b University of Chinese Academy of Sciences, Beijing 100049, China
Aflatoxin, one of the most toxic mycotoxins, is produced in Aspergillus flavus, Aspergillus parasiticus and Aspergillus metabolic processes [1-3]. Aflatoxin has harmful effect on human and animal health [1-3]. Aflatoxin B1 (AFB1) is the most commonly and toxic component in aflatoxins family, and has been classified as Group 1 carcinogen by the International Agency for Research in Cancer [1-3]. Agriculture products (e.g., corn, nuts, wheat) and food stuff are easy to be contaminated by AFB1. Countries worldwide have set strict regulation limits of AFB1 in various products [3, 4]. Therefore, development of sensitive and reliable methods for monitoring AFB1 is of great importance. A simple and rapid method is highly demanded for AFB1 detection on sites.
Traditional methods for detection of AFB1 include thin layer chromatography (TLC), high performance liquid chromatography (HPLC), and liquid chromatography–tandem mass spectrometry (LC–MS) [4-6]. However, these methods are complicated, costly, time-consuming, and require sophisticated instruments and trained personnel [4, 6]. Immunoassays can simplify processing procedures and are relatively fast [7], but the antibodies meet limitations in preparation, stability, and cost.
Aptamers, short single stranded DNA or RNA, can bind to targets (e.g., small molecules or proteins) with high affinity and selectivity. Aptamers are selected from a random sequence oligonucleotide library through an in vitro process named systematic evolution of ligands by exponential enrichment (SELEX) [8, 9]. In comparison to antibodies, aptamers possess remarkable merits, such as ease of generation, low cost, good thermal stability, easy chemical synthesis and modification with various functional groups, thus they are promising in broad analytical applications [10-15].
Aptamers against AFB1 were successfully selected [16, 17], and aptamer-based assays have been developed for AFB1, including colorimetric assay, electrochemical assay, fluorescence assay, chemiluminescence assay, dipstick assay, and etc. [17-22]. Wang et al. described a method based on aptamer induced assembly of fluorescent nitrogen-doped carbon dots (N, C-dots) on gold nanoparticles [20]. The fluorescence of N, C-dots was quenched by aptamer modified gold nanoparticles. Upon addition of AFB1, aptamers on gold nanoparticles reacted with AFB1 and released N, C-dots, causing fluorescence recovery of N, C-dots [20]. Lu et al. employed grapheneoxide (GO) as aquencherand aptamermodified quantum dots (QDs) to develop an assay for AFB1 [21]. The binding of AFB1 to aptamer caused QD-labelled aptamer was displaced from graphene oxide, and the fluorescence increased [21].
Molecular beacon techniques are attractive for biosensing and bioassays for nucleic acids, showing advantages in sensitivity, simplicity, and rapidity [23-26]. The use of aptamer probes has allowed to develop aptamer based fluorescent molecular beacon assays for detection of proteins and small molecules, e.g., Tat protein ofHIV-1, thrombin, cocaine, adenosine triphosphate (ATP), basedon binding-induced structure change and the subsequent fluorescence decrease or fluorescence increase [25, 27-33].
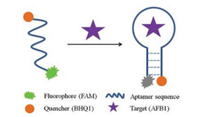
Taking advantage of aptamer molecular beacon, herewe reported a simple aptamer molecular beacon assay for rapid AFB1 detection.Fig. 1 shows the principle of our aptamer molecular beacon assay for AFB1. Fluorescein (FAM) and black hole quencher1 (BHQ1) are conjugatedat the 5′ end and the 3′ end of one aptamer against AFB1, respectively. In the absence of AFB1, FAM andBHQ1in theaptamerareapart fromeach other. In the presence of AFB1, aptamer binds with AFB1, forming a hairpin structure and drawing FAM and BHQ1 into close proximity, and the fluorescence of FAM is quenched by the adjacent BHQ1. Detection of AFB1 is achieved by measuring the fluorescence change of the aptamer probe.
