b National Engineering Research Center for Solid Preparation Technology of Chinese Medicines, Jiangxi University of Traditional Chinese Medicines, Nanchang 330006, China
Multi-walled carbon nanotubes (MWCNTs) are carbon-based nanoparticles (NPs) of tubular structures and a high aspect ratio (length-to-width ratio), which have already been used in many commercial products [1]. For example, MWCNTs are used in electronics [2], energy storage [3], and nanomedicine [4, 5]. The increasing use of MWCNTs may lead to exposure to NPs in daily life and this raises concerns about their impact on human health. Indeed, previous studies already found that MWCNT exposure leads to adverse effects, including but not limited to oxidative stress [6], DNA damage [7], and cell death [8]. However, it is still necessary to further study the toxic effects of MWCNTs and the molecular mechanisms that mediate these effects.
Recently, we found that MWCNT exposure alters the mRNA levels of endoplasmic reticulum (ER) stress and autophagic genes, which is likely responsible for the cytotoxicity of MWCNT to human umbilical vein endothelial cells (HUVECs) [9] and alveolarHUVEC co-cultures [10], as well as MWCNT-induced lipid accumulation in hepatocytes [11]. The ER is an organelle that is crucial for proper protein folding, lipid synthesis, and calcium storage. Extensive studies have shown that NP exposure perturbs the normal function of the ER and activates the ER stress response, which consequently mediate the adverse health effects [12, 13]. ER stress can also activate autophagy in an attempt to degrade organelles, macromolecules, and particles, although the association of ER stress and autophagy is still largely unknown and requires further study [14]. Interestingly, cumulative evidence suggests that NPs promote autophagic dysfunction, leading to toxicological responses [8, 12]. Although previous studies have shown that NPs can affect the ER stress/autophagy response, the influence of excessive nutrients on this response may have been overlooked. Because the ER stress/autophagy response can also be modulated by excessive nutrients [15, 16], co-exposure to NPs and excessive nutrients may induce a different ER stress/autophagy response.
Therefore, the purpose of this study is to evaluate the different ER stress/autophagy responses in macrophages derived from THP- 1 monocytes (denoted as THP-1 macrophages) following exposure to carboxylated MWCNTs (c-MWCNTs) pre-treated with either BSA or BSA-complexed free fatty acid (denoted as FFA). C-MWCNTs were used because MWCNTs are modified with carboxyl groups to improve their stability and to conjugate therapeutic molecules [17]. Recently, we found that c-MWCNTs promote lipid accumulation, probably by inducing autophagic dysfunction [11]. Free fatty acids were studied because they are typical lipids that can influence the ER stress/autophagy response [15, 16]. Moreover, we and others recently reported that the adsorption of free fatty acids onto NP surface changed the colloidal stability and/or toxicity of NPs [18-20]. However, the possible impact of lipids on the NPinduced ER stress/autophagy response has not been studied in detail. THP-1 macrophages were used in the current study because they are phagocytes that can phagocytose both particles and excessive lipids and our recent studies have shown that free fatty acids alter NP-induced toxicological responses in THP-1 macrophages [19, 21].
The overall study design is illustrated in Fig. S1 (Supporting information). After pre-incubation of c-MWCNTs with BSA or FFA, the changes in atomic force microscopy (AFM) height, hydrodynamic size, and zeta potential were determined to indicate the surface coating effects of BSA and FFA to c-MWCNTs. THP-1 macrophages were incubated with c-MWCNTs coated with BSA or FFA, and then cell counting kit-8 (CCK-8) as well as neutral red uptake assays were used to indicate cytotoxicity. To indicate oxidative stress levels, intracellular reactive oxygen species (ROS) and glutathione (GSH) were measured. Real-time RT-PCR was performed to measure the mRNA levels of the ER stress genes, DDIT-3 (DNA damage-inducible transcript 3; also named as CHOP [C/EBP homologous protein]), XBP-1s (spliced X-box binding protein 1), and HSPA5 (heat shock protein family A (Hsp70) member 5) and the autophagic genes, ATG5 (autophagy related 5), PLIN2 (perilipin 2), and BECN1 (beclin 1), whereas Western blot was done to measure the protein levels of chop, p-chop, LC3B, ATG5, and beclin-1. Finally, since excessive lipids can promote lipid accumulation and the ER stress/autophagy response is closely related with lipid droplet formation in macrophage foam cells [22-24], this study determined lipid accumulation by oil red staining, cytokine release by ELISA, CD36 and MSR1 (macrophage scavenger receptor 1) mRNA levels by real-time RT-PCR, as well as scavenger receptor protein levels by Western blot.
