b Key Laboratory of Hunan Province for Water Environment and Agriculture Product Safety, Changsha 410083, China;
c College of Chemistry & Molecular Engineering, Zhengzhou University, Zhengzhou 450001, China
Biological thiols, cysteine (Cys), homocysteine (Hcy) and glutathione (GSH), play important physiological functions in organisms [1-3]. They can promote the actions of enzymes in organism, and regulate the normal redox state of cells, and are important signal molecules in physiological activity. As a result, abnormal levels of thiols can lead to a variety of diseases. For example, the lack of Cys can cause slow growth, liver damage, hair fading, skin diseases, lethargy, and edema [4]. Excessive levels of Hcy in plasma of cells are closely related to cardiovascular disease, Alzheimer's disease and osteoporosis [5, 6]. GSH, an anti-oxidant agent, can protect the body from oxidative stress and free radical damage which are believed to correlate with leukemia, cancer and AIDS [7-9]. Therefore, the sensitive and selective detection of biothiols in living samples is of great significance.
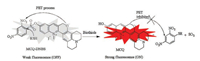
Compared with the traditional detection techniques, fluorescence analysis has the advantages of simple operation, high sensitivity, and non-invasive detection in biological samples. Thus, a lot of fluorescent probe have been developed to detect various ions and biomolecules in the past decade [10-22]. Though there are many fluorescent probes for biothiols reported, most of them may suffer from small Stokes shift, short emission wavelengths, high detection limit and/or poor water solubility. Recently, we developed a new kind of fluorophores, methylated chromenoquinolines (MCQs) having a red emission (up to 613 nm), a large Stokes shift (up to 115 nm) (. S1 in Supporting information), and a good water solubility [23], which are favorable for the construction of fluorescent probes. Based on a chromenoquinoline derivative, we designed a fluorescent probe, MCQ-DNBS, for biothiol detection using a common sensing group, 2, 4-dinitrobezensulfonate (DNBS) (Scheme 1) [24-27]. The fluorescent quantum yield (ΦFl) of MCQ in PBS buffers is of 0.07. The fluorescence of the MCQ-DNBS would be quenched due to the strong photo induced electron transfer (PET) process between MCQ and DNBS moiety. Upon the addition of biothiols to the solution of MCQ-DNBS, the nucleophilic biothiols would cleave DNBS moiety in this probe to release MCQ, which would emit a strong red fluorescence signal with a large Stokes shift upon excited (Scheme 2). It is noteworthy that the positive charge in this probe would provide an excellent water-solubility, which is desirable for biological applications. Therefore, probe MCQ-DNBS could serve as a turn-on fluorescence probe for biothiol detection with both a red emission and a large Stokes shift in aqueous media. The synthetic route of probe MCQ-DNBS was illustrated in Scheme 1.

|
Download:
|
| Scheme 1. Synthetic route of probe MCQ-DNBS. (a) CH2Cl2, Et3N, 0 ℃, 1 h, yield 21%. | |
|
Download:
|
| Scheme 2. The proposed sensing mechanism of MCQ-DNBS with biothiols. | |
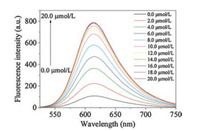
Due to the good water solubility of MCQ-DNBS, the fluorescence responses of probe MCQ-DNBS (10.0 μmol/L) to Cys, Hcy and GSH were studied in pure HEPES buffer (20.0 mmol/L, pH 7.4). The solution of MCQ-DNBS exhibited an intense absorption band centered at 517 nm, and was weakly fluorescent (ΦFl≈0.004). In the presence of Cys, the absorption spectrum of the solution of this probe slightly blue-shifted to 498 nm, and exhibited a strong red fluorescence with a maximum at 613 nm (Fig. 1 and Fig. S2 in Supporting information). The long emission wavelength of MCQDNBS was desirable for biological applications due to the deep tissue penetration, minimized photodamage to living organisms and the interference from background. The intensity of the fluorescent signal increased with increasing the added amount of Cys and leveled off when the concentration of Cys was up to 16.0 μmol/L (Fig. 1 and Fig. S3 in Supporting information). As seen in Fig. S3, there was a good linear relationship (R = 0.99865) between fluorescence intensity (613 nm) and the concentration of Cys in a range of 0.0–10.0 μmol/L. The detection limit for Cys was as low as 2.0 ×10-8 mol/L based on signal to noise (S/N = 3). Furthermore, this probe displayed a large Stokes shift (115 nm) in the sensing process, which could improve the sensing accuracy and avoid the interference from excitation signals. As for Hcy and GSH, similar results were obtained and the detection limits were 2.3 ×10-8 mol/L for Hcy and 2.1 ×10-8 mol/L for GSH (Figs. S4 and S5 in Supporting information), which were lower than or as low as the detection limits of previously reported fluorescent probes (Table S1 in Supporting information). These results clearly suggested that MCQ-DNBS could detect biothiols with high sensitivity in aqueous media.
