Adenosine triphosphate (ATP, nucleoside triphosphates), is the major energy source power in living organisms. Moreover, it plays a key role in regulation of many biological pathways and is extracellular signal mediator in many biological processes [1-3]. So ATP has been used as an indicator for many diseases such as cardiovascular diseases [4], Parkinson's disease [5], and Alzheimer's disease [6, 7]. Consequently, the monitoring of ATP and other micromolecules in biological sample has attracted considerable attention [8-12]. At present, owning to the high affinity between ATP and its aptamer [13], aptamer-based fluorescence methods have attracted increasing interest for they are not only sensitive, selective, operated simply and quantified feasibly, but also could be read out easily, need low sample volume [14-16]. For example, Zhang's group [16] developed DNA nanoprism with a split aptamer for ATP detection in cells. However, some of them need nanomaterials as quencher, the poor stability of nanomaterials limits the probe's performance [14]. Others need expensive double-labelled probes [16]. So it is still necessary to search some simple methods with only single-labelled probe for ATP detection in complex media.
Owning to the remarkable and unique properties, the applications of supramolecular assembly have attracted tremendous interest on the horizon of analytical science [17-20]. Cyclodextrins (CDs), the most commonly used host molecules, which consisted of D-glucopyranose units, are a serious of torus-shaped cyclic oligosaccharides. The outer surface of CDs is hydrophilic and inner cavity is mildly lipophilic. Therefore, CDs are not only water-soluble, but also can form host–guest inclusion complexes with less polar fluorophores which can geometrically fit the lipophilic inner cavity. This molecular recognition could significantly change the microenvironment of fluorophores from hydrophilic phase to the lipophilic phase and could manifests in analytical domain such as the enhancement of sensitivity [17, 21]. Some cyclodextrin-based methods have been developed for detection of mycotoxin, nucleic acid, tryptophan, aflatoxins, etc., with enhanced fluorescence spectral response and higher sensitivity [22-26]. Asforcyclodextrin contained polymers (polyβ-CD), there are multiple exterior β-CDs attached on polymers, which could interact with fluorophores and so the recognition ability of polyβ-CDishigherthan β-CDsmonomer and the affinity of the β-CDs-fluorophores complex is higher than β-CD monomer-fluorophores, with enhanced and stabilized fluorescence [21, 27-31]. Markus Hollas [21] has reported that the overall complexation constant of the β-cyclodextrin polymers and pyrene increased by more than 2 orders of magnitude compared to monomer β-cyclodextrin, which perhaps be attribute to the excellent fluorescence enhancement of pyrene by β-cyclodextrin polymer. Due to the advantages of polyβ-CD, a series of remarkable method that based on electroneutral and cationic β-cyclodextrin polymers fluorescence enhancement have been developed for T4 polynucleotide kinase activity [27], alkaline phosphatase [28] and DNA [32] sensitive detection. Furthermore, the fluorescence probes used in this work need only single-labeled by pyrene, which was inexpensive, stable and ordered conveniently from TaKaRa Bio. Inc. (Dalian, China).
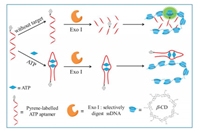
Herein, electroneutral polyβ-CD was exploited for facile and sensitive ATP detection based on the fact that the fluorescence emission of pyrene could be greatly enhanced by electroneutral polyβ-CD through host-guest interaction. The working mechanism of the sensing method was illustrated in Scheme 1. The proposed method consists of a 3'-pyrene-labelled ATP-binding aptamer as the fluorescence probe, electroneutral polyβ-CD as the fluorescence enhancement component, ATP as target. The probe could be digested by exonuclease Ⅰ (Exo Ⅰ) to generate mononucleotides with pyrene labelled on, and the pyrene labelled on the DNA fragment could facilely enter into the cavity of polyβ-CD, result in prominent enhanced fluorescence. While in the presence of ATP, the target and its aptamer could combine together and obtained hairpin complex could not be cleaved by Exo Ⅰ. The pyrene attached on probe DNA was difficult to enter into the lipophilic cavity of polyβ-CD, for the steric hindrance of the formed hairpin complex. Then the fluorescence emission of pyrene kept weak. Therefore the ATP could be easily detected by simply measuring the fluorescence intensity of the system.
|
Download:
|
| Scheme 1. Schematic diagram of β-cyclodextrin polymer fluorescence enhancement based assay for sensitive ATP detection. | |
The electroneutral polyβ-CD was synthesized according to the literatures [21] and in accordance with our previous work [32]. The preparation and characterization of cyclodextrin polymer was detailed in the Supporting information. We measured the molecular weight of polyβ-CD by waters-515 gel permeation chromatography and found it was about 94.4 KD.