A representative AFM image (Fig. S5 in Supporting information) indicated uniform surfaces of c-MWCNTs. The average diameter and length of uncoated c-MWCNTs were measured as 21 ±3.1 nm (range 15–30 nm) and 223.4 ±77.5 nm (range 100–450 nm), respectively. The average AFM diameter of c-MWCNTs increased to 44.9 ± 21.1 nm after BSA pre-incubation and 52.3 ± 21.5 nm after FFA pre-incubation (Fig. 1A). It should be noted that, after either BSA or FFA incubation, c-MWCNTs appeared to be less adhesive to the mica surface, leading to relatively fewer NPs in each independent area. This may be due BSA and FFA changing the surface charge and/or dispersity of c-MWCNTs, as indicated by DLS measurements shown below. C-MWCNTs exhibited two peaks of hydrodynamic size at approximately 190 nm and 5580 nm, and a negative zeta potential, with a peak at approximately -29 mV. After pre-incubation with BSA or FFA, c-MWCNTs showed slightly increased average hydrodynamic size as well as decreased absolute zeta potential value and polydispersity index (PDI). This tendency was more obvious for FFA than for BSA (Figs. 1B and C, Table S4 in Supporting information). In combination, it can be concluded from these data that BSA and FFA were coated onto the surface of MWCNTs. Previous studies have shown similar effects, in that incubation of NPs with proteins resulted in the formation of protein corona that altered the colloidal stability of carbon-based NPs [25-28]. In the current study, our data further indicated that FFA could coat the surface of MWCNTs as effectively as BSA.

|
Download:
|
| Fig. 1. Changes in colloidal aspects of c-MWCNT (code XFM24) brought by BSA or FFA pre-incubation. (A) AFM diameter. Data were reported as the mean ± SD (n = 20 for each sample). (B) Distribution of hydrodynamic size. (C) Distribution of zeta potential. | |
The interference assay indicates no direct reaction between cMWCNTs and CCK-8 (Table S1 in Supporting information) or neutral red (Table S2 in Supporting information). CCK-8 (Fig. S6A in Supporting information) or neutral red uptake assays (Fig. S6B in Supporting information) showed no cytotoxicity of c-MWCNTs pre-incubated with BSA or FFA. In contrast, the positive control (2% Triton X-100) significantly induced cytotoxicity as indicated by both assays (P < 0.01). Meanwhile, exposure to BSA-coated cMWCNTs or FFA-coated c-MWCNTs did not significantly (P > 0.05) affect intracellular ROS (Fig. S7A in Supporting information) or GSH (Fig. S6B) levels in THP-1 macrophages. The lack of cytotoxicity could be due to that the c-MWCNTs used in this study had relatively short fibers. We have recently shown that long fibers are cytotoxic to human endothelial cells [9, 10]. Pre-incubation with FFA did not significantly augment the cytotoxicity of c-MWCNTs, which could be explained by that lipids, at non-cytotoxic concentrations, do not enhance the cytotoxic potential of NPs [29]. The lack of oxidative stress may also account for the lack of cytotoxicity of these c-MWCNTs, since oxidative stress is generally accompanied with cytotoxicity of MWCNTs [6].