|
Download:
|
| Fig. 1. Fluorescence spectral changes of probe MCQ-DNBS (10.0 μmol/L) with Cys (0.0–20.0 μmol/L) in HEPES buffer (20.0 mmol/L, pH 7.4) for 120 min. λex = 498 nm. Excitation/emission slit width: 5.0 nm/5.0 nm. | |
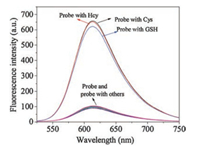
In order to evaluate the selectivity and anti-interference ability of probe MCQ-DNBS for the detection of biothiols, 20 kinds of amino acids were used as analytes to react with this probe, including Asp, Ala, Val, Phe, His, Leu, Ser, Ile, Trp, Lys, Arg, Pro, Gly, Met, Tyr, Glu, Thr, Cys, Hcy and GSH. As shown in Fig. 2, the addition of Cys, Hcy and GSH to the solution of this probe resulted in a significant fluorescence enhancement, while other amino acids caused negligible fluorescence change. Moreover, MCQ-DNBS still exhibited a good performance for the detection of biothiols under the co-existence of other amino acids (Figs. S7 and S8 in Supporting information). Therefore, it could be concluded that probe MCQ-DNBS had a good selectivity for Cys, Hcy and GSH.
|
Download:
|
| Fig. 2. Fluorescence response of probe MCQ-DNBS (10.0 μmol/L) to biothiols and other amino acids in HEPES buffer (20.0 mmol/L, pH 7.4) (Asp, Ala, Val, Phe, His, Leu, Ser, Ile, Trp, Lys, Arg, Pro, Gly, Met, Tyr, Glu and Thr, 0.1 mmol/L for each; 16.0 μmol/L for Cys, 10.0 μmol/L for Hcy, 10.0 μmol/L for GSH). | |
We also investigated the time-dependent fluorescence of probe MCQ-DNBS with Cys, Hcy and GSH, respectively. As shown in Fig. S9 (Supporting information), there was no fluorescence change when probe MCQ-DNBS was in the absence of biothiols within 120 min, indicating its good stability. In contrast, the addition of biothiols to the solution of probe MCQ-DNBS (10.0 μmol/L) quickly resulted in strong fluorescence signals, which were enhanced with increasing the reaction time and were maximized within 100 min for GSH (10.0 μmol/L), 120 min for Cys (16.0 μmol/L) and 120 min for Hcy (10.0 μmol/L) with rate constants of kobs[Cys] = 0.02269 min-1, kobs[Hcy] = 0.03088 min-1, kobs[GSH] = 0.02979 min-1 (Fig. S10 in Supporting information). Moreover, the response rate could be remarkably accelerated with the aid of CTAB, a surfactant (Fig. S11 in Supporting information).
The sensing mechanism of probe MCQ-DNBS to biothiol was investigated by NMR and mass spectral analysis of the isolated reaction product of MCQ-DNBS with Cys. As shown in Figs. S12 and S15 (Supporting information), 1H NMR spectra of the reaction product is consistent with that of MCQ. In addition, the high resolution mass spectrum of the reaction product displayed a peak at m/z 359.1765, which was nearly identical to the exact molecular weight of MCQ ([M-H]- = 359.1754) (Fig. S13 in Supporting information). These experimental results confirmed that Cys reacted with MCQ-DNBS to cleave the DNBS moiety from the probe and to release the fluorescent dye MCQ.