|
Download:
|
| Fig. 1. Schematic diagram of fluorescence assay for AFB1 by using an aptamer beacon probe. In the absence of AFB1, FAM and BHQ1 are far from each other. In the presence of the AFB1, the target binding induces the aptamer to form a hairpin structure and brings FAM and BHQ1 into proximity, causing fluorescence quenching. AFB1 can be detected by measuring the fluorescence intensity change of probe. | |
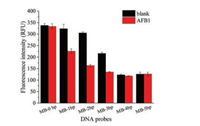
One DNA aptamer against AFB1 (5′-TGGGCACGTGTTGTCTCT CTGTGTCTCGTGCCCT-3′) was predicted to have a stem-loop secondary structure [16] (Fig. S1A in Supporting information). To develop a successful aptamer molecular beacon assay for AFB1, we truncated a series of base pairs off the terminals of the stem of this aptamer and designed the molecular beacon probes having different numbers of base pairs in the stem part (shown in Table S1 and Fig. S1B in Supporting information). The numbers of base pairs in the stem of aptamer probe are critical for the aptamer beacon to detect AFB1. Fluorescence intensities of the FAM-labeled aptamer probes (excitation wavelength: 485 nm; the emission wavelength: 528 nm) were recorded in the absence or in the presence of AFB1. MB-0bp gave high fluorescence intensity, but it showed negligible fluorescence change in the presence of AFB1. Upon addition of 200 nmol/L AFB1, MB-1bp, MB-2bp and MB-3bp probes showed fluorescence quenching, and the MB-2bp allowed for larger fluorescence change than MB-1bp and MB-3bp (Fig. 2). For MB- 4bp and MB-5bp, they showed lower fluorescence in the absence of AFB1 and did not show remarkable fluorescence change in the presence of AFB1. It suggests that MB-4bp and MB-5bp possibly have stable stem structures, and FAM and BHQ1 of them are already adjacent in the absence of AFB1. MB-0bp might lose binding affinity to AFB1 because the stem structure was completely disrupted. The probe MB-2bp (5′-FAM-CGTGTTGTCTC TCTGTGTCTCG-BHQ1-3′, the underline nucleotides were the complementary nucleotides in the stem) was preferred due to the larger AFB1-binding induced fluorescence change and relatively good binding affinity.
|
Download:
|
| Fig. 2. Fluorescence responses of aptamer probes (25 nmol/L) having different numbers of base pairs in the stem in the absence of or in the presence of AFB1 (200 nmol/L). The binding buffer contained 10 mmol/L HEPES (pH 7.0), 20 mmol/L MgCl2, and 0.1% Tween 20. | |
We investigated the effect of MgCl2 concentration in the binding buffer on the fluorescence response of MB-2bp to addition of AFB1. The tested binding buffer contained 10 mmol/L HEPES (pH 7.0), 0.1% Tween 20, and various concentrations of MgCl2. As shown in Fig. S2 (Supporting information), with increase of MgCl2 the fluorescence intensity of aptamer probe in the absence of AFB1 decreased, and then the fluorescence was not changed at higher concentrations of MgCl2 (higher than 50 mmol/L). Fluorescence change caused by AFB1 binding (fluorescence intensity of the blank sample (Fblank) was subtracted from fluorescence intensity of the sample containing 200 nmol/L AFB1 (FAFB1), denoted as △F (△F = Fblank-FAFB1)) gradually increased with the increase of MgCl2, and then it reached the maximum value when the concentration of MgCl2 was 20 mmol/L. Further increase of MgCl2 concentration had slight effect on △F. It suggests the use of MgCl2 in the binding buffer helps to improve the fluorescence change of the aptamer probe caused by AFB1. We further investigated the influence of NaCl in the binding buffer on the fluorescence response of MB-2bp upon addition of AFB1. Fig. S3 (Supporting information) shows how the fluorescence intensity of MB-2bp changed with varying concentrations of NaCl in the binding buffer without containing MgCl2 in the absence of or in the presence of AFB1. In both cases, the fluorescence intensity of the MB-2bp increased firstly and then decreased with further increase of NaCl. The fluorescence intensity of MB-2bp reached maximum value at 20 mmol/L NaCl. In the presence of AFB1, when NaCl concentrations were higher than 120 mmol/L in the binding buffer, the addition of AFB1 caused fluorescence decrease, and the fluorescence change caused by AFB1 increased with increase of NaCl. It shows that MgCl2 and NaCl have different influences on the fluorescence responses of MB-2bp.
When the binding buffer containing MgCl2(20 mmol/L)wasused, with increase of NaCl, the fluorescence intensity of MB-2bp in the absence of AFB1 gradually increased and then reached a plateau value at about 20mmol/L NaCl (Fig. S4 in Supporting information). In the presence of AFB1, the AFB1-binding induced net fluorescence change did not alter much with increase of NaCl. When NaCl concentration was higher than 20 mmol/L, the fluorescence change caused by AFB1 was relatively higher. The use of high concentration of NaCl may reduce the possible nonspecific interactions between aptamer and interferences. Therefore, basedon the above results, the binding buffer containing 20mmol/L MgCl2 and 120mmol/L NaCl was used in further studies.