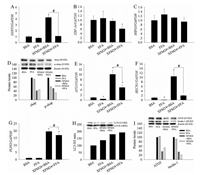
C-MWCNTs pre-incubated with BSA (P < 0.01), but not the ones pre-incubated with FFA (P > 0.05), significantly increased DDIT3 expression (Fig. 2A). Moreover, exposure to BSA-coated c-MWCNTs led to significantly higher DDIT3 expression compared with exposure to FFA-coated c-MWCNTs (P < 0.01). XBP-1s mRNA levels were only significantly decreased by c-MWCNTs pre-incubated with FFA (P < 0.05, Fig. 2B). The expression of HSPA5 remained unaltered by c-MWCNTs either pre-incubated with BSA or FFA (P > 0.05, Fig. 2C). The protein levels of chop and p-chop were decreased in THP-1 macrophages by the exposure of BSA or FFA pre-incubated c-MWCNTs (Fig. 2D). For the biomarkers of autophagy, the mRNA levels of ATG5 were significantly downregulated by FFA exposure (P < 0.01) but up-regulated by cMWCNTs coated with BSA or FFA (P < 0.01). However, BSA-coated c-MWCNTs induced significantly higher mRNA levels of ATG5, compared to FFA-coated c-MWCNTs (P < 0.01, Fig. 2E). The mRNA levels of BECN1 were significantly down-regulated by FFA (P < 0.01), remained unaltered by FFA-coated c-MWCNTs (P > 0.05), but significantly up-regulated by BSA-coated c-MWCNTs (P < 0.01, Fig. 2F). Similarly, BECN1 mRNA levels were significantly higher by BSA-coated c-MWCNTs compared with FFA-coated c-MWCNTs (P < 0.01). For PLIN2 (Fig. 2G), both BSA-coated c-MWCNTs and those pre-incubated with FFA significantly increased PLIN2 expression (P < 0.01), but PLIN2 expression was significantly higher by BSA-coated c-MWCNTs than FFA-coated c-MWCNTs (P < 0.01). For autophagic proteins, exposure to BSA-coated c-MWCNTs or FFA increased the ratio of LC3-II/LC3-I (Fig. 2H), but decreased the protein levels of ATG5 and beclin-1 (Fig. 2I).
|
Download:
|
| Fig. 2. The changes of ER stress/autophagy pathways. THP-1 macrophages were exposed to 128 μg/mL of c-MWCNTs (code XFM24) pre-incubated with BSA or FFA for 24 h. After exposure, ER stress genes DDIT3 (A), XBP-1s (B), and HSPA5 (C), as well as autophagic genes ATG5 (E), BECN1 (F), and PLIN2 (G), were measured by real-time RT-PCR. Protein levels (control = 100%) of chop, p-chop (D), LC3-I, LC3-II (H), ATG5 and BECN1 (I) were measured by Western blot. Data represent mean ± SE of three independent experiments (n = 3). *P < 0.05, #P < 0.01 compared with control. | |
These data showed that BSA-coated c-MWCNTs significantly increased the mRNA levels of ER stress/autophagic genes, but at the protein level, only increased the conversion of LC3-I to LC3-II. The significant increase in DDIT3 expression by BSA-coated c-MWCNTs may indicate a modest increase in ER stress, which we have recently observed in MWCNT-exposed human endothelial cells [9], alveolar-endothelial co-cultures [10], and HepG2 cells [11]. The induction of autophagic biomarkers by BSA-coated c-MWCNTs is also in general agreement with previous observations that carbonaceous NPs can activate autophagy [8]. In the present study, it was further shown that the induction of the ER stress/ autophagic pathway by NPs may be modulated by the presence of biological molecules, since FFA-coated c-MWCNTs induced a significantly lower level of activation of ER stress/autophagic genes. In addition, the protein levels of chop, p-chop, ATG5, and beclin-1 were also decreased after exposure to FFA-coated cMWCNTs. Although it is necessary to further confirm the activation of the ER stress/autophagic pathway by MWCNTs and characterize the association between ER stress and autophagy, results from this study indicate that the biological microenvironment at the NP entry site, as well as the interactions between NPs and nutrients, should be taken into account in order to better evaluate the possible influence of NPs on the ER stress/autophagic signaling pathway.