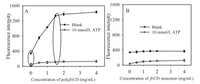
The fluorescence enhancement capacity of polyβ-CD to pyrene was first investigated and shown in Fig. 1. If polyβ-CD or β-CD monomer was all absent, the fluorescence was weak (in Fig. 1, the points when the concentration of polyβ-CD or β-CD monomer was 0 mg/mL). In the case of polyβ-CD presented (Fig. 1A), the fluorescence of the system (the plots of 10 mmol/L ATP) kept weak. While in case of the 0 mmol/L ATP (the plots of 0 mmol/L ATP in Fig. 1A), the fluorescence of the system increased with the increasing concentration of polyβ-CD from 0 to 4.0 mg/mL and was more than quadrupled in the case of 1.5 mg/mL polyβ-CD. In the case of various concentrations of CD monomer presented (Fig. 1B), the fluorescence emission of the system in the presence/absence of ATP kept low and little changed. The results were in accordance with our previous report [27]. It was reported that the overall complexation constant of the polyβ-CD and pyrene increased by more than two orders of magnitude [21] compared with β-CD monomer, which may be resulted in the prominent fluorescence enhancement capability of polyβ-CD. The prominent fluorescence enhancement capability of polyβ-CD could greatly improve the sensitivity of our proposed method.

|
Download:
|
| Fig. 1. Fluorescence emission intensity of the system in the case of polyβ-CD (A) or β-CD monomer (B) presented. The concentration of fluorescence probe was 50 nmol/L. The activity of Exo Ⅰ and reaction time was 80 U/mL and 30 min respectively. The error bars were the standard deviations of triplicate measurements in this work. | |
To investigate the feasibility of our proposed method, the fluorescence emission of the system with 0 or 10 mmol/L ATP respectively was detected (Fig. 2). When ATP was absent, the fluorescence intensity of the system was high. When ATP was introduced, it bound to the 3'-pyrene-labelled aptamer and hindered the digestion of the aptamer. The steric hindrance of ATP-aptamer hairpin complex inhibited the host-gest interaction of pyrene and polyβ-CD, so the fluorescence intensity kept weak. These results confirmed the feasibility of our proposed strategy.

|
Download:
|
| Fig. 2. Fluorescence emission spectra of the system when the ATP was 0 and 10 mmol/L respectively. The polyβ-CD and probe included in the system were 1.5 mg/mL and 50 nmol/L, respectively. The activity and reaction of Exo Ⅰ was 80 U/mL and 30 min respectively. | |
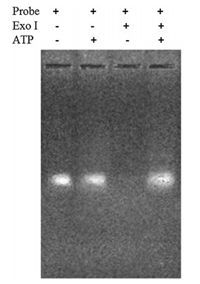
We further used gel electrophoresis to verity ATP-induced protection of 3'-pyrene-labelled aptamer from digestion (Fig. 3). In the absence of Exo Ⅰ and ATP, bright bands were obtained (from left to right, the first and second lane, respectively), indicating no 3'-pyrene-labelled aptamer digestion. No obvious band was observed in the third lane in the case of Exo Ⅰ present and ALP absent, indicating digestion of 3'-pyrene-labelled aptamer by Exo Ⅰ. Conversely, in the fourth lane (Exo Ⅰ and ALP were all present), we could obtained a bright band, indicated that 3'-pyrene-labelled aptamer was protected from digestion by ATP.
|
Download:
|
| Fig. 3. The image of agarose gel electrophoresis. | |
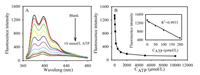
To characterize the performance of our proposed analytical method, the fluorescence emission spectra of the system at various amount of ATP under the optimum condition (the optimization of experimental conditions was detailed in Fig. S1 in Supporting information) was measured. As shown in Fig. 4, it was found that the fluorescence intensity of the system decreased with incremental ATP from 0 to 10 mmol/L. Fig. 4B showed that the fluorescence response was inverse proportional to ATP in the range of 0.00 - 200 μmol/L with a regression equation expressed as F = 1294.9 - 4.1030×CATP (μmol/L) (Fig. 4B, inserted) and the correlation coefficient was 0.9933. The detection limit was 11 μmol/L (S/N = 3). The high sensitivity of the proposed method should be due to the excellent fluorescence enhancement capability of polyβ-CD.