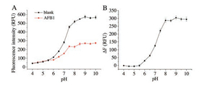
The solution pH may affect the binding between aptamer and AFB1 and the fluorescence intensity of FAM. We tested fluorescence response of the MB-2bp upon AFB1 in the buffers with pH values ranging from 4.0 to 10.0 (shown in Fig. 3). As Fig. 3A shows, the fluorescence intensity of MB-2bp increased with increase of pH, and reached the maximum level when pH was at 9.0. The fluorescence intensitychange △F caused by AFB1 binding was also greatly enhanced with pH increase in the range from 5.5 to 8.0. When the pH value increased from 8.0 to 10.0, the △ did not further change. The binding buffer with pH 8.0 was applied in our assay.
|
Download:
|
| Fig. 3. (A) Effect of buffer pH on fluorescence response of MB-2bp before and after addition of AFB1 (200 nmol/L). (B) Effect of buffer pH on the fluorescence change (△F) caused by AFB1. Concentration of MB-2bp was 25 nmol/L. Buffers containing 10 mmol/L HEPES, 20 mmol/L MgCl2, 120 mmol/L NaCl, and 0.1% Tween 20 were used. | |
We also tested the influence of incubation time in the optimized binding buffer (10 mmol/L HEPES (pH 8.0), 20 mmol/L MgCl2, 120 mmol/L NaCl, and 0.1% Tween 20) (Fig. S5 in Supporting information). It indicated the aptamer probe showed quick response to AFB1. Longer time incubation caused small decrease of fluorescence change (△F). Finally, 15 min incubation at room temperature was chosen because the maximum AFB1 binding induced fluorescence change was achieved at this condition.
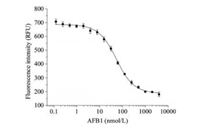
Detection of AFB1 using MB-2bp was achieved under the optimized conditions and the detailed procedure was described in Supporting information. As Fig. 4 shows, fluorescence intensity of MB-2bp gradually decreased with increase of AFB1. When 25 nmol/L MB-2bp was applied, the detection limit of AFB1 was determined to be about 3.9 nmol/L. The dynamic detection range was from 3.9 nmol/L to 4 μmol/L. The fluorescence of MB-2bp decreased more than 70% when AFB1 was at 4 μmol/L. The dissociation constant (Kd) of the aptamer probe MB-2bp was estimated to be 50 ± 2 nmol/L by non-linear fitting from the fluorescence response curve of MB-2bp, which is higher than that reported value (12 nmol/L) for the original aptamer [16]. It suggests that truncation of stem decreased the affinity the aptamer because the stability of stem became weak. The use of higher concentrations of aptamer probe allowed high fluorescence intensity of probe and larger net fluorescence decrease (Fig. S6 in Supporting information), but the target-binding induced percentage changes of fluorescence intensity were close. Under the same experimental conditions, the aptamer probes MB-1bp and MB-3bp also gave fluorescence response to AFB1 (Fig. S7 in Supporting information), and the detection limits of AFB1 were 15.6 nmol/L (MB-1bp) and 3.9 nmol/L (MB-3bp), respectively. The Kds of MB-1bp and MB-3bp were estimated to be 136 ± 5 nmol/L and 15 ± 2 nmol/L respectively. It shows MB-2bp showed better assay performance for detection of AFB1. The sensitivity of this fluorescence assay is comparable to that of a few reported aptamer-based assays for AFB1 (Table S2 in Supporting information) [21, 34-38], and it is lower than that of the assays using signal amplifications [39, 40]. As comparison, our assay is simple and rapid (Table S2). The electrochemical assays needed long time for preparation of modified electrode and detection of targets [18, 19].
|
Download:
|
| Fig. 4. Detection of AFB1 using aptamer probe MB-2bp. MB-2bp (25 nmol/L) was incubated with varying concentrations of AFB1 in the binding buffer (10 mmol/L HEPES (pH 8.0), 20 mmol/L MgCl2, 120 mmol/L NaCl, and 0.1% Tween 20). | |
To test the selectivity of the aptamer molecular beacon assay, we assessed the other five kinds of mycotoxins (OTA, OTB, FB1, FB2 and ZAE) (Fig. S8 in Supporting information). AFB1 (200 nmol/L) was tested along with OTB (2 μmol/L) and other mycotoxins concentrations (4 μmol/L). The addition of AFB1 caused great decrease of fluorescence intensity of MB-2bp. The fluorescence intensity of MB-2bp did not significantly change when other mycotoxins were tested. It suggests that the tested other mycotoxins did not cause interference to AFB1 detection. In addition, the presence of OTA, OTB, FB1, FB2 or ZAE in the AFB1 samples did not interfere with the detection of AFB1. The results show the assay is selective for AFB1 detection.