The expression of CD36 (Fig. 3A) and MSR1 (Fig. 3B) were significantly up-regulated in FFA-coated c-MWCNT-exposed THP-1 macrophages (P < 0.01). Moreover, FFA-coated c-MWCNTs led to significantly higher mRNA levels of CD36 and MSR1 compared to those pre-incubated with BSA. Western blot analysis indicated that the protein levels of CD36 increased following exposure to BSAcoated or FFA-coated c-MWCNTs, but MSR1 protein levels were only modestly increased by BSA-coated c-MWCNTs (Fig. 3C).

|
Download:
|
| Fig. 3. mRNA and protein levels of scavenger receptors. THP-1 macrophages were exposed to 128 μg/mL c-MWCNTs (code XFM24) pre-incubated with BSA or FFA for 24 h, and mRNA levels of CD36 (A) and MSR1 (B) were measured by real-time RT-PCR. Data represent mean ± SE of three independent experiments (n = 3). Protein levels (control = 100%) of CD36 and MSR1 (C) were measured by western blotting. *P < 0.05, #P < 0.01 compared with control. | |
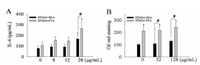
The interference assay suggests that the interactions between c-MWCNTs and the analyzed cytokines were only minimal (Fig. S4 in Supporting information). BSA-coated or FFA-coated c-MWCNTs dose-dependently increased IL-6 concentration (P < 0.05), with a significantly increased IL-6 release after exposure to 128 μg/mL cMWCNTs that were pre-incubated with BSA (P < 0.05) or FFA (P < 0.01, Fig. 4A). However, FFA-coated MWCNTs induced a significantly higher IL-6 release than those pre-incubated with BSA (P < 0.05). Exposure to 32 μg/mL and 128 μg/mL of BSA-coated or FFA-coated c-MWCNTs significantly reduced sMCP-1 release (P < 0.05, Fig. S8A in Supporting information). However, it should be noted that the decrease in sMCP-1 was only modest and was not dose-dependent. Exposure to BSA-coated or FFA-coated MWCNTs did not significantly affect IL-8 release (P > 0.05, Fig. S8B in Supporting information). The results from oil red staining showed a significant increase of lipid accumulation after exposure to 128 μg/mL BSA-coated c-MWCNTs (P < 0.05, Fig. 4B). FFA exposure also significantly enhanced lipid accumulation (P < 0.01), and 32 μg/mL and 128 μg/mL of MWCNTs that were pre-incubated with FFA led to significantly higher lipid accumulation than BSA-coated MWCNTs (P < 0.01).