|
Download:
|
| Fig. 4. (A) Fluorescence emission spectra of the system with various concentration of ATP (0, 10, 50, 100, 200, 500, 1000, 2000, 5000, 10, 000 μmol/L, respectively); (B) Plot of F vs. ATP concentration. Inset chart is the linear relation plot of the fluorescence emission intensity to the concentration of ATP from 0 to 200 μmol/L. | |
High specificity was necessary for an analytical method. GTP, CTP and UTP are structures analogues with ATP and usually coexisting in biology samples. So GTP, CTP and UTP were chosen as interfering substances whose concentration was the same with the target (Fig. 5). (F0-F)/F0 was employed as fluorescence factor in this work (F0, the fluorescence intensities of the system in the absence of target; F, the fluorescence intensities of the system in the presence of target). The values of (F0-F)/F0 were about only 0.1 in the presence of analogues, while as for ATP, (F0-F)/F0 was above 0.9, 9 times the value of these analogues, which indicated that the proposed method has satisfied specificity for discriminating ATP from its analogues. The high specificity is possible attributed to the high affinity aptamers of biological micromolecules [33, 34].

|
Download:
|
| Fig. 5. Specificity study. ATP, GTP, CTP and UTP were all 10 mmol/L. | |
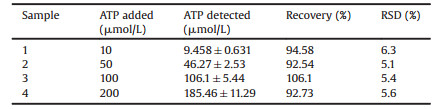
Human serum sample was chosen to inspect the application of our proposed method in biological sample detection. We spiked ATP standard solution in 10% human serum to simulate the biological sample (Table 1). The standard addition recovery was within the scope of 92.54%–106.1%, and relative standard deviation (RSD) was below 6.3%, which indicated that the proposed method can be used in biological samples detection with satisfied results.
|
|
Table 1 ATP detection in human serum samples (n = 3). |
In conclusion, combining with the prominent fluorescent enhancement of polyβ-CD to pyrene and the specific high affinity aptamer, a facile and sensitive fluorescence method has been constructed. One of the advantages of the developed method is that the probe needs only single-labelled, avoids complicated doublelabel or the use of nanomaterial. Besides, owning to the prominent fluorescence enhancement capability of polyβ-CD, the method has high sensitivity (the detection limit for ATP: 11 μmol/L). Moreover, it provided high specificity of ATP detection against its structures analogues. More importantly, it can be used in the determination of ATP in the biological samples, and holds great potential for the detection of micromolecules in biological samples.
AcknowledgmentsThis work was financially supported by the National Natural Science Foundation of China (Nos. 21305002 and 21705002) and the Scientific Research Foundation of Anhui Agricultural University (No. yj2017-19).
Appendix A. Supplementary dataSupplementary material related to this article can be found, in the online version, at doi:https://doi.org/10.1016/j.cclet.2019.03.055.
| [1] |
X. Feng, Y. Jia, P. Cai, et al., ACS Nano 10 (2016) 556-561. DOI:10.1021/acsnano.5b05579 |
| [2] |
D.R. Palleros, K.L. Raid, L. Shi, et al., Nature 365 (1993) 664-666. DOI:10.1038/365664a0 |
| [3] |
X. Zheng, R. Peng, X. Jiang, et al., Anal. Chem. 89 (2017) 10941-10947. DOI:10.1021/acs.analchem.7b02763 |