To assess the performance of this assay in complex sample matrix, we detected AFB1 spiked in diluted liquor wine with the binding buffer. As shown in Fig. S9 (Supporting information), detection of AFB1 spiked in 10-fold diluted liquor wine was achieved, and the sensitivity was a little lower than that in binding buffer. The detection limit of AFB1 in the 10-fold diluted liquor was 7.8 nmol/L, and dynamic range was from 7.8 nmol/L to 4 μmol/L. We also applied our method to detect AFB1 in corn flour samples. AFB1 at different levels in corn flour sample was successfully detected with recovery ranging from 81% to 133% (Table S3 in Supporting information). We compared the results using the aptamer beacon method and that determined from the commercial ELISA kit for AFB1 (Table S3). The determined level of AFB1 was close to that obtained from the ELISA kit, showing our assay is comparable to the ELISA kit. These results show this assay has potential for analysis of AFB1 in complex sample matrix.
In summary, we successfully developed a simple aptamer molecular beacon assay for AFB1. Fluorescein and BHQ1 quencher were labelled on the 5′ end and the 3′ end of an aptamer against AFB1, respectively. The presence of AFB1 caused fluorescence quenching of aptamer probe as the target binding induced formation of a hairpin structure and brought the fluorescein and quencher into proximity. Detection of AFB1 was achieved by measuring the change of fluorescence intensity. This assay was specific for AFB1 and showed potential for AFB1 detection in real samples. This aptamer molecular beacon assay shows advantages in simplicity and rapidity.
AcknowledgmentsWe thank for financial support from the National Natural Science Foundation of China (Nos. 21575153, 21435008, 21874146), Strategic Priority Research Program of the Chinese Academy of Sciences (No. XDB14030200), and the Key Research Program of the Chinese Academy of Sciences (No. KFZD-SW-203).
Appendix A. Supplementary dataSupplementary material related to this article canbefound, in the online version, at doi:https://doi.org/10.1016/j.cclet.2019.01.029.
| [1] |
P. Kumar, D.K. Mahato, M. Kamle, T.K. Mohanta, S.G. Kang, Front. Microbiol 7 (2016) 2170.. |
| [2] |
P.E. Jackson, J.D. Groopman, Best Pract. Res. Clin. Gastroenterol. 13 (1999) 545-555. |
| [3] |
U.P. Sarma, P.J. Bhetaria, P. Devi, A. Varma, Indian J. Clin. Biochem. 32 (2017) 124-133. DOI:10.1007/s12291-017-0649-2 |
| [4] |
H. Yao, Z. Hruska, Di Mavungu J.D., World Mycotoxin J. 8 (2015) 181-191. DOI:10.3920/WMJ2014.1797 |
| [5] |
S.M. Herzallah, Food Chem. 114 (2009) 1141-1146. DOI:10.1016/j.foodchem.2008.10.077 |
| [6] |
H.K. Abbas, W.T. Shier, B.W. Horn, M.A. Weaver, J. Toxicol.-Toxin Rev. 23 (2008) 295-315. |
| [7] |
P. Li, Q. Zhang, W. Zhang, TrAC-Trend. Anal. Chem. 28 (2009) 1115-1126. DOI:10.1016/j.trac.2009.07.003 |
| [8] |
A.D. Ellington, J.W. Szostak, Nature 346 (1990) 818-822. DOI:10.1038/346818a0 |
| [9] |
C. Tuerk, L. Gold, Science 249 (1990) 505-510. DOI:10.1126/science.2200121 |
| [10] |
J.W. Liu, Z.H. Cao, Y. Lu, Chem. Rev. 109 (2009) 1948-1998. DOI:10.1021/cr030183i |
| [11] |
F. Li, H. Zhang, Z. Wang, et al., Anal. Chem. 87 (2015) 274-292. DOI:10.1021/ac5037236 |
| [12] |
W. Zhou, P.J. Huang, J. Ding, J. Liu, Analyst 139 (2014) 2627-2640. DOI:10.1039/c4an00132j |
| [13] |
A.B. Iliuk, L. Hu, W.A. Tao, Anal. Chem. 83 (2011) 4440-4452. DOI:10.1021/ac201057w |
| [14] |
H.Q. Chen, J. Xu, F. Yuan, et al., Chin. Chem. Lett. 24 (2013) 79-81. DOI:10.1016/j.cclet.2012.12.008 |
| [15] |
X. Nie, X. Ning, Y.Y. Zhao, et al., Chin. Chem. Lett. 28 (2017) 619-624. DOI:10.1016/j.cclet.2016.11.013 |
| [16] |
L.C. Le, J.A. Cruz-Aguado, G.A. Penner, Patent, PCT/CA2010/001292, 2012.
