Monoamine oxidase B (MAO-B) belongs to the flavin monoamine oxidase family, which catalyzes the oxidation of biogenic and xenobiotic amines and plays an important role in the catabolism of neuroactive and vasoactive amines in the central nervous system and peripheral tissues. MAO dysfunction is thought to be responsible for several psychiatric and neurological disorders. Especially, the level of MAO-B is related to Alzheimer's disease and Parkinson's disease [1, 2]. Therefore, detecting MAO-B in human nerve cells is of great significance. Various technologies have been developed for the detection of MAO-B. Compared with technologies like positron emission tomography (PET) [3, 4], magnetic resonance imaging (MRI) [5, 6], and enzyme-linked immunosorbent assay (ELISA) that exploits bio-reducible agents, fluorescence imaging as a powerful non-invasive method with high sensitivity is regarded particularly suitable for monoamine oxidase detection [7-14].
In recent years, many fluorescent probes have been reported for MAO-B, most of which explored the fluorescence-enhancement strategy by removing the reducible group and releasing the fluorescence signal of the fluorophore based on MAO-B's redox reaction [10] (Scheme 1a). However, one shortage of this design strategy is that the original fluorescence intensity of the probes is hard to form a zero background before reacting with MAO, which would influence the sensitivity of the detection. Therefore, developing "off-on" fluorescent probe from a zero background is highly desirable. Probe via "covalent-assembly" strategy (raised by Yang group) could provide an approach to get "off-on" or ratiometric fluorescent probe, since a new fluorophore can be formed via covalent reaction of two fragments [15]. The fundamental feature of "covalent-assembly" probe is that it warrants both a colorimetric change and a turn-on fluorescent signal from a zero background. Moreover, the conjugative covalent-assembly of the backbone may cause a "red shift" of the fluorescence spectra, since a new fluorescent conjugated structure is developed.

|
Download:
|
| Scheme 1. (a) The reaction process of probes with MAO-B in most previous research. (b) Previously reported "covalent assembly" MAO-B probe. (c) The "covalent assembly" strategy applied in this work. | |
In fact, several "covalent-assembly" probes have been successfully applied in the detection of small molecules including sulfurcontaining species (SCS) and reactive oxygen species (ROS) [16-24]. However, such strategy is hardly applied in the detection of MAO [12, 13]. Sames group [13] reported a coumarin switch for MAO-B, based on a sequence of enzymatic oxidation and intramolecular cyclization (Scheme 1b). In this work, a different design strategy was applied in the "covalent-assembly" to detect MAO-B. Instead of expanding the conjugate planes of the fluorophore, we newly built a fluorescent skeleton. As indicated in Scheme 1c, after reacting with MAO-B, the re-exposed oxygen anion will attack the nearest carbonyl carbon atom, leading to the formation of the fluorophore and the off-on fluorescence effect.
As a kind of important fluorophore, coumarin has large conjugate system, strong molecular electron transfer ability, good thermodynamic and photochemical stability. As a result, a large number of coumarin based fluorescent probes have been reported for anions, metal ions and small molecules [25-28]. As an important reactant of coumarin in synthesis, salicylaldehyde compound has a hydroxyl moiety which is the key reaction group. If the hydroxyl group was protected, the formation of coumarin will be restricted because of the lack of nucleophilic group. Once the protecting group was removed, coumarin skeleton would be formed. In the light of "covalent-assembly" strategy, we want to develop a novel approach for MAO-B detection via this cascade reaction. In this work, two salicylaldehyde derivatives decorated with the 3-aminopropoxy group were designed for the detection of MAO-B. With the sequence of enzymatic oxidation by MAO-B and intramolecular cyclization, fluorescence signal of coumarin would be turned on. Furthermore, the development of inhibitors of MAOB is an important item in the treatment of neurodegenerative diseases [29, 30]. We therefore detected the inhibition of pargyline (N-benzyl-N-methylprop-2-yn-1-amine) to MAO-B using our probes. It was proved that both DEAN-MA and DEAB-MA were capable of screening MAO-B inhibitors. Importantly, it has been suggested that the expression level of MAO-B in nerve cells is higher than non-nerve cells [31], we therefore applied our probes to the imaging of MAO-B in both HL-7702 cells (human liver cells) and U87 cells (human astrocyte).
Firstly, the enzyme activity of MAO-B was tested to be 4.8 U/mg via UV-vis absorption spectra (Fig. S1 in Supporting information) [32]. Then the reaction conditions for probe DEAN-MA and DEABMA with MAO-B were studied, including the effects of pH and the concentration of probes. The enzymatic reaction was carried out in the pH range of 5.3 - 9.0. And It was discovered that the fluorescence barely changed under acidic conditions (pH ≤7.0), while a significant enhancement of the probe could be observed in biological conditions (pH around 7.4) after incubation with MAO-B for 1 h. This indicated that DEAN-MA and DEAB-MA should be able to serve as sensitive MAO-B probes under physiological conditions (Fig. S2 in Supporting information). Following that, the optimum concentration of the probes to react with MAO-B (10 μg/mL) were studied. As indicated in Fig. S3 (Supporting information), the fluorescence intensity reached a plateau when the concentration of the probes reached 10 μmol/L. The following experiments were therefore all carried out using 10 μmol/L as the concentrations of the probes.
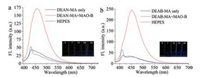
As seen from Fig. 1, fluorescence studies revealed that probe DEAB-MA and DEAN-AM had very weak fluorescence in HEPES buffer. Reaction of probe DEAN-MA and DEAB-AM with MAO-B leaded to the change of fluorescence signal at 456 nm from "off" state to "on" state. These results indicated that both fluorescence probes DEAN-MA and DEAB-MA were MAO-B reactive.
|
Download:
|
| Fig. 1. (a) The fluorescence response to MAO-B (10 μg/mL) and the image of fluorescence response of probe DEAN-MA (10 μmol/L) to different concentration of MAO-B. (b) The fluorescence response of probe DEAB-MA (10 μmol/L) to MAO-B (10 μg/mL) and the image of fluorescence response of probe DEAN-MA (10 μmol/L) to different concentration of MAO-B. Spectra collected 1 h after MAO-B addition. λex: 360 nm (DEAB-MA), 375nm (DEAN-MA). Solvent: HEPES buffer (100 mmol/L, pH 7.4, 1% DMSO and 5% glycerol included) [14]. | |
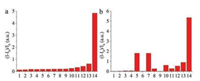
We then studied the specificity and selectivity of the probes towards MAO-B. We examined various potential interfering species in parallel under the same conditions, including H2O2, serine, glutamic acid, cysteine, glutathione, GSH, high-cysteine, some oxidoreductase and other related enzymes. As depicted in Fig. 2, probe DEAN-MA exhibited high selectivity to MAO-B over the species tested, which may be ascribed to the specific oxidation of the substrate by the enzyme. Probe DEAB-MA also showed high signal to background ratio after reacting with MAO-B, however, interferences from L-GSH and L-cysteine were also hard to omit. Therefore, only DEAN-MA was applied in the living cell imaging experiments. Furthermore, both DEAN-MA and DEAB-MA showed selectivity towards MAO-B to MAO-A under the same concentration, which indicates that probes are more suitable for detecting MAO-B than MAO-A.
|
Download:
|
| Fig. 2. The signal-to-background ratio of probe DEAN-MA (a) and DEAB-MA (b) (10 μmol/L) to several distractors. Spectra was collected 1h after substance addition. The concentration of enzymes except MAOs was 1mg/mL, and the concentration of h2O2 and amino acids was 1 mmol/L. Insert: 1. horse radish peroxidase, 2. glucoseoxidase, 3. H2O2, 4. sparsely cotton like thermophilic lipase, 5. L-GSH, 6. L-cysteine, 7. L-serine, 8. pancrelipase, 9. trypsin, 10. L-high cysteine, 11. D-glutamic acid, 12. acyl transferase, 13. MAO-A (10 μg/mL), 14. MAO-B (10 μg/mL). | |
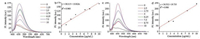
Fig. 3 shows the fluorescence change of probes with MAO-B at various concentrations. As is seen, the fluorescence intensity of the probes increased gradually with increasing concentration of MAOB. A good linearity between the fluorescence intensity and the concentration of MAO-B range of 0-10 μg/mL with an equation of I456 = 13.82209 × [MAO-B] +38.11 (R2 = 0.960) was observed. The detection limit (k =3) of DEAN-MA and DEAB-MA was respectively determined to be 0.6 ng/mL and 0.2 ng/mL [33], which was better or comparable to previously published MAO probes.