|
Download:
|
| Fig. 4. The formation of macrophage foam cells. THP-1 macrophages were exposed to c-MWCNTs (code XFM24) pre-incubated with BSA or FFA for 24 h. (A) IL-6 release was determined by ELISA. (B) Lipid accumulation was quantified by oil red staining (control = 100%). Data represent mean ± SE of 4 independent experiments (n = 3). *P < 0.05, #P < 0.01 compared with control. | |
ER stress and autophagy are involved in lipid droplet biogenesis and foam cell formation. For ER stress, it is well-accepted that prolonged ER stress can promote lipid droplet formation [22, 24]. For autophagy, it selectively breaks down lipid droplet-stored lipids, a process known as lipophagy. Dysfunction of this process has been suggested to promote macrophage foam cell formation and is closely related with the development of atherosclerosis [23]. Here, the results showed that both BSA-coated c-MWCNTs and those pre-incubated with FFA significantly increased IL-6 release and lipid accumulation (Fig. 4), with FFA pre-incubated ones being most effective. BSA-coated or FFA-coated c-MWCNTs significantly increased the release of IL-6, but not sMCP-1 or IL-8, which may indicate that they only modestly induced inflammatory response in THP-1 macrophages. The modest increase in lipid accumulation and IL-6 release may be due to the induction of expression of the ER stress marker, DDIT3, since it is a transcription factor that regulates lipogenesis [22] and inflammatory responses [30]. In cells exposed to c-MWCNTs that were pre-incubated with FFA, the mRNA and protein levels of the autophagic markers, ATG5 and BECN1, were down-regulated (Fig. 2). ATG5 and BECN1 are crucial regulators of lipophagy [24, 31]. Our recent study showed that lipophagy may be involved in MWCNT-induced lipid accumulation in human hepatocytes, since inhibition of autophagy significantly augmented MWCNT-induced lipid accumulation [11]. Thus, it is possible that defective induction of autophagy by exposure to FFA-coated MWCNTs may impair the metabolism of lipid droplets and consequently, contribute to lipid accumulation in THP-1 macrophages. Another explanation for the increased lipid accumulation is the up-regulation of mRNA and protein levels of scavenger receptors (CD36 and MSR1) due to FFA-coated c-MWCNT exposure (Fig. 3). Similar observations have been previously reported, with NP exposure up-regulating scavenger receptors, leading to macrophage foam cell formation in vitro [32, 33].
It has been suggested that the interactions between nutrients and NPs should be evaluated to reflect the toxicity of NPs in the biological environment, but previous studies in this area have focused primarily on the effect of nutrients on colloidal aspects of NPs [34, 35]. The influence of nutrients on NP-induced signaling pathways has not been studied in detail. As a pilot study, we used FFA, which are known to influence ER stress/autophagy and lipid accumulation [15, 16]. Recently we found that 3-hydroxyflavone promoted the cytotoxicity of ZnO NPs, probably because 3- hydroxyflavone modulated NP-induced ER stress [36]. In nanomedicinal studies, modulation of ER stress/autophagy has been shown to influence the therapeutic effects of NPs [12, 37, 38]. Moreover, researchers have developed NPs based on nutrients, such as albumin [39] and lipids [40, 41]. The data from this study may indicate a need to evaluate the impact of excessive nutrients when assessing the effects of NPs, especially the nutrients that are known to modulate ER stress/autophagy.
Collectively, the results from this study showed that preincubation with BSA or FFA can coat the surface of c-MWCNTs and alter c-MWCNT-induced expression of ER stress/autophagy genes, but not cytotoxicity or oxidative stress in THP-1 macrophages. Specifically, we propose that FFA-coated MWCNTs exert their effects via the mechanism summarized in Fig. S9 (Supporting information). After exposure to FFA-coated MWCNTs, lipids are stored in lipid droplets, with the help of the ER. Although FFAcoated MWCNTs down-regulated ER stress, which may limit lipid droplet biogenesis, autophagy (metabolism of lipid droplets by autophagy is termed lipophagy) was also down-regulated, which can impair lipid droplet metabolism, leading to lipid accumulation. Additionally, MWCTs pre-incubated with FFA also up-regulated scavenger receptors (CD36 and MSR1), which may have contributed to increased cholesterol influx and foam cell formation. Currently, different kinds of NPs, such as functional magnetic NPs [42] and polymers [43-45], have been developed for disease therapy. The results from this study may indicate that it is necessary to assess the impact of biological molecules on the endpoints of NPs intended for biomedical uses.