The effect of pH on the performance of this probe was also investigated. As shown in Fig. 3, probe MCQ-DNBS exhibited weak fluorescence at all pH values. In the presence of biothiols, a significant fluorescent enhancement could be observed within a pH range of 5–13, suggested MCQ-DNBS was able to detect biothiols under physiological condition.

|
Download:
|
| Fig. 3. pH Effect on the fluorescence intensity of probe MCQ-DNBS (10.0 μmol/L) in the absence/presence of biothiols. | |
Finally, the application of probe MCQ-DNBS was performed in the detection of biothiols in living cells. Cytotoxic assays of probe MCQ-DNBS in HeLa cells were performed, 91% of cell viability were obtained when cells were treated with 10.0 μmol/L of probe MCQDNBS for 24 h indicating probe MCQ-DNBS was non-toxic (Fig. S14 in Supporting information). When HeLa cells were incubated with probe MCQ-DNBS (10.0 μmol/L) for 30 min at 37 ℃, a strong red fluorescence signals gave off from inside cells (Fig. 4). In contrast, when cells were pre-treated with thiol-blocking agent (N-ethylmaleimide, NEM, 1.0 mmol/L) at 37 ℃ for 30 min and then incubated with probe MCQ-DNBS (10.0 μmol/L) at 37 ℃ for 30 min, only weak fluorescence signals were observed. These imaging experiments indicated that probe MCQ-DNBS was permeable to cell membrane and could detect intracellular biothiols in living cells.

|
Download:
|
| Fig. 4. Images of living HeLa Cells. Top row: cells incubated with probe MCQ-DNBS (10.0 μmol/L) for 30 min at 37 ℃. Bottom row: cells pretreated with N-ethylmaleimide (1.0 mmol/L) for 30 min and incubated with probe MCQ-DNBS (10.0 μmol/L) for 30 min at 37 ℃. Left column: bright-field; middle column: fluorescence; right column: merged. Cells were magnified 60 times. | |
In conclusion, we developed a fluorescent probe for the sensitive and selective probe detection of biothiols. This probe could work well in pure aqueous media due to its good water solubility. The red emission coupled with the large Stokes shift (115 nm) of this probe makes it suitable for the detection of biothiols in vitro and in vivo, which was demonstrated by the preliminary biological experiments
AcknowledgmentsThis work was supported by the National Natural Science Foundation of China (No. U1608222), Special Fund for Agroscientific Research in the Public Interest of China (No. 201503108), Fundamental Research Funds for the Central Universities of Central South University (No. 2018zzts109) and the State Key Laboratory of Chemo/Biosensing and Chemometrics (No. 2016005).
Appendix A. Supplementary dataSupplementary material related to this article can be found, in the online version, at doi:https://doi.org/10.1016/j. cclet.2018.12.017.