|
Download:
|
| Fig. 3. The fluorescence intensity of DEAN-MA (a) and DEAB-MA (10 μmol/L) (c) after reacting with MAO-B (the concentration of MAO-B range is 0 -10 μg/mL) for 1h and their linear graphs for DEAN-MA (b) and DEAB-MA (d). | |
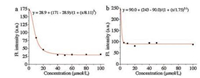
The probe can also be used for the screening of potential MAO-B inhibitors. As shown in Fig. 4, with increasing concentrations of pargyline (N-benzyl-N-methylprop-2-yn-1-amine), a typical inhibitor of MAO-B, the emission intensity gradually decreased. The IC50 value of pargyline obtained by using the logistic fit of the experimental data was 9.3 μmol/L for DEAN-MA and 2.6 μmol/L for DEAB-MA. This result suggests that DEAN-MA and DEAB-MA have the potential of being applied to the screening of MAO-B inhibitors.
|
Download:
|
| Fig. 4. The fluorescence intensity of DEAN-MA (a) and DEAB-MA (b) (10 μmol/L) reacting with MAO-B (10 μg/mL) for one more hour after deferent concentration of inhibitor incubated with MAO-B 1 h. | |
The reaction mechanism was proved by ESI-MS of the reacting product and the fluorescence spectra. The peak at m/z 255.0655 [M-Na+] of fluorescent molecule EMCC was found in the reaction mixturewith DEAB-MA and MAO-B, and the peaks at m/z 269.0802 [M-H+] and 291.0616 [M-Na+] of EBCC were found in the reaction mixture with DEAN-MA (Fig. S4 in Supporting information). The results of fluorescence spectra showed that the emission spectra of the theoretical product EMCC was the same tothe reaction mixture of DEAB-MA and MAO-B, and DEAN-MA showed the same result. These results proved the proposed reaction mechanism between the probes and MAO-B (Scheme 1c, Fig. S5 in Supporting information).
Followingthat, wefound the DEAN-MAcould response to MAO-B at λex =405nm as well (Fig. S6 in Supporting information), so the cytotoxicity of probe DEAN-MA to U87 and HL-7702 cells were studied (Fig. S7 in Supporting information). The results showed that the cells viability was still more than 80% when the concentration of DEAN-MA reached 10 μmol/L. Therefore, it was safe to say that DEAN-MA possessed relatively low cytotoxicity and the following fluorescence imaging experiments could be carried out with 10 μmol/L DEAN-MA.
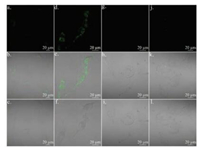
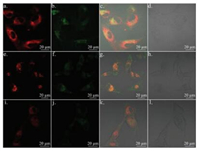
Fluorescence imaging of HL-7702 cells and U87 cells was shown in Fig. 5. The fluorescence in U87 cells was higher than that in HL- 7702 cells, through which we successfully demonstrated that the level of MAO-B in nerve cells is higher than somatic cells including liver epithelial cells. This result is in line with the previous report [28]. We then studied the distribution of the probe inside cells. As shown in Fig. 6, there was no obvious colocation effect between probe DEAN-AM and mitochondria tracker red, lysosome tracker red, or endoplasmic reticulum tracker red in U87 cells. The result suggested that probe DEAN-AM was distributed in various organelles after diffusion into cells.
|
Download:
|
| Fig. 5. The fluorescence image of control of U87 cells (a, b, c) and incubated with DEAN-MA (d, e, f), control of HL-7702 cells (g, h, i) and incubated with DEAN-MA (j, k, l). a, d, g, j for blue channel; b, e, h, k for overlay; c, f, i, l for bright. | |
|
Download:
|
| Fig. 6. The fluorescence image of U87 cells incubated with DEAN-MA and mitotracker (a, b, c, d), lyso-tracker (e, f, g, h), ER-tracker (i, j, k, l). a, e, i for red channel of tracker; b, f, j for blue channel of DEAN-MA; c, g, k for overlay; d, h, l for bright. | |
In summary, the tandem reaction-based probes were used to detect MAO-B selectively in vitro under the suitable pH value (pH 7.4) and temperature (37 ℃). The 5.8-fold signal enhancement for DEAN-AM and 6.4-fold signal enhancement for DEAB-AM were detected and both of them showed selectivity between MAO-B and MAO-A. Importantly, the novel design strategy of utilizing targeted enzyme to build a fluorophore in the reaction process could be of great importance for the future design of fluorescent probes. Meanwhile, probe DEAB-MA showed stronger fluorescence response and less selectivity, which may be caused by the different substituent on the benzene ring from DEAN-MA. Moreover, the inhibition assay suggested the potential use of DEAB-MA and DEAN-MA on the screening of MAO-B inhibitors. Furthermore, fluorescence imaging results corresponded with previous published results about the higher level of MAO-B in nerve cells. Altogether, these results proved the great value and potential of DEAB-MA and DEAN-MA in the future study of MAO-B.
AcknowledgmentsThis work was financially supported by the National Natural Science Foundation of China (Nos. 21572147 and 21472131). We also thank the Comprehensive Training Platform of Specialized Laboratory, College of Chemistry, Sichuan University for sample analysis.
Appendix A. Supplementary dataSupplementary material related to this article can be found, in the online version, at doi:10.1016/j.cclet.2018.05.018.
| [1] |
D.E. Edmondson, C. Binda, A. Mattevi, Arch. Biochem. Biophys. 464 (2007) 269-276. DOI:10.1016/j.abb.2007.05.006 |
| [2] |
J.K. Mallajosyula, S.J. Chinta, S. Rajagopalan, et al., Neurotox. Res. 16 (2009) 186-193. DOI:10.1007/s12640-009-9032-2 |
| [3] |
X. Zhao, C. Wang, W.G. Cui, et al., Sci. Rep. 5 (2015) 9043-9055. DOI:10.1038/srep09043 |
| [4] |
A.F. Brooks, X. Shao, C.A. Quesada, et al., ACS. Chem. Neurosci. 6 (2011) 1965-1971. |
| [5] |
R. Ueki, K. Yamaguchi, H. Nonaka, et al., J. Am. Chem. Soc. 134 (2012) 12398-12401. DOI:10.1021/ja305051u |
| [6] |
K. Yamaguchi, R. Ueki, H. Nonaka, et al., J. Am. Chem. Soc. 133 (2011) 14208-14211. DOI:10.1021/ja2057506 |
| [7] |
W.H. Zhou, M.P. Valley, J. Shultz, et al., J. Am. Chem. Soc. 1 (2006) 3122-3123. |
| [8] |
S. Long, L. Chen, Y. Xiang, et al., Chem. Commun. 48 (2012) 7164-7166. DOI:10.1039/c2cc33089j |
| [9] |
X. Wu, L. Li, W. Shi, et al., Anal. Chem. 88 (2016) 1440-1446. DOI:10.1021/acs.analchem.5b04303 |
| [10] |
L.L. Li, K. Li, Y.H. Liu, et al., Sci. Rep. 6 (2016) 31217-31225. DOI:10.1038/srep31217 |
| [11] |
X. Li, J. Yu, Q. Zhu, et al., Analyst 139 (2014) 6092-6095. DOI:10.1039/C4AN01195C |
| [12] |
D. Kim, S.H. Baik, S. Kang, et al., ACS. Cent. Sci. 2 (2016) 967-975. DOI:10.1021/acscentsci.6b00309 |
| [13] |
G. Chen, D.J. Yee, N.G. Gubernator, et al., J. Am. Chem. Soc. 127 (2005) 4544-4545. DOI:10.1021/ja0428457 |
| [14] |
D. Kim, S. Sambasivan, H. Nam, et al., Chem. Commun. 48 (2012) 6833-6835. DOI:10.1039/c2cc32424e |
| [15] |
T.T. He, H.H. He, X. Luo, et al., Science (Wash., DC, United States) 47 (2015) 945-954.