AcknowledgmentsThis work was financially supported by the National Natural Science Foundation of China (No. 21707114) and Scientific Research Foundation of Hunan Provincial Education Department (No. 17A205).
| [1] |
M.E. Vance, T. Kuiken, E.P. Vejerano, et al., Beilstein J. Nanotechnol. 6 (2015) 1769-1780. DOI:10.3762/bjnano.6.181 |
| [2] |
T. Kuang, L. Chang, F. Chen, et al., Carbon 105 (2016) 305-313. DOI:10.1016/j.carbon.2016.04.052 |
| [3] |
M.F. de Volder, S.H. Tawfick, R.H. Baughman, A.J. Hart, Science 339 (2013) 535-539. DOI:10.1126/science.1222453 |
| [4] |
H. Gong, R. Peng, Z. Liu, Adv. Drug Deliv. Rev. 65 (2013) 1951-1963. DOI:10.1016/j.addr.2013.10.002 |
| [5] |
B.S. Wong, S.L. Yoong, A. Jagusiak, Adv. Drug Deliv. Rev. 65 (2013) 1964-2015. DOI:10.1016/j.addr.2013.08.005 |
| [6] |
P. Moller, D.V. Christophersen, D.M. Jensen, et al., Arch. Toxicol. 88 (2014) 1939-1964. DOI:10.1007/s00204-014-1356-x |
| [7] |
P. Moller, N.R. Jacobsen, Crit. Rev. Toxicol. 47 (2017) 867-884. |
| [8] |
L. Ou, B. Song, H. Liang, et al., Part. Fibre Toxicol. 13 (2016) 57. DOI:10.1186/s12989-016-0168-y |
| [9] |
J. Long, Y. Xiao, L. Liu, Y. Cao, J. Nanobiotechnol. 15 (2017) 80. DOI:10.1186/s12951-017-0318-x |
| [10] |
S. Chang, X. Zhao, S. Li, et al., Ecotoxicol. Environ. Saf. 161 (2018) 569-577. DOI:10.1016/j.ecoenv.2018.06.025 |
| [11] |
C. Zhao, Y. Zhou, L. Liu, et al., Food Chem. Toxicol. 121 (2018) 65-71. DOI:10.1016/j.fct.2018.08.033 |
| [12] |
Y. Cao, J. Long, L. Liu, et al., Life Sci. 186 (2017) 33-42. DOI:10.1016/j.lfs.2017.08.003 |
| [13] |
Y. Cao, Y. Gong, L. Liu, et al., J. Appl. Toxicol. 37 (2017) 1359-1369. DOI:10.1002/jat.v37.12 |
| [14] |
H.O. Rashid, R.K. Yadav, H.R. Kim, H.J. Chae, Autophagy 11 (2015) 1956-1977. DOI:10.1080/15548627.2015.1091141 |
| [15] |
G.S. Hotamisligil, E. Erbay, Nat. Rev. Immunol. 8 (2008) 923-934. DOI:10.1038/nri2449 |
| [16] |
G.S. Hotamisligil, Nature 542 (2017) 177-185. DOI:10.1038/nature21363 |
| [17] |
J. Saleem, L. Wang, C. Chen, Adv. Healthc. Mater. 7 (2018) e1800525. DOI:10.1002/adhm.v7.20 |
| [18] |
X. Fang, L. Jiang, Y. Gong, et al., Chem. Biol. Interact. 278 (2017) 40-47. DOI:10.1016/j.cbi.2017.10.002 |
| [19] |
Q. Jiang, X. Li, S. Cheng, et al., Environ. Toxicol. Pharmacol. 48 (2016) 103-109. DOI:10.1016/j.etap.2016.10.014 |
| [20] |
M.R. Go, S.H. Bae, H.J. Kim, J. Yu, S.J. Choi, Front. Microbiol. 8 (2017) 1013. DOI:10.3389/fmicb.2017.01013 |
| [21] |
Y. Gong, X. Li, G. Liao, et al., RSC Adv. 8 (2018) 15380-15388. DOI:10.1039/C8RA02509F |
| [22] |
A. Baiceanu, P. Mesdom, M. Lagouge, F. Foufelle, Nat. Rev. Endocrinol. 12 (2016) 710-722. DOI:10.1038/nrendo.2016.124 |