|
| [17] |
N.M. Danesh, H.B. Bostan, K. Abnous, et al., TrAC-Trend. Anal. Chem. 99 (2018) 117-128. |
| [18] |
N. Hao, Y. Zhang, H. Zhong, et al., Anal. Chem. 89 (2017) 10133-10136. DOI:10.1021/acs.analchem.7b03132 |
| [19] |
L. Wu, F. Ding, W. Yin, et al., Anal. Chem. 89 (2017) 7578-7585. DOI:10.1021/acs.analchem.7b01399 |
| [20] |
B. Wang, Y. Chen, Y. Wu, et al., Biosens. Bioelectron. 78 (2016) 23-30. DOI:10.1016/j.bios.2015.11.015 |
| [21] |
Z. Lu, X. Chen, Y. Wang, X. Zheng, C.M. Li, Microchim. Acta 182 (2014) 571-578. |
| [22] |
K.Y. Goud, A. Sharma, A. Hayat, et al., Anal. Biochem. 508 (2016) 19-24. DOI:10.1016/j.ab.2016.05.018 |
| [23] |
Y. Li, X. Zhou, D. Ye, Biochem. Biophys. Res. Commun. 373 (2008) 457-461. DOI:10.1016/j.bbrc.2008.05.038 |
| [24] |
K. Wang, Z. Tang, C.J. Yang, et al., Angew. Chem. Int. Ed. 48 (2009) 856-870. DOI:10.1002/anie.v48:5 |
| [25] |
J. Zheng, R. Yang, M. Shi, et al., Chem. Soc. Rev. 44 (2015) 3036-3055. DOI:10.1039/C5CS00020C |
| [26] |
S. Tyagi, F.R. Kramer, Nat. Biotechnol. 14 (1996) 303-308. DOI:10.1038/nbt0396-303 |
| [27] |
A.A. Goulko, F. Li, C. Le X., TrAC-Trend. Anal. Chem. 28 (2009) 878-892. DOI:10.1016/j.trac.2009.03.014 |
| [28] |
R. Yamamoto, T. Baba, P.K. Kumar, Genes Cells 5 (2000) 389-396. DOI:10.1046/j.1365-2443.2000.00331.x |
| [29] |
N. Hamaguchi, A. Ellington, M. Stanton, Anal. Biochem. 294 (2001) 126-131. DOI:10.1006/abio.2001.5169 |
| [30] |
J.J. Li, X. Fang, W. Tan, Biochem. Biophys. Res. Commun. 292 (2002) 31-40. DOI:10.1006/bbrc.2002.6581 |
| [31] |
M.N. Stojanovic, P. de Prada, D.W. Landry, J. Am. Chem. Soc. 123 (2001) 4928-4931. DOI:10.1021/ja0038171 |
| [32] |
Z. Tang, P. Mallikaratchy, R. Yang, et al., J. Am. Chem. Soc. 130 (2008) 11268-11269. DOI:10.1021/ja804119s |
| [33] |
N. Li, C.M. Ho, J. Am. Chem. Soc. 130 (2008) 2380-2381. DOI:10.1021/ja076787b |
| [34] |
F.S. Sabet, M. Hosseini, H. Khabbaz, M. Dadmehr, M.R. Ganjali, Food Chem. 220 (2017) 527-532. DOI:10.1016/j.foodchem.2016.10.004 |
| [35] |
J. Zhang, Z. Li, S. Zhao, Y. Lu, Analyst 141 (2016) 4029-4034. DOI:10.1039/C6AN00368K |
| [36] |
M. Hosseini, H. Khabbaz, M. Dadmehr, M.R. Ganjali, J. Mohamadnejad, Acta Chim. Slov. 62 (2015) 721-728. |
| [37] |
L. Chen, F. Wen, M. Li, et al., Food Chem. 215 (2017) 377-382. DOI:10.1016/j.foodchem.2016.07.148 |
| [38] |
W.B. Shim, M.J. Kim, H. Mun, M.G. Kim, Biosens. Bioelectron. 62 (2014) 288-294. DOI:10.1016/j.bios.2014.06.059 |
| [39] |
Y. Seok, J.Y. Byun, W.B. Shim, M.G. Kim, Anal. Chim. Acta 886 (2015) 182-187. DOI:10.1016/j.aca.2015.05.041 |
| [40] |
X. Guo, F. Wen, N. Zheng, et al., Biosens. Bioelectron. 56 (2014) 340-344. DOI:10.1016/j.bios.2014.01.045 |
 2019, Vol. 30
2019, Vol. 30 

