAMAM.

|
Download:
|
| Fig. 1. The characterization of ROS-PAMAM (G4.0). (a) 1H NMR, inset: a partial enlargement picture of (a); (b) FTIR; (c) Transmission electron microscopy (TEM) images of ROS-PAMAM and (d) siRNA/ROS-PAMAM. | |
Generally, the cytotoxicity of nanoparticles should be firstly considered before they are used in vivo [27, 28]. To investigate the cytotoxicity of ROS-PAMAM, both MTT assay and LDH release assay were used. Two cell lines (A549 cells and MCF-7 cells) were exposure to different concentrations of ROS-PAMAM dendrimers for 24 h and the same concentrations of PAMAM and Polyethyleneimine (PEI) were included for comparison. PAMAM has a core of ethylenediamine instead of thioketal, as a result, it is not responsive to reactive oxygen species. PEI as a model gene transfection vector is often used in daily experiments, which however is considered too toxic [29]. Fig. 2 shows the viability of A549 cells and MCF-7 cells after 24h of incubation with ROSPAMAM, PAMAM and PEI. As expected, A549 cells and MCF-7 cells treated with ROS-PAMAM showed no cytotoxicity even at a concentration of 200 μg/mL in comparison with the IC50 value of PAMAM is about 100 μg/mL and that of PEI is about 50 μg/mL, confirming the low cytotoxicity of ROS-PAMAM. The results from both methods were consistent. Such results suggest that ROSPAMAM demonstrated low toxicity than non-responsive PAMAM or PEI, and it is more secure to be used as a gene carrier for siRNA delivery. To explain the low toxicity of ROS-PAMAM, the particle size (hydrodynamic diameter) and zeta potential changes of ROSPAMAM before and after treated with ROS were measured. As shown in Fig. S6 (Supporting information), both particle size and zeta potential were reduced after treated with ROS. Dendrimers bearing -NH2 termini display concentration and usually generation-dependent cytotoxicity, the smaller size and less surface chargedistribution of ROS-PAMAM in ROS-abundant conditions, as the characters of low generations dendrimers, may contribute its low cytotoxicity [11, 14-17].

|
Download:
|
| Fig. 2. The cytotoxicity analysis of ROS-PAMAM dendrimers. Cell viability was assessed by MTT assay and LDH release assay upon exposure to different concentrations of ROSPAMAM dendrimers for 24h. PAMAM and PEI were included for comparison. Results are presented as percent cell viability (mean ±S.D.) from three independent experiments using A549 cells and MCF-7 cells. MTT assay (a) A549 cells, (b) MCF-7 cells; LDH release assay (c) A549 cells, (d) MCF-7 cells. | |
To investigate the cell uptake behaviour of ROS-PAMAM, FITC labelled ROS-PAMAM was synthesized according to our previous studies [30]. Then, the cellular uptake of ROS-PAMAM was calculated by observing the location of FITC-ROS-PAMAM in subcellular compartments of A549 cells by confocal laser scanning microscopy (CLSM). As shown in Fig. S7 (Supporting information), the green fluorescence intensity increased by extending the incubation time from 1 h to 6 h, suggesting that the ROS-PAMAM could be endocytosed continuously in a time-dependent manner. Next, we co-stained FITC-ROS-PAMAM in A549 cells with LysoTracker Red DND-99 (lysosomal marker). After 6 h incubation, the green and red fluorescence signals from FITC-ROS-PAMAM and lysosome marker, respectively, were found to overlap (Fig. S7). This overlap indicated that the internalized ROS-PAMAM was translocated via lysosomal pathway after entering cells. Based on these results, it could be inferred that ROS-PAMAM can be efficiently translocated to endosomes in A549 cells.
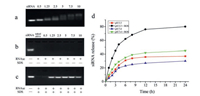
The interaction between negatively charged siRNA and cationic vehicles is known to strongly influence its loading efficiency of vehicles [31, 32]. To investigate whether ROS-PAMAM could complex with siRNA, agarose gel electrophoresis was performed at different mass ratios of vector to siRNA (w/w) ranging from 0.5:1 to 10:1 (Figs. 3a–c) [33]. As Fig. 3a shows, in comparison with the motility of naked siRNA, the movement of siRNA loaded on ROSPAMAM was gradually retarded. The fluorescence intensity of uncomplexed siRNA decreased with increasing w/w ratio from 0.5 to 10 and nearly disappeared when the w/w ratio was greater than 5, suggesting that ROS-PAMAM has strong condensation ability towards siRNA and higher w/w ratios are favourable. The stability of siRNA/ROS-PAMAM complexes were then examined by RNase digestion to mimic physiological condition. Un-complexed siRNA was detected with ethidium bromide stain. Once siRNA was either degraded by RNase or completely bound with gene vehicles, the ethidium bromide fluorescence disappeared. Fig. 3b shows that the fluorescence of siRNA/ROS-PAMAM complexes disappeared after RNase digestion, which means that within the complexes there is no naked gene present. Then, as Fig. 3c shows, after SDS treatment, siRNA was released from vehicle by SDS replacement. The fluorescence intensity gradually increased with the mass ratio from 0.5 to 7.5, indicating increasing number of siRNA was released from ROS-PAMAM. These results further suggested that complexed siRNA was protected by ROS-PAMAM instead of being degraded by RNase. However, we also found that the fluorescence intensity decreased at the mass radio of 10, it was possibly because that the vehicle was over dosage, as a result, a little siRNA on the surface of ROS-PAMAM was degraded. Thus, gene delivery experiments were carried out at an optimized w/w ratio of 7.5.

|
Download:
|
| Fig. 3. (a–c) Analysis of siRNA/ROS-PAMAM complex formation at various mass ratios by agarose gel electrophoresis using 2% agarose in Tris-acetate running buffer. The siRNA/ROS-PAMAM complexes were prepared at pH 7.4. (a) Agarose gel retardation assay of siRNA/ROS-PAMAM complexes under various vector/siRNA mass ratios (0.5:1, 1.25:1, 2.5:1, 5:1, 7.5:1, and 10:1 from left to right). (b) Protection of siRNA against RNase digestion. (c) siRNA release from complexes by competitive binding of SDS with vehicles after enzymatic degradation. Naked siRNA was used as a control (first lane from the left). (d) siRNA release profile of siRNA/ROS-PAMAM complex under different conditions. | |
Then, we investigated the release behaviour of siRNA from siRNA/ROS-PAMAM complexes [34]. We used 100 mmol/L H2O2 and 1.6 mmol/L CuCl2 in PBS buffer to produce ROS according to a previous report [35]. Fig. 3d shows the amount of siRNA cumulatively released under different conditions. In the pH 7.4 and absence of ROS, we observed that there is no apparent siRNA (about 30%) release after 24 h. Then, under a single stimulus of ROS or partial acidic environment (pH 5.5), the amount of siRNA released was slightly improved to 40% but the effect was not obvious. In contrast, in the presence of ROS under slightly acidic environment, the siRNA release rate significantly increased, and nearly 80% siRNA was released within 12 h. For intracellular release of siRNA from ROS-PAMAM, it is known that cationic particles bind with high affinity to lipid groups on the surface membrane and are endocytosed in the tight-fitting vesicles. Once ROS-PAMAM nanoparticles enter into an acidifying lysosomal compartment (pH 5.5) [36], the unsaturated amino groups of ROS-PAMAM are capable of sequestering protons that are supplied by the v-ATPase (proton pump). This process keeps the pump functioning and leads to the retention of one Cl- ion and one water molecule per proton, which causes lysosomal swelling and rupture, leading to particle deposition in the cytoplasm [37]. The protonation of the central tertiary amines can increase the volume of internal cavity of ROSPAMAM, which promotes siRNA release from siRNA/ROS-PAMAM complexes [38]. Then, cancer cells can constantly generate high level of intracellular ROS, including H2O2, hydroxyl radial and superoxide, in comparison with normal cells [39, 40]. As a result, the internalized ROS-PAMAM disassembles, which further promotes the release of siRNA into the cytoplasm.
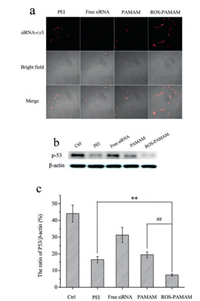
To investigate the siRNA delivery efficacy by ROS-PAMAM, confocal laser scanning microscopy (CLSM) were used to measure the uptake of cy5-labeled siRNA. We used PAMAM as a reference to investigate whether ROS-PAMAM responded to ROS and facilitated the release of siRNA in vivo. As shown in Fig. 4a, siRNA/ROSPAMAM complexes exhibited strong fluorescence intensity in A549 cells, while the cy5-labeled siRNA delivered by PEI or PAMAM showed relatively weak fluorescence. Moreover, the fluorescence intensities of cy5-labelled siRNA uptake by A549 cells were determined by flow cytometry analysis (Fig. S8 in Supporting information), The results were in agreement with the above data determined by confocal laser scanning microscopy. That is, after treatment with siRNA/ROS-PAMAM, the fluorescent intensity of A549 cells increased much more than the other two carriers, suggesting that ROS-PAMAM can increase the transfection efficiency.
|
Download:
|
| Fig. 4. (a) Confocal microscope images of intracellular uptake of cy5-labeled siRNA (red) complexed with PEI, PAMAM, ROS-PAMAM after incubation with A549 cells for 12 h. Free siRNA was used as a control. (b) The A549 cells were incubated with free p53 siRNA, PEI, PAMAM and ROS-PAMAM complexed with p53 siRNA for 12 h, respectively. The p53 knockdown efficacy was detected by western blot. (c) Quantitative analysis of p53 knockdown efficacy. | |
Moreover, to further evaluate the effectiveness of siRNA transfection by ROS-PAMAM in vitro, we used p53 siRNA that specifically knockdown p53 protein [41]. p53 siRNA was delivered via ROS-PAMAM, PAMAM, PEI, and the level of p53 was measured by Western Blot (Fig. 4b). In A549 cells treated with p53 siRNA/ ROS-PAMAM at a mass ratio of 7.5, significant p53 knockdown was observed and the level of p53 was reduced by nearly 83% (Fig. 4c), compared to a slight reduction in the cells treated with siRNA/PEI (62%) and siRNA/PAMAM (51%). From those results, it can be concluded that ROS-PAMAM is an effective non-viral gene carrier.
With the advent of nanotechnology, concerns about the potential adverse health effects of nanomaterials have been expressed. It is of great importance to find nanomaterials with lower toxicity, or to make appropriate improvements to reduce nanotoxicity. Here, we prepared a ROS-responsive dendrimer, which exhibits low cytotoxicity than non-responsive PAMAM or polyethyleneimine. The low toxicity mechanism of ROS-PAMAM may be attributed to its easily cleavable capability in ROS abundant conditions, which reduced the size and surface charge quantity of PAMAM. Dendrimers bearing -NH2 termini display concentration and usually generation-dependent cytotoxicity, the smaller size and less surface charge distribution of ROS-PAMAM in ROSabundant conditions, as the characters of low generations dendrimers, may contribute its low toxicity.
High siRNA transfection efficiency is the most important factor in gene modification of cells. The targeted delivery of siRNA to tumour cells was carried out to check the transport efficiency of ROS-PAMAM. Results showed that this vector demonstrated high siRNA binding affinity and protected siRNA from enzyme degradation. Moreover, confocal laser scanning microscopy demonstrate the high gene delivery efficiency of ROS-PAMAM at a vector/siRNA w/w ratio of 7.5:1. Furthermore, the targeted knockdown effect on p53 expression was much higher than that of PEI and non-responsive PAMAM using our new delivery carrier. This indicates that ROS-PAMAM has the potential to be used as an efficient in vivo gene carrier. We believe that ROS-PAMAM is a promising delivery vehicle and can contribute to the design of novel carriers with low cytotoxicity and high delivery efficiency.
AcknowledgmentThis project was supported by the National Natural Science Foundation of China (Nos. 31571020, 31570856 and 31300697).
Appendix A. Supplementary dataSupplementary material related to this article can be found, in the online version, at doi:https://doi.org/10.1016/j.cclet.2019.03.040.
| [1] |
Y. Deng, C.C. Wang, K.W. Choy, et al., Gene 538 (2014) 217-227. DOI:10.1016/j.gene.2013.12.019 |
| [2] |
X. Li, L. Zhao, Q. Liang, et al., J. Biomed. Nanotechnol. 13 (2017) 280-289. DOI:10.1166/jbn.2017.2350 |
| [3] |
W.W. Wang, W.Z. Li, N. Ma, G. Steinhoff, Curr. Pharm. Biotechnol. 14 (2013) 46-60. |
| [4] |
Q. Zheng, D. Lin, L. Lei, et al., J. Biomed. Nanotechnol. 13 (2017) 1565-1580. DOI:10.1166/jbn.2017.2489 |
| [5] |
M.S. Shim, Y. Xia, Angew. Chem. Int. Ed. 52 (2013) 6926-6929. DOI:10.1002/anie.201209633 |
| [6] |
P. Kesharwani, A. Gothwal, A.K. Iyer, et al., Drug Discov. Today 23 (2018) 300-314. DOI:10.1016/j.drudis.2017.06.009 |
| [7] |
H.R. Ihre, O.L. Padilla De Jesús, F.C. Szoka, et al., Bioconjugate Chem. 13 (2002) 443-452. DOI:10.1021/bc010102u |
| [8] |
J. Wang, G.S. Williamson, M.G. Lancina III, H. Yang, J. Biomed. Nanotechnol. 13 (2017) 1089-1096. DOI:10.1166/jbn.2017.2436 |
| [9] |
Y. Fan, S. Yuan, M. Huo, et al., Nanomed. Nanotechnol. 13 (2017) 1399-1410. DOI:10.1016/j.nano.2017.01.008 |
| [10] |
V.M. Thanh, T.H. Nguyen, T.V. Tran, et al., Mater. Sci. Eng. C-Mater 82 (2018) 291-298. DOI:10.1016/j.msec.2017.07.051 |
| [11] |
N. Malik, R. Wiwattanapatapee, R. Klopsch, et al., J. Control. Release 65 (2000) 133-148. DOI:10.1016/S0168-3659(99)00246-1 |
| [12] |
S.P. Gautam, R.K. Keservani, T. Gautam, et al., Ars Pharm. 56 (2015) 155-159. DOI:10.4321/S2340-98942015000300004 |
| [13] |
M.A. Dobrovolskaia, A.K. Patri, J. Simak, et al., J. Mol. Pharm. Org. Process Res. 9 (2012) 382-393. |
| [14] |
C.C. Lee, J.A. MacKay, J.M. Fréchet, F.C. Szoka, Nat. Biotechnol. 23 (2005) 1517-1526. DOI:10.1038/nbt1171 |
| [15] |
R. Duncan, L. Izzo, Adv. Drug Deliver. Rev. 57 (2005) 2215-2237. DOI:10.1016/j.addr.2005.09.019 |
| [16] |
R.B. Kolhatkar, K.M. Kitchens, P.W. Swaan, et al., Bioconjugate Chem. 18 (2007) 2054-2060. DOI:10.1021/bc0603889 |
| [17] |
B.H. Zinselmeyer, S.P. Mackay, A.G. Schatzlein, I.F. Uchegbu, Pharmaceut. Res. 19 (2002) 960-967. DOI:10.1023/A:1016458104359 |
| [18] |
V. Mishra, U. Gupta, N. Jain, J. Biomater. Sci. Polym. E 20 (2009) 141-166. DOI:10.1163/156856208X386246 |
| [19] |
Y. Shen, X. Ma, B. Zhang, et al., Chem.-Eur. J. 17 (2011) 5319-5326. DOI:10.1002/chem.201003495 |
| [20] |
M.W. Amjad, P. Kesharwani, M.C.I.M. Amin, A.K. Iyer, Prog. Polym. Sci. 64 (2017) 154-181. DOI:10.1016/j.progpolymsci.2016.09.008 |
| [21] |
D.W. Pack, A.S. Hoffman, S. Pun, P.S. Stayton, Nat. Rev. Drug Discov. 4 (2005) 581-593. DOI:10.1038/nrd1775 |
| [22] |
H. Hatakeyama, H. Akita, H. Harashima, Adv. Drug Deliver. Rev. 63 (2011) 152-160. DOI:10.1016/j.addr.2010.09.001 |
| [23] |
K. Chen, H. Cai, H. Zhang, et al., Acta Biomater. 84 (2019) 339-355. DOI:10.1016/j.actbio.2018.11.050 |
| [24] |
S. Tripathy, P.K. Mohanty, Int. J. Pharm. Sci. Res. 8 (2017) 1-16. |
| [25] |
I.I.C. Chio, D.A. Tuveson, Trends Mol. Med. 23 (2017) 411-429. DOI:10.1016/j.molmed.2017.03.004 |
| [26] |
H.Y. Lee, E.I. Parkinson, C. Granchi, et al., ACS Chem. Biol. 12 (2017) 1416-1424. DOI:10.1021/acschembio.7b00015 |
| [27] |
T. Liao, Q. Deng, B. Wu, et al., Biomed. Mater. 12 (2017) 015018. DOI:10.1088/1748-605X/aa52ca |
| [28] |
K. Luo, B. He, Y. Wu, Y. Shen, Z. Gu, Biotechnol. Adv. 32 (2014) 818-830. DOI:10.1016/j.biotechadv.2013.12.008 |
| [29] |
G. Guo, M. Tortorella, B. Zhang, Y. Wang, J. Colloid Interface Sci. 504 (2017) 633-644. DOI:10.1016/j.jcis.2017.05.009 |
| [30] |
L.B. Du, X.X. Miao, Y.L. Gao, H.Y. Jia, K. Liu, Nanomed. Nanotechnol. 10 (2014) 1411-1420. DOI:10.1016/j.nano.2014.04.001 |
| [31] |
A. Sizovs, P.M. McLendon, S. Srinivasachari, T.M. Reineke, Top. Curr. Chem. 296 (2010) 131-190. |
| [32] |
L.M. Guan, S.P. Huang, Z. Chen, et al., J. Nanopart. Res. 17 (2015) 1-9. DOI:10.1007/s11051-014-2856-6 |
| [33] |
L. Han, J. Zhao, X. Zhang, et al., ACS Nano 6 (2012) 7340-7351. DOI:10.1021/nn3024688 |
| [34] |
V. Dehousse, N. Garbacki, S. Jaspart, et al., Int. J. Biol. Macromol. 46 (2010) 342-349. DOI:10.1016/j.ijbiomac.2010.01.010 |
| [35] |
E. Fadillioglu, Z. Kurcer, H. Parlakpinar, et al., Arch. Pharm. Res. 31 (2008) 705-712. DOI:10.1007/s12272-001-1216-3 |
| [36] |
W.L. Chen, F. Li, Y. Tang, et al., Int. J. Nanomed. Nanosurg. 12 (2017) 4241-4256. DOI:10.2147/IJN.S129748 |
| [37] |
A.E. Nel, L. Mädler, D. Velegol, et al., Nat. Mater. 8 (2009) 543-557. DOI:10.1038/nmat2442 |
| [38] |
F. Joris, L. De Backer, T. Van de Vyver, et al., J. Control. Release 269 (2018) 266-276. DOI:10.1016/j.jconrel.2017.11.019 |
| [39] |
D. Trachootham, J. Alexandre, P. Huang, Nat. Rev. Drug Discov. 8 (2009) 579-591. DOI:10.1038/nrd2803 |
| [40] |
Z. Zhou, J. Song, L. Nie, X. Chen, Chem. Soc. Rev. 45 (2016) 6597-6626. DOI:10.1039/C6CS00271D |
| [41] |
Z.C. Edwards, E.W. Trotter, P. Torres-Ayuso, et al., Cancer Res. 77 (2017) 4961-4972. DOI:10.1158/0008-5472.CAN-17-0267 |
 2020, Vol. 31
2020, Vol. 31 