| [23] |
F. Cingolani, M.J. Czaja, Trends Endocrinol. Metab. 27 (2016) 696-705. DOI:10.1016/j.tem.2016.06.003 |
| [24] |
T.C. Walther, J. Chung, R.V. Farese Jr., Annu. Rev. Cell Dev. Biol. 33 (2017) 491-510. DOI:10.1146/annurev-cellbio-100616-060608 |
| [25] |
Q. Dai, J. Guo, Y. Yan, et al., Biomacromolecules 18 (2017) 431-439. DOI:10.1021/acs.biomac.6b01545 |
| [26] |
G. Duan, S.G. Kang, X. Tian, et al., Nanoscale 7 (2015) 15214-15224. DOI:10.1039/C5NR01839K |
| [27] |
W. Hu, C. Peng, M. Lv, et al., ACS Nano 5 (2011) 3693-3700. DOI:10.1021/nn200021j |
| [28] |
X. Zhao, D. Lu, F. Hao, R. Liu, J. Hazard. Mater. 292 (2015) 98-107. DOI:10.1016/j.jhazmat.2015.03.023 |
| [29] |
Y. Gong, L. Liu, J. Li, Y. Cao, J. Nanopart. Res. 19 (2017) 335. DOI:10.1007/s11051-017-4038-9 |
| [30] |
A.D. Garg, A. Kaczmarek, O. Krysko, et al., Trends Mol. Med. 18 (2012) 589-598. DOI:10.1016/j.molmed.2012.06.010 |
| [31] |
M. Ouimet, V. Franklin, E. Mak, et al., Cell Metab. 13 (2011) 655-667. DOI:10.1016/j.cmet.2011.03.023 |
| [32] |
Y. Suzuki, S. Tada-Oikawa, G. Ichihara, et al., Toxicol. Appl. Pharmacol. 278 (2014) 16-25. DOI:10.1016/j.taap.2014.04.010 |
| [33] |
C. Guo, R. Ma, X. Liu, et al., Sci. Total Environ. (2018) 631-632 570-579. |
| [34] |
D.J. McClements, H. Xiao, P. Demokritou, Adv. Colloid Interface Sci. 246 (2017) 165-180. DOI:10.1016/j.cis.2017.05.010 |
| [35] |
Y. Cao, J. Li, F. Liu, et al., Environ. Toxicol. Pharmacol. 46 (2016) 206-210. DOI:10.1016/j.etap.2016.07.023 |
| [36] |
Y. Luo, C. Wu, L. Liu, et al., Appl. Toxicol. 38 (2018) 1206-1214. DOI:10.1002/jat.v38.9 |
| [37] |
J. Xu, H. Wang, Y. Hu, et al., Adv. Sci. 6 (2019) 1801233. DOI:10.1002/advs.v6.8 |
| [38] |
Y. Zhang, R. Sha, L. Zhang, et al., Nat. Commun. 9 (2019) 4236. |
| [39] |
H. Wang, J. Wu, L. Xu, et al., Chem. Commun. (Camb.) 53 (2017) 2618-2621. DOI:10.1039/C6CC08978J |
| [40] |
H. Wang, Z. Lu, L. Wang, et al., Cancer Res. 77 (2017) 6963-6974. DOI:10.1158/0008-5472.CAN-17-0984 |
| [41] |
H. Wang, J. Chen, C. Xu, et al., Theranostics 7 (2017) 3638-3652. DOI:10.7150/thno.20028 |
| [42] |
Z. Chen, C. Wu, Z. Zhang, et al., Chin. Chem. Lett. 29 (2018) 1601-1608. DOI:10.1016/j.cclet.2018.08.007 |
| [43] |
Y. Qiao, J. Wan, L. Zhou, et al., Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 11 (2019) e1527. DOI:10.1002/wnan.1527 |
| [44] |
S. Gao, G. Tang, D. Hua, et al., J. Mater. Chem. B 7 (2019) 709-729. DOI:10.1039/C8TB02491J |
| [45] |
H. Wang, L. Zhou, K. Xie, et al., Theranostics 8 (2018) 3949-3963. DOI:10.7150/thno.26161 |
 2019, Vol. 30
2019, Vol. 30 