| [4] |
H. Yokoshiki, M. Sunagawa, T. Seki, et al., Am. J. Physiol. 274 (1998) C25-C37. DOI:10.1152/ajpcell.1998.274.1.C25 |
| [5] |
A.V. Gourine, E. Llaudet, N. Dale, et al., Nature 436 (2005) 108-111. DOI:10.1038/nature03690 |
| [6] |
L. Annunziato, G. Pignataro, R.G. Di, Pharmacol. Rev. 56 (2004) 633-654. DOI:10.1124/pr.56.4.5 |
| [7] |
C. Zhang, R.A. Rissman, J. Feng, J. Alzheimers Dis. 44 (2015) 375-378. DOI:10.3233/JAD-141890 |
| [8] |
X. Zhang, C. Song, K. Yang, et al., Anal. Chem. 90 (2018) 4968-4971. DOI:10.1021/acs.analchem.7b05442 |
| [9] |
Z. Qing, L. Hou, L. Yang, et al., Anal. Chem. 88 (2016) 9759-9765. DOI:10.1021/acs.analchem.6b02720 |
| [10] |
L. Yang, Z. Qing, C. Liu, et al., Anal. Chem. 88 (2016) 9285-9292. DOI:10.1021/acs.analchem.6b02667 |
| [11] |
X. Zhang, R. Kong, Q. Tan, et al., Talanta 169 (2017) 1-7. DOI:10.1016/j.talanta.2017.03.050 |
| [12] |
Q. Tan, R. Zhang, R. Kong, et al., Microchim. Acta 185 (2018) 44-50. DOI:10.1007/s00604-017-2603-7 |
| [13] |
D.E. Huizenga, J.W. Szostak, Biochemistry 34 (1995) 656-665. DOI:10.1021/bi00002a033 |
| [14] |
X.M. Hai, N. Li, K. Wang, et al., Anal. Chim. Acta 998 (2018) 60-66. DOI:10.1016/j.aca.2017.10.028 |
| [15] |
B. Huang, Z. Geng, S. Yan, et al., Anal. Chem. 89 (2017) 8816-8821. DOI:10.1021/acs.analchem.7b01212 |
| [16] |
X. Zheng, R. Peng, X. Jiang, et al., Anal. Chem. 89 (2017) 10941-10947. DOI:10.1021/acs.analchem.7b02763 |
| [17] |
L. Szente, J. Szemán, Anal. Chem. 85 (2013) 8024-8030. DOI:10.1021/ac400639y |
| [18] |
A. Harada, Acc. Chem. Res. 34 (2001) 456-464. DOI:10.1021/ar000174l |
| [19] |
K. Uekama, F. Hirayama, T. Irie, Chem. Rev. 98 (1998) 2045-2076. DOI:10.1021/cr970025p |
| [20] |
A.R. Hedges, Chem. Rev. 98 (1998) 2035-2044. DOI:10.1021/cr970014w |
| [21] |
M. Hollas, M.A. Chung, J. Adams, J. Phys. Chem. B 102 (1998) 2947-2953. DOI:10.1021/jp9800719 |
| [22] |
C.M. Maragos, M. Appell, V. Lippolis, et al., Food Addit. Contam. A 25 (2008) 164-171. DOI:10.1080/02652030701564555 |
| [23] |
G. Zhilong, Z. Zhujun, Microchim. Acta 126 (1997) 325-328. DOI:10.1007/BF01242340 |
| [24] |
C. Cucci, A.G. Mignani, C. Dall'Asta, et al., Sens. Actuators B-Chem. 126 (2007) 467-472. DOI:10.1016/j.snb.2007.03.036 |
| [25] |
Z. Qing, X. He, J. Huang, et al., Anal. Chem. 86 (2014) 4934-4939. DOI:10.1021/ac500834g |
| [26] |
J. Sun, S. Wang, F. Gao, Langmuir 32 (2016) 12725-12731. DOI:10.1021/acs.langmuir.6b03002 |
| [27] |
C. Song, X. Yang, K. Wang, et al., Chem. Commun. 51 (2015) 1815-1818. DOI:10.1039/C4CC08991J |
| [28] |
C. Song, X. Yang, K. Wang, et al., Spectrochim. Acta A 156 (2016) 131-137. DOI:10.1016/j.saa.2015.12.001 |
| [29] |
P. Liu, S. Sun, X. Guo, et al., Anal. Chem. 87 (2015) 2665-2671. DOI:10.1021/ac503301q |
| [30] |
M. Shaikh, J. Mohanty, M. Sundararajan, et al., J. Phys. Chem. B 116 (2012) 12450-12459. DOI:10.1021/jp3087368 |
| [31] |
L. Wang, C. Zhong, P. Xue, et al., J. Org. Chem. 76 (2011) 4874-4883. DOI:10.1021/jo2007829 |
| [32] |
C. Song, B. Li, X. Yang, et al., Analyst 142 (2017) 224-228. DOI:10.1039/C6AN02269C |
| [33] |
J.A. Cruz-Aguado, G. Penner, Anal. Chem. 80 (2008) 8853-8855. DOI:10.1021/ac8017058 |
| [34] |
J.A. Cruz-Aguado, G. Penner, J. Agric. Food Chem. 56 (2008) 10456-10461. DOI:10.1021/jf801957h |
 2019, Vol. 30
2019, Vol. 30