| [1] |
A. Fava, A. Iliceto, E. Camera, J. Am. Chem. Soc. 79 (1957) 833-838. DOI:10.1021/ja01561a014 |
| [2] |
Y.M. Go, D.P. Jones, Crit. Rev. Biochem. Mol. Biol. 48 (2013) 173-181. DOI:10.3109/10409238.2013.764840 |
| [3] |
N. Tyagi, K.C. Sedoris, M. Steed, et al., Am. J. Physiol. Heart Circ. Physiol. 289 (2005) 2649-2656. DOI:10.1152/ajpheart.00548.2005 |
| [4] |
S. Saeed, Anal. Chem. 73 (2001) 5972-5978. DOI:10.1021/ac010541m |
| [5] |
H. Refsum, P.M. Ueland, O. Nygård, S.E. Vollset, Annu. Rev. Med. 49 (1998) 31-62. DOI:10.1146/annurev.med.49.1.31 |
| [6] |
P. Sachdev, R. Parslow, C. Salonikas, et al., Arch. Neurol. 61 (2004) 1369-1376. DOI:10.1001/archneur.61.9.1369 |
| [7] |
W.A. Kleinman, J.P. Richie, Biochem. Pharmacol. 60 (2000) 19-29. DOI:10.1016/S0006-2952(00)00293-8 |
| [8] |
S.C. Lu, Mol. Asp. Med. 30 (2009) 42-59. DOI:10.1016/j.mam.2008.05.005 |
| [9] |
D.M. Townsend, K.D. Tew, H. Tapiero, Biomed. Pharmacother. 57 (2003) 145-155. DOI:10.1016/S0753-3322(03)00043-X |
| [10] |
M. Lan, J. Wu, W. Liu, et al., Sens. Actuators B-Chem. 156 (2011) 332-337. DOI:10.1016/j.snb.2011.04.042 |
| [11] |
X. Li, S. Qian, Q. He, et al., Org. Biomol. Chem. 8 (2010) 3627-3630. DOI:10.1039/c004344c |
| [12] |
F. Qi, X. Liu, L. Yang, et al., Tetrahedron 72 (2016) 6909-6913. DOI:10.1016/j.tet.2016.08.020 |
| [13] |
S.P. Wang, W.J. Deng, D. Sun, et al., Org. Biomol. Chem. 7 (2009) 4017-4020. DOI:10.1039/b909760k |
| [14] |
M. Wei, P. Yin, Y. Shen, et al., Chem. Commun. 49 (2013) 4640-4642. DOI:10.1039/c3cc39045d |
| [15] |
J. Shao, H. Guo, S. Ji, J. Zhao, Biosens. Bioelectron. 26 (2011) 3012-3017. DOI:10.1016/j.bios.2010.12.004 |
| [16] |
W. Jiang, Y. Cao, Y. Liu, W. Wang, Chem. Commun. 46 (2010) 1944-1946. DOI:10.1039/b926070f |
| [17] |
W. Qu, L. Yang, Y. Hang, et al., Sens. Actuators B-Chem. 211 (2015) 275-282. DOI:10.1016/j.snb.2015.01.117 |
| [18] |
M. Li, P. Cui, K. Li, et al., Chin. Chem. Lett. 29 (2018) 992-994. DOI:10.1016/j.cclet.2017.11.011 |
| [19] |
Y.H. Li, J.F. Yang, C.H. Liu, J.S. Li, R.H. Yang, Chin. Chem. Lett. 24 (2013) 96-98. DOI:10.1016/j.cclet.2013.01.037 |
| [20] |
L. Song, L.M. Ma, Q. Sun, et al., Chin. Chem. Lett. 27 (2016) 330-334. DOI:10.1016/j.cclet.2015.12.012 |
| [21] |
Y. Yang, H. Wang, Y.L. Wei, et al., Chin. Chem. Lett. 28 (2017) 2023-2026. DOI:10.1016/j.cclet.2017.08.051 |
| [22] |
P. Zhang, Z.Q. Guo, C.X. Yan, W.H. Zhu, Chin. Chem. Lett. 28 (2017) 1952-1956. DOI:10.1016/j.cclet.2017.08.038 |
| [23] |
X. Liu, Y. Li, X. Ren, et al., Chem. Commun. 54 (2018) 1509-1512. DOI:10.1039/C7CC08154E |
| [24] |
H. Maeda, H. Matsuno, M. Ushida, et al., Angew. Chem. Int. Ed. 44 (2005) 2922-2925. DOI:10.1002/(ISSN)1521-3773 |
| [25] |
X.D. Jiang, J. Zhang, X. Shao, W. Zhao, Org. Biomol. Chem. 10 (2012) 1966-1968. DOI:10.1039/c2ob07046d |
| [26] |
S. Chen, P. Hou, B. Zhou, et al., RSC Adv. 3 (2013) 11543-11546. DOI:10.1039/c3ra41554f |
| [27] |
X. Liu, L. Gao, L. Yang, et al., RSC Adv. 5 (2015) 18177-18182. DOI:10.1039/C5RA00255A |
 2019, Vol. 30
2019, Vol. 30 