|
| [16] |
J. Chan, S.C. Dodani, C.J. Chang, Nat. Chem. 4 (2012) 973-984. DOI:10.1038/nchem.1500 |
| [17] |
H. Kojima, N. Nakatsubo, K. Kikuchi, et al., Anal. Chem. 70 (1998) 2446-2453. DOI:10.1021/ac9801723 |
| [18] |
Z. Zhou, F. Wang, G. Yang, et al., Anal. Chem. 89 (2017) 11576-11582. DOI:10.1021/acs.analchem.7b02910 |
| [19] |
X. Chen, F. Wang, J.Y. Hyun, et al., Chem. Soc. Rev. 45 (2016) 2976-3016. DOI:10.1039/C6CS00192K |
| [20] |
Z. Li, Y. Xu, H. Zhu, et al., Chem. Sci. 8 (2017) 5616-5621. DOI:10.1039/C7SC00373K |
| [21] |
B. Li, Z. He, H. Zhou, et al., Chin. Chem. Lett. 28 (2017) 1929-1934. DOI:10.1016/j.cclet.2017.08.055 |
| [22] |
Q. Liu, L. Xue, D.J. Zhu, et al., Chin. Chem. Lett. 25 (2014) 19-23. DOI:10.1016/j.cclet.2013.11.024 |
| [23] |
M.Y. Li, P. C. Cui, K. Li, et al., Chin. Chem. Lett. 29 (2018) 992-994. DOI:10.1016/j.cclet.2017.11.011 |
| [24] |
Z.Q. Xu, J.H. Chen, L.L. Hu, Y. Tan, J. Yin, Chin. Chem. Lett. 28 (2017) 1935-1942. DOI:10.1016/j.cclet.2017.07.018 |
| [25] |
Q. Lin, C. Bao, S. Cheng, et al., J. Am. Chem. Soc. 134 (2012) 5052-5055. DOI:10.1021/ja300475k |
| [26] |
K.K. Yu, K. Li, J.T. Hou, et al., Tetrahedron. Lett. 54 (2013) 5771-5774. DOI:10.1016/j.tetlet.2013.08.046 |
| [27] |
J. Yang, K. Li, J.T. Hou, et al., Science (Wash., DC, United States) 6 (2017) 1-6.
|
| [28] |
J.T. Hou, K.P. Ko, W.X. Ren, et al., ACS Sens. 2 (2017) 1512-1516. DOI:10.1021/acssensors.7b00544 |
| [29] |
J.B. Wu, T.P. Lin, J.D. Gallagher, et al., J. Am. Chem. Soc. 137 (2015) 2366-2374. DOI:10.1021/ja512613j |
| [30] |
Z.V. Chirkova, M.V. Kabanova, S.I. Filimonov, et al., Bioorg. Med. Chem. Lett. 25 (2015) 1206-1211. DOI:10.1016/j.bmcl.2015.01.061 |
| [31] |
L. Li, C.W. Zhang, G.Y.J. Chen, et al., Nat. Commun. 5 (2014) 3276-3286. DOI:10.1038/ncomms4276 |
| [32] |
M. Braun, J.M. Kim, R.D. Schmid, Appl. Microbio. Biotech. 37 (1992) 594-598. |
| [33] |
C.D. Sun, W. Shi, Y.C. Song, W. Chen, H.M. Ma, Chem. Commun. 47 (2011) 8630-8638. |
 2019, Vol. 30
2019, Vol. 30 

