b College of Environment and Ecology, Chengdu University of Technology, Chengdu 610059, China
Uranium, one of the actinides, is a highly radioactive element and also well-known for its chemical toxicity. It was reported that high doses exposure of uranium could give rise to DNA damage [1], lung disease [2], cancer [3], etc. The World Health Organization (WHO) stipulated the maximum uranium concentration in drinking water to be 20 ng/mL [4]. The assessment of uranium contamination requires a simple and sensitive monitoring method. Current techniques for detection of uranium (commonly UO22+ in water) mainly focuses on sophisticated instrumental methods, including atomic emission spectrometry [5], atomic absorption spectrometry [6], and inductively coupled plasma mass spectrometry (ICP-MS) [7, 8]. These techniques are highly accurate and sensitive with industrial standards, but they are costly, available only in large centralized laboratories and require extensive sample pretreatment. Metal sensor is a good alternative tool for detection of UO22+ owing to its potential applications in on-site, real-time, or in-situ measurement.
Recently, DNA has been demonstrated to be ideal for construction of metal ions sensor because it can selectively bind metal ions via both the phosphate backbone and nucleobases [9]. Two main classes of functional DNA were developed for metal sensing: aptamers [10-12] and DNAzymes [13-19]. Compared to aptamer-based sensing, DNