2 Hunan Key Laboratory of Biomedical Materials and Devices, Hunan University of Technology, Zhuzhou 412008, China;
3 The First Affiliated Hospital and the Oncological Institute of Hainan Medical College, Haikou 571101, China
In recent decades, nanomaterials have become widely used in biomedical sciences, owing to their unique physicochemical properties, including thermal conductivity, optical properties and controllable size [1]. Acting as carriers to deliver a variety of drugs to primary, metastatic, and multidrug-resistant cancers through passive and active targeting is one of the most important functions of nanomaterials [2-4]. Some carefully designed nanomaterials can controllably release drugs by light [5, 6], pH [7, 8], magnetic [9, 10], enzyme [11, 12] and redox-responsive factors [13, 14], which greatly reduces damage to normal tissues. In addition, a small number of nanomaterials can damage tumor cells directly through some special mechanisms. For example, some nanomaterials can cause the rapid accumulation of the generation of reactive oxygen species (ROS) which directly induce tumor cells apoptosis [15, 16]. Although many nanomaterials have been shown to be low-toxic or non-toxic to normal cells, to the human body, many nanomaterials are still considered as foreign invaders. For this reason, when immune cells contact some nanomaterials, the innate immune system is rapidly activated by initiating the inflammatory pathway. Therefore, the effect of biomedical nanomaterials on immunity should be clarified, especially for clinical nanodrugs.
In the human immune system, macrophages act as "guardians" against foreign invaders [17]. In response to different stimuli in microenvironments, resting macrophages (M0) undergo functional changes and are polarized into plastic phenotypes which exist within a spectrum between a pro-inflammatory M1 and an antiinflammatory M2 [18-20]. The classic pro-inflammatory M1 phenotype exerts antiproliferative and cytotoxic activities, resulting partly from its ability to secrete reactive nitrogen and oxygen species (e.g., hydrogen peroxide, NO, peroxynitrite, and superoxide) and proinflammatory cytokines including IL-6 (interleukin 6) and TNF-α (tumor necrosis factor-α) [21-23]. In contrast to the M1 phenotype, M2 macrophages have an anti-inflammatory function which turns off damaging immune system activation by generating anti-inflammatory cytokines, such as IL-10 (interleukin 10) and TGF-β (transforming growth factor-β) [23, 24]. Recently, studies have focused on the interaction between nanomaterials and macrophages [25-27]. For instance, super-paramagnetic iron oxide induced M2-type macrophages turn to a high CD86+, TNF-α positive M1-type macrophages [28]. Multiwalled carbon nanotubes activate macrophages to a M1/M2 mixed status [27]. These results show the interaction between nanomaterials and macrophages varies, depending on the types of nanoparticles. Therefore, determination of polarization direction of macrophages stimulated by nanomaterials will assist the development of better biomedical applications of these nanomaterials.
In a previous study, we synthesized a novel nanomedicine (Cy5.5-MSA-G250), consisting of coomassie brilliant blue (G250) and cyanine 5.5 (Cy5.5) with mouse serum albumin (MSA) [29]. Cy5.5-MSA-G250 nanoparticles (CMGNPs) have tumor cell-selective cytotoxicity which can induce tumor cell death without damaging normal cells. In addition CMGNPs are gradually degraded in vivo and excreted in feces and urine. These properties make CMGNPs of great interest for applications in cancer treatment, but the effect of CMGNPs on the immune system is still unclear.
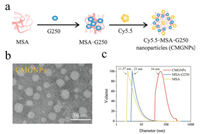
In this work, we aim to initially study the effects of CMGNPs on macrophages. For this purpose, we first prepared CMGNPs according to the previous study [29]. As shown in Fig. 1a, MSAG250 was synthesized by simply mixing MSA and G250 owing to G250 can bind with proteins via hydrophobic interactions and van der Waals' forces [30]. Next, CMGNPs were prepared by covalent conjugation of Cy5.5 to MSA-G250. TEM images clearly showed that only Cy5.5-MSA-G250 has nanostructures, forming spherical nanoparticles with an average diameter of 27.5 ± 5.7 nm (Fig. 1b and Fig. S1 in Supporting information). Meanwhile, as shown in Fig. 1c, the hydrodynamic size of well-dispersed CMGNPs characterized by DLS using the volume model was approximately 54 nm with a polydispersity index (PDI) of 0.259, much larger than that of MSA (11.57 ±2.11 nm, PDI: 0.233) and MSA-G250 (23 ± 4.23 nm, PDI: 0.229), respectively. It is noteworthy that the hydrodynamic size of the micelles measured by DLS was much larger than measured by TEM. This is because the samples were dried, and shrank in size before TEM measurement. The size of CMGNPs measured by both TEM images and DLS analysis suggested that this nanomaterial allows high tumor accumulation and quite a good blood compartment pharmacokinetics [31], and is therefore well suited for cancer treatment.
|
Download:
|
| Fig. 1. Synthesis and characterization of CMGNPs. (a) Schematic illustration of the synthesis of CMGNPs. (b) TEM image of CMGNPs. (c) The hydrodynamic size distribution of MSA, MSA-G250 and CMGNPs. | |
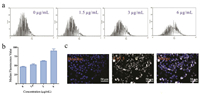
Then, we evaluated the cellular uptake of CMGNPs. Since CMGNPs display the characteristic fluorescence emission peak of Cy5.5, a near-infrared cyanine dye (with a maximum emission wavelength at 700 nm), we investigated the cellular uptake behaviors by measuring the fluorescent intensity of CMGNPtreated RAW264.7 cells. After incubation with CMGNPs for 1 h, the fluorescence intensity of RAW264.7 cells significantly increased as the concentration of CMGNPs increased (Figs. 2a and b), suggesting that CMGNPs are quickly uptaken into RAW264.7 cells. Moreover, confocal microscopic images also demonstrated that a large number of CMGNPs were endocytosed into RAW264.7 cells (Fig. 2c).
|
Download:
|
| Fig. 2. Cellular uptake of CMGNPs. The fluorescence intensity (a) and the median fluorescence value (b) of RAW264.7 cells treated with different doses of CMGNPs (from 0 to 6 μg/mL) for 1 h. (c) Confocal microscopic images of RAW264.7 cells incubating with CMGNPs (6 μg/mL) for 1 h. | |
Phagocytosis plays a crucial role in macrophage-mediated host defense, which leads to internalization and distraction of pathogens [32]. To determine whether CMGNPs affect the phagocytosis of macrophage, we measured the internalization of FITC-labeled dextran by flow cytometry. As shown in Figs. S2a and b (Supporting information), compared with non-treated RAW264.7 cells, CMGNP-treated RAW264.7 cells showed a greater uptake of FITC-dextran, and the relative cell uptake increased to 154.1%, which is only slightly smaller than that of the positive control (treated with LPS). The results indicated that CMGNPs enhances the phagocytosis capacity of RAW264.7 cells. These findings also mean that CMGNP-treated macrophages may shift to active defense. Furthermore, microscope images showed that the cell morphology changed significantly after incubation with CMGNPs. As illustrated in Fig. S3 (Supporting information), the unexposed RAW264.7 cells and G250-treated RAW264.7 cells are dominated by round, spherical cells, whereas the cells incubated with CMGNPs show a distorted morphology with increased length of lamellipodia (Fig. S3). In addition, RAW264.7 cells treated with MSA-G250 complex, similar to the way they treated with CMGNPs, showed polarized morphology. These morphology changes after treated with CMGNPs also suggest that the CMGNPs may induce macrophage polarization. Moreover, we investigated the effect of CMGNPs on the relative cell viability of RAW264.7 cells by MTT assay. As Fig. S4 (Supporting information) indicates, the result of the MTT assay showed that high concentrations of CMGNPs inhibit the proliferation of RAW264.7 cells to a certain extent, but the inhibitory effect of medium or low concentrations of CMGNPs is not obvious.
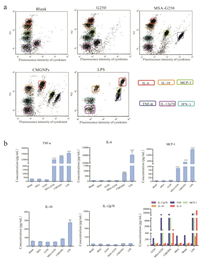
In order to determine the polarization status of RAW264.7 cells upon CMGNPs exposure, we examined the level of expression of cytokines, including IL-6, IL-10, MCP-1(monocyte chemoattractant protein-1), TNF-α, and IL-12p70 (interleukin 12p70), using the CBA Mouse Inflammation Kit. As shown in Figs. 3a and b, TNF-α, a typical proinflammatory cytokine, increased sharply after incubation with CMGNPs (6 μg/mL of G250 concentration), which was over 40 times higher than that for untreated RAW264.7 cells. In addition, the expression of IL-6 and MCP-1 reached 10-fold and 81-fold higher levels after incubation with NPs (6 μg/mL of G250 concentration), respectively, compared with the untreated sample. Meanwhile, G250 did not trigger any significant changes of cytokines. MSA-G250 also induced TNF-α and MCP-1 up-regulation, but it did not affect IL-6 expression. These results illustrate that CMGNPs and MSA-G250 induce macrophage polarization towards the proinflammatory M1 phenotype but G250 does not. Therefore, we speculate that the ability to stimulate RAW264.7 cell polarization originally derives from the combination of MSA and G250, and the conjugation of Cy5.5 to form nanoparticles further enhances this immunostimulatory capacity. Many studies indicate that malignant cells build an immunosuppressive microenvironment which impels macrophage polarization towards a M2-like phenotype [3, 10]. In such an environment, macrophages no longer act as loyal guardians of our body but become accomplices of tumor cells. Thus, the M1 macrophage-induced ability of CMGNPs may help the achievement of better anti-tumor effects.
|
Download:
|
| Fig. 3. Cytokine expression of RAW264.7 cells after different treatments. (a) Cytokine expression level of RAW264.7 cells after treated with G250, MSA-G250, CMGNPs, and LPS as positive control. (b) Data analysis of cytokine expression level of RAW264.7 cells after treated with G250, MSA-G250, CMGNPs and LPS (LPS: 100 ng/mL; others: 6 μg/mL of G250 content). | |
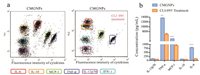
TLR4 is one of the essential pattern-recognition receptors (PRRs) of macrophages, and plays a key role in pathogen recognition and activation of antigen-presenting cells [33, 34]. To date, several studies have revealed that nanomaterials, such as graphene oxide activate TLR4 signaling and increase the secretion of pro-inflammatory cytokines [35]. To determine whether TLR4 is involved in the induction of immune responses induced by CMGNPs, we used TLR4 inhibitor CLI-095 for pre-treatment in cells exposed to CMGNPs. As shown in Figs. 4a and b, TLR4 inhibitor markedly represses the expression of TNF-α, MCP-1 and IL-6 (by approximately 50%, 18.7%, and 12.5%, respectively) in RAW264.7 cells after CMGNPs treatment, suggesting that TLR4 plays an indispensable role in CMGNP-induced inflammation.

|
Download:
|
| Fig. 4. Changes of cytokine expressions of CMGNP-treated RAW264.7 cells pretreatment with CLI-095. (a) Cytokine expression level of CMGNP-treated RAW264.7 cells pretreated or not with CLI-095 measured by flow cytometry. (b) Data analysis of cytokine expression level of CMGNP-treated RAW264.7 cells pretreated or not with CLI-095 (CMGNPs: 6 μg/mL; CLI-095: 1 μg/mL). | |
In conclusion, in vitro experiments clearly show that CMGNPs not only enhance the phagocytosis capacity of RAW264.7 cells, but also promote M1 polarization, associated with changes in cell morphology and increased expression of inflammatory cytokines. Moreover, the observed M1 macrophage' polarization triggered by CMGNPs is abolished after adding TLR4 inhibitor, CLI095, suggesting that TLR4 is involved in CMGNP-induced inflammation. This ability to induce M1 polarization may be beneficial in assisting to CMGNPs to achieve better anticancer effects in clinical trials.
AcknowledgmentsWe gratefully acknowledge the laboratory facilities and equipment support provided by the public research lab of Hainan medical college. This work was financially supported by the National Natural Science Foundation of China (Nos. 81660592, 81660301 and 81860037) and the Key New and High-Tech Project of the Department of Science and Technology of Hainan Province (No. ZDYF2016023).
Appendix A. Supplementary dataSupplementary material related to this article canbefound, in the online version, at doi:https://doi.org/10.1016/j.cclet.2019.03.001.
| [1] |
A.P. Blum, J.K. Kammeyer, A.M. Rush, et al., J. Am. Chem. Soc. 137 (2015) 2140-2154. DOI:10.1021/ja510147n |
| [2] |
E. Blanco, H. Shen, M. Ferrari, Nat. Biotechnol. 33 (2015) 941-951. DOI:10.1038/nbt.3330 |
| [3] |
X. Li, L. Hong, T. Song, et al., J. Biomed. Nanotechnol. 13 (2017) 747-757. DOI:10.1166/jbn.2017.2383 |
| [4] |
Z. Xi, B. Zheng, Nanosci. Nanotechnol. Lett. 10 (2018) 309-319. DOI:10.1166/nnl.2018.2624 |
| [5] |
M.S. Yavuz, Y. Cheng, J. Chen, et al., Nat. Mater. 8 (2009) 935-939. DOI:10.1038/nmat2564 |
| [6] |
Q. Liu, C. Zhan, D.S. Kohane, Bioconjugate Chem. 28 (2017) 98-104. DOI:10.1021/acs.bioconjchem.6b00448 |
| [7] |
J. Liu, Y. Huang, A. Kumar, et al., Biotechnol. Adv. 32 (2014) 693-710. DOI:10.1016/j.biotechadv.2013.11.009 |
| [8] |
L. Li, Y. Lu, M. Zhang, et al., Nanosci. Nanotechnol. Lett. 10 (2018) 13-22. DOI:10.1166/nnl.2018.2588 |
| [9] |
K. Zhu, Y. Ju, J. Xu, et al., Acc. Chem. Res. 51 (2018) 404-413. DOI:10.1021/acs.accounts.7b00407 |
| [10] |
G.E. Choi, M.S. Kang, Y.J. Kim, et al., J. Nanosci. Nanotechnol. 19 (2019) 675-679. DOI:10.1166/jnn.2019.15910 |
| [11] |
Q. Hu, P.S. Katti, Z. Gu, et al., Nanoscale 6 (2014) 12273-12286. DOI:10.1039/C4NR04249B |
| [12] |
R. Chandrawati, Exp. Biol. Med. 241 (2016) 972-979. DOI:10.1177/1535370216647186 |
| [13] |
L. Han, X.Y. Zhang, Y.L. Wang, et al., J. Control. Release 259 (2017) 40-52. DOI:10.1016/j.jconrel.2017.03.018 |
| [14] |
L. Qiu, L. Zhao, C. Xing, et al., Chin. Chem. Lett. 29 (2018) 301-304. DOI:10.1016/j.cclet.2017.09.048 |
| [15] |
S.K. Dash, T. Ghosh, S. Roy, et al., J. Appl. Toxicol. 34 (2014) 1130-1144. DOI:10.1002/jat.v34.11 |
| [16] |
D. Trachootham, J. Alexandre, P. Huang, et al., Nat. Rev. Drug Discov. 8 (2009) 579-591. DOI:10.1038/nrd2803 |
| [17] |
S.J. Galli, N. Borregaard, T.A. Wynn, Nat. Immunol. 12 (2011) 1035-1044. DOI:10.1038/ni.2109 |
| [18] |
J. Linares, A.B. Fernandez, M.J. Feito, et al., J. Mater. Chem. B 4 (2016) 1951-1959. DOI:10.1039/C6TB00014B |
| [19] |
M. Motskin, D.M. Wright, K. Muller, et al., Biomaterials 30 (2009) 3307-3317. DOI:10.1016/j.biomaterials.2009.02.044 |
| [20] |
X. Miao, X. Leng, Q. Zhang, Int. J. Mol. Sci. 18 (2017) 336. DOI:10.3390/ijms18020336 |
| [21] |
K.M. Hotchkiss, G.B. Reddy, S.L. Hyzy, et al., Acta Biomater. 31 (2016) 425-434. DOI:10.1016/j.actbio.2015.12.003 |
| [22] |
D.M. Mosser, J.P. Nat. Rev. Immunol. 8 (2008) 958-969. DOI:10.1038/nri2448 |
| [23] |
M. Song, T. Liu, C. Shi, et al., ACS Nano 10 (2016) 633-647. DOI:10.1021/acsnano.5b06779 |
| [24] |
P.J. Murray, Nat. Rev. Immunol. 11 (2011) 723-737. DOI:10.1038/nri3073 |
| [25] |
A. Marucco, E. Gazzano, D. Ghigo, et al., Nanotoxicology 10 (2016) 1-9. |
| [26] |
C. Farrera, B. Fadeel, Eur. J. Pharma. Biopharm. European. 95 (2015) 3-12. DOI:10.1016/j.ejpb.2015.03.007 |
| [27] |
J. Meng, X. Li, C. Wang, et al., ACS Appl. Mater. Interfaces 7 (2015) 3180-3188. DOI:10.1021/am507649n |
| [28] |
S. Zanganeh, G. Hutter, R. Spitler, et al., Nat. Nanotech. 11 (2016) 986-994. DOI:10.1038/nnano.2016.168 |
| [29] |
Z. Lu, F.Y. Huang, R. Cao, et al., ACS Appl. Mater. Interfaces (2018) 26028-26038. |
| [30] |
Y.J. Wei, K.A. Li, et al., Talanta 44 (1997) 923-930. DOI:10.1016/S0039-9140(96)02140-6 |
| [31] |
S.D. Perrault, C. Walkey, T. Jennings, et al., Nano Lett. 9 (2009) 1909-1915. DOI:10.1021/nl900031y |
| [32] |
Z.S. Gan, Q.Q. Wang, J.H. Li, et al., Mediat. Inflamm. 2017 (2017) 8570818. |
| [33] |
W. Wei, H.T. Xiao, W.R. Bao, et al., J. Ethnopharmacol. 179 (2016) 243-252. DOI:10.1016/j.jep.2015.12.060 |
| [34] |
P.R. Rauta, M. Samanta, H.R. Dash, et al., Immunol. Lett. 158 (2014) 14-24. DOI:10.1016/j.imlet.2013.11.013 |
| [35] |
J. Ma, R. Liu, X. Wang, et al., ACS Nano 9 (2015) 10498-10515. DOI:10.1021/acsnano.5b04751 |
 2019, Vol. 30
2019, Vol. 30 

