Cells are naturally exposed to complex extracellular matrix (ECM) in vivo, in which the native proteins, rigidity and topographic structure of substrate corporately impact on cell behaviors [1-3]. Inversely, cell response to these external stimuli is the basis of cell functions. It is well known that ECM proteins usually have micrometer or nanometer scales topographic structures and cells biologically respond to structures when interact with ECM proteins commonly known as "contact guidance" [4]. As a consequence, cells morphology, adhesion and cytoskeleton arrangement are distinctly changed under contact guidance [5-7]. Thus, topographic cues innately existing in extracellular microenvironment are essential in construction of biological models in vitro for accurately revealing physiological or pathological processes.
Cell migration underlies many biological processes including embryogenesis, wound healing, immune response, vascular disease and tumor metastasis, etc. [8, 9]. The highly integrated and extremely complicated cell migration process is regulated by numerous extracellular biophysical and biochemical factors [9-11]. However, the traditional research method for adherent cell migration in vitro is commonly scratch assay [12] where cells are cultured in planar substrate lacking of topographic cues. As the advanced development of micro- and nanofabrication techniques, a series of anisotropic and isotropic topographic structures, such as micro/nano grooves, fibers, pillars, and other particular shaped structures has been successfully fabricated [13-17]. It helps us to widely explore the topography roles in cell migration.
Vaccinia virus (VACV), usually as a model virus of poxviridae family, can induce cells intrinsic migration due to virus F11-mediated inhibition of RhoA signaling [20, 21]. The basic steps of cell migration induced by VACV are thought to be consisted of cell polarization, protrusions formation at leading edge, body contraction and adhesion release at the rear [18-20]. During migration process, actin cytoskeleton and microtubule of infected cells were distinctly reconfigured [18-20]. Recent years, VACV-induced cell migration has been proposed as a supplementary mechanism for VACV rapid spread in vivo [21-24]. Researchers widely suppose that these moving infectious bodies inevitably increase the risk of the neighboring or distant normal cells to be infected [20, 22, 23]. Changes in VACV-infected cells migration behaviors are likely to influence virus spread. However, the limited researches involved in VACV-induced cell migration are almost conducted on two-dimensional flat surface ignoring the effect of topographic cue on migration behaviors. Therefore, it is significant to study VACV-induced cell migration in the presence of topography for comprehensively understanding the infected cells behaviors in vivo, as well as for revealing the relationship of virus-induced cell migration and virus spread.
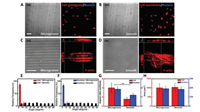
Herein, we developed an open microfluidic chip with microgrooved poly(dimethylsiloxane) (PDMS) substrate by soft lithography technique (Fig. S1 in Supporting information). Results showed that the uniform and parallel microgrooves were successfully fabricated and replicated on PDMS membrane (Fig. S2 in Supporting information). Due to the proportion of cells aligned along the micro-grating axis was obviously increased with decreased width of microgroove (Fig. S3 in Supporting information), we explored the effect of topography on cells morphology on 3 μm-wide microgrooved substrate. Differed from the random extension on two-dimension smooth substrate, Vero cells exhibited highly alignment along the micro-grating axis with elongated cells body and nucleus on microgrooved substrate (Figs. 1A, B and E-G). Meanwhile, cellular actin cytoskeleton was reconfigured and accumulated in the direction parallel to the microgroove compared to the random cross in the whole cytoplasm on smooth substrate (Figs. 1C and D, Fig. S4 in Supporting information). Nevertheless, the projected spreading areas of cell body and nucleus were no detected significant difference on microgrooved and smooth substrates (Fig. 1H). It was indicated that designed microgrooved PDMS substrate provided an ordered and directional physical surface as an effective topographic cue to regulate cells morphology viacontact guidance. Cells morphology on designed micro-scale topographic surface has many characteristics of that in an oriented fibrillar ECM to mimic the real extracellular topography in a certain extent [2]. It provides an opportunity for revealing the behaviors of VACV-infected cells in vivo.
|
Download:
|
| Fig. 1. Characterization of cell morphology and actin cytoskeleton organization on microgrooved and smooth substrates. Vero cells body and nucleus aligned uniformly in the axis parallel to the microgrooves (A) while located randomly on smooth substrate (B), Scale bars: 100 μm. (C) and (D) were z-stacked confocal images to describe the F-actin arrangement of two representative cells on microgrooved and smooth substrates, respectively. Scale bars: 20 μm. (E-H) were the results of statistical analysis for alignment angle, aspect ratio, and projected spreading area both in cells body and nucleus, respectively (n = 100). | |
Using this chip, we explored the topography roles in cell migration induced by VACV. The uninfected cell behaviors were firstly monitored on microgrooved substrate for 14 h and results revealed that microgrooved structure could not trigger uninfected Vero cells to migrate without additional stimuli (Fig. S5A and Movie S3 in Supporting information). A recombinant VACV expressing green fluorescent proteins (GFPs) was used in our experiments. The behaviors of VACV-infected cells at MOI 5 were observed from 2 hpi to 16 hpi for 14 h. Images displayed that infected cells migrated randomly on smooth PDMS substrate (Fig. S5B and Movie S2 in Supporting information). In contrast, on microgrooved substrate, vast majority of infected cells preferred to migrate along the axis parallel to microgrooves (Fig. S5C and Movie S1 in Supporting information). The consistent results were also observed in trajectory of representative single migratory cell (Figs. 2A and B). Moreover, the proportion of migratory cells along positive or negative micro-grating axis was equally and very few cells were reversed during migration (Movie S1). Subsequently, migration behaviors of infected cell were quantitatively analyzed. Results indicated that on microgrooved substrate, the migration path length and displacement of infected cells were both raised from 306.4 ± 72.4 μm and 194.5 ± 67.5 μm to 426.1 ± 46.7 μm and 355.4 ± 67.5 μm, respectively (Figs. 2C and D). The migration velocity was obviously increased between 6 hpi and 14 hpi on microgrooved substrate (Fig. 2E). The directionality reflected the persistence of cell migration in one direction. Infected migratory cells exhibited increased directionality of 0.83 ± 0.11 under topographic cue compared to that of 0.65 ± 0.13 on smooth substrate (Fig. 2F). Results mentioned above illustrated that topographic cue regulated the VACV-infected cell behaviors including the velocity and direction of cell migration.

|
Download:
|
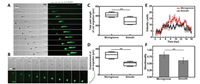
| Fig. 2. VACV-induced cell migration on different topographic substrates. (A) and (B) were confocal sequential images to describe the migration paths of two representative infected cells on microgrooved and smooth substrate. Green arrows diagrams to the top or right of the micrographs in (A) and (B) represented the direction and distance of cell movement each hour. Scale bars: 50 μm. (C-F) Statistical analysis of the motility of VACV-infected cells on microgrooved and smooth substrates including the total path length, displacement, velocity and directionality of cell migration, respectively (n = 90). | |
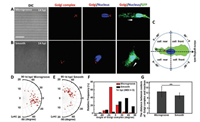
Generally, under two-dimensional culture, Golgi complex in migratory cell would be reoriented in front of the nucleus towards cellular leading edges, which is closely related to the direction of cell migration [9, 10]. To further explore the topography influence on subcellular structures of VACV-infected migratory cells, we observed and statistically analyzed the locations of Golgi complex in migratory cells. Results showed that on microgrooved substrate, Golgi complex was positioned behind the nucleus with respect to the direction of cell migration in more than 73% infected migratory cells (Figs. 3A and D-F). While a front position appeared in more than 86% infected migratory cells on smooth substrate (Figs. 3B and D-F). The quantification method of orientation angle of Golgi complex was shown in Fig. 3C. Furthermore, the orientation angle of Golgi complex with a concentrated distribution within -30° to 0° on microgrooved substrate indicated that the Golgi complex locations were confined within the micro-grating axis in infected migratory cells (Figs. 3D and F). In contrast, on smooth substrate orientation angle of Golgi complex in infected migratory cells was uniformly distributed from 0° to 90° (Figs. 3E and F). In addition, whether on microgrooved or smooth substrate, Golgi complex of infected migratory cells exhibited usual juxtanuclear localization without obvious displacement from the nucleus (Figs. 3A and B). The increased distance between the center points of Golgi complex and nucleus indirectly validated that the nucleus was elongated on microgrooved substrate (Fig. 3G). Nevertheless, microgrooved structure would not induce the relocations of Golgi complex in uninfected cells (Fig. S6 in Supporting information). Results mentioned above suggested that topography as an external cue regulated the reorientation process of Golgi complex in VACV-induced cell migration. In fact, the positions of Golgi complex and centrosome in migratory cells are dependent on the geometrical constraints of substrate [27]. The rearward position of Golgi complex would not affect the occurrence of cell migration. It indicates that a front position of Golgi complex in common mode observed on planar substrate in vitro is not only due to the polarization and migration of cells but rather by the fact that topographic cues are removed. This finding demonstrates that topographic cue is a vital external factor determined the positions the Golgi complex within a cell during migrating.
|
Download:
|
| Fig. 3. Topography influence on the orientation of Golgi complex in VACV-infected migratory cells. (A) and (B) were the fluorescence images to recognize the Golgi complex (red color) and nucleus (blue color), and GFPs expressed in cytoplasm (green color) of VACV-infected migratory cells on microgrooved substrate and smooth substrate. The white arrowheads represented the direction of cell migration. (C) The schematic diagram outlined the quantification method of Golgi complex orientation angle. The positive direction of micro-grating axis was related to the direction of cell migration. (D) and (E) Polar diagrams (R: the distance between the center points of Golgi complex and nucleus, θ: orientation angle) reflected the locations distribution of Golgi complex in infected migratory cells at 14 hpi. (F) and (G) Statistical analysis of orientation angle and the distance between the center points of Golgi complex and nucleus in infected migratory cells at 14 hpi (n = 57). Scale bar: 20 μm. | |
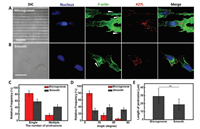
Controlling the formation of lamellipodial protrusions can modulate the direction of cell migration [10, 11]. To investigate the effect of topography on the protrusion formation in migratory cells, actin filaments were stained with phalloidin to recognize the morphologically distinct protrusion. Results illustrated more than 75% infected migratory cells preferred to extend single elongated protrusion confined within micro-grating axis on microgrooved substrate, while nearly 50% infected cells appearing multiple protrusions were randomly stretched on smooth substrate (Figs. 4A-D). In addition, the average length of protrusion was raised from 18.75 ± 7.14 μm to 29.16 ± 9.66 μm when infected migratory cells were exposed to topographic cue (Fig. 4E). For the individual migratory cell on microgrooved substrate, the length of protrusion was even up to nearly 50 μm (Fig. S7 in Supporting information).
|
Download:
|
| Fig. 4. Topography influence on the protrusion formation and the distribution of virions in migratory cells. To distinguish the red color of Dylight 649 from iFluorTM 555, iFluorTM 555 was painted in pseudo green color to recognize F-actin. (A) and (B) were the fluorescence images to recognize nucleus (blue color), virions (red color) and F-actin (green color) of migratory cells at 14 hpi (MOI = 5) on microgrooved substrate and smooth substrate, respectively. White arrowheads represented the extending direction of protrusions. The number, alignment angle and the length of protrusion in infected migratory cells on microgrooved and smooth substrates were statistical analyzed in (C-E), respectively (n = 85). Scale bars: 20 μm. | |
It has been reported that some types of virus like herpesvirus and poxvirus can induce actin/microtubule-filled projections to provide "superhighways" for virus reaching out and contacting with neighboring cells to enhance the spread of infection [22, 23, 25]. In our system, migratory cells infected with VACV carried lots of virions due to the replication of progeny virions in cytoplasm after infection [21]. These virions would be concentrated distribution in the protrusion of migratory cells influenced by cell polarization (Figs. 4A and B). The directional elongated actin-filled protrusion containing numerous virions is likely to facilitate virus delivery between migratory cells and distal uninfected cells. Moreover, compared to the cell-free spread mode of virus, cell-to-cell spread is thought to be a highly efficient dissemination route for enveloped virus, which can evade neutralizing antibodies, reduce surfing distance and increase local density of viral particles, etc. [26]. However, the cell-to-cell spread of virus is limited to that between the adjacent cells. In our study, VACV-infected cells exhibited accelerated directional cell migration on microgrooved substrate (Fig. 2). Changes in migration behaviors and marked structures of migratory cells in presence of topographic cue reflect that a smart and selective response of infected cells to substrate surface, which provides a new sight to understand the VACV rapid and direct spread behaviors in vivo.
In summary, our research reported that the accelerated and directionally persistent cell migration induced by VACV on microgrooved substrate. Vero cells on microgrooved substrate were highly aligned and elongated accompanied by the directed arrangement of actin cytoskeleton, exhibiting the morphological characteristics of cells exposed to topographic cues in vivo. On microgrooved substrate, the migration behaviors of VACV-infected cells were distinct changed compared to the observation of random migration in traditional scratch assay. Topographic structures of substrate guided the actin cytoskeleton reorganization of infected migratory cells via contact guidance leading to a dominant elongated protrusion along the axis of microgroove, which was mainly responsible for the increased directionality of cell migration. Moreover, topographic cue affected the relocation process of Golgi complex in migratory cells to cause the rearward position confined within the micro-grating axis, conflicting with the common mode of that in front position under traditional two-dimension cell culture. It appears that the locations of Golgi complex in migratory cells are significant influenced by the extracellular environment. Here, VACV-infected cells smartly respond to the substrate topography performing the extended actin-containing protrusion filled with numerous virions, as well as accelerated movement. These changes in morphology and motility of VACV-infected cells on microgrooved substrate are seemed to be beneficial for the rapid and direct spread of VACV. As we all know, cells exposed to ECM in vivo with higher geometric constraints. Our work demonstrates that topography is a vital external cue to regulate the migration behaviors induced by VACV. This finding opens a window for us to understand the VACV-induced cell migration in vivo, and lays a foundation of construction the physiologically-representative model for virology investigation in vitro. Elucidating the interactions between the virus, host cells and substrate topography can help us to reveal the relationship of virus-induced cell migration and virus rapid spread.
AcknowledgmentsThis work was supported by the National Natural Science Foundation of China (Nos. 21775111, 21475099) and the National Science and Technology Major Project of China (No. 2018ZX10301405).
Appendix A. Supplementary dataSupplementary material related to this article canbefound, in the online version, at doi:https://doi.org/10.1016/j.cclet.2019.05.043.
| [1] |
S.V. Plotnikov, A.M. Pasapera, B. Sabass, C.M. Waterman, Cell 151 (2012) 1513-1527. DOI:10.1016/j.cell.2012.11.034 |
| [2] |
A.D. Doyle, R.J. Petrie, M.L. Kutys, K.M. Yamada, Curr. Opin. Cell Biol. 25 (2013) 642-649. DOI:10.1016/j.ceb.2013.06.004 |
| [3] |
C.J. Bettinger, R. Langer, J.T. Borenstein, Angew. Chem. Int. Ed. 48 (2009) 5406-5415. DOI:10.1002/anie.200805179 |
| [4] |
D. Hoffman-Kim, J.A. Mitchel, R.V. Bellamkonda, Annu. Rev. Biomed. Eng. 12 (2010) 203-231. DOI:10.1146/annurev-bioeng-070909-105351 |
| [5] |
A. Ray, O. Lee, Z. Win, et al., Nat. Commun. 8 (2017) 14923. DOI:10.1038/ncomms14923 |
| [6] |
C.D. Paul, W.C. Hung, D. Wirtz, K. Konstantopoulos, Annu. Rev. Biomed. Eng. 18 (2016) 159-180. DOI:10.1146/annurev-bioeng-071114-040654 |
| [7] |
L. Liu, K. Yang, Z. Dai, et al., Chin. Chem. Lett. 30 (2019) 672-675. DOI:10.1016/j.cclet.2018.11.006 |
| [8] |
T. Yuan, D. Gao, S. Li, Y. Jiang, Chin. Chem. Lett. 30 (2019) 331-336. DOI:10.1016/j.cclet.2018.07.013 |
| [9] |
A.J. Ridley, M.A. Schwartz, K. Burridge, et al., Science 302 (2003) 1704-1709. DOI:10.1126/science.1092053 |
| [10] |
R.J. Petrie, A.D. Doyle, K.M. Yamada, Nat. Rev. Mol. Cell Biol. 10 (2009) 538-549. DOI:10.1038/nrm2729 |
| [11] |
C. Wang, N. Xu, Y.J. Yang, et al., Integr. Biol. 9 (2017) 903-911. DOI:10.1039/C7IB00151G |
| [12] |
C.C. Liang, A.Y. Park, J.L. Guan, Nat. Protoc. 2 (2007) 329-333. DOI:10.1038/nprot.2007.30 |
| [13] |
E.K. Yim, R.M. Reano, S.W. Pang, et al., Biomaterials 26 (2005) 5405-5413. DOI:10.1016/j.biomaterials.2005.01.058 |
| [14] |
A.D. Doyle, F.W. Wang, K. Matsumoto, K.M. Yamada, J. Cell Biol. 184 (2009) 481-490. DOI:10.1083/jcb.200810041 |
| [15] |
A. Ray, Z.M. Slama, R.K. Morford, S.A. Madden, P.P. Provenzano, Biophys. J. 112 (2017) 1023-1036. DOI:10.1016/j.bpj.2017.01.007 |
| [16] |
E.I. Liang, E.J. Mah, A.F. Yee, M.A. Digman, Integr. Biol. 9 (2017) 145-155. DOI:10.1039/C6IB00193A |
| [17] |
S.S. Chen, X.M. Lu, Q.H. Lu, Chin. Chem. Lett. 28 (2017) 818-826. DOI:10.1016/j.cclet.2016.10.039 |
| [18] |
C.M. Sanderson, M. Way, G.L. Smith, J. Virol. 72 (1998) 1235-1243. DOI:10.1128/JVI.72.2.1235-1243.1998 |
| [19] |
I. Morales, M.A. Carbajal, S. Bohn, et al., Traffic 9 (2008) 1283-1298. DOI:10.1111/j.1600-0854.2008.00762.x |
| [20] |
F. Valderrama, J.V. Cordeiro, S. Schleich, F. Frischknecht, M. Way, Science 311 (2006) 377-381. DOI:10.1126/science.1122411 |
| [21] |
K.L. Roberts, G.L. Smith, Trends Cell Biol. 16 (2008) 472-479. DOI:10.1016/j.tim.2008.07.009 |
| [22] |
Q. Sattentau, Nat. Rev. Microbiol. 6 (2008) 815-826. DOI:10.1038/nrmicro1972 |
| [23] |
U.F. Greber, M. Way, Cell 124 (2006) 741-754. DOI:10.1016/j.cell.2006.02.018 |
| [24] |
C. Beerli, A. Yakimovich, S. Kilcher, et al., Nat. Microbiol. 4 (2019) 216-225. DOI:10.1038/s41564-018-0288-2 |
| [25] |
V. Doceul, M. Hollinshead, L. van der Linden, G.L. Smith, Science 327 (2010) 873-876. DOI:10.1126/science.1183173 |
| [26] |
P. Zhong, L.M. Agosto, J.B. Munro, W. Mothes, Curr. Opin. Virol. 3 (2013) 44-50. DOI:10.1016/j.coviro.2012.11.004 |
| [27] |
F. Pouthas, P. Girard, V. Lecaudey, et al., J. Cell. Sci. 121 (2008) 2406-2414. DOI:10.1242/jcs.026849 |
 2020, Vol. 31
2020, Vol. 31 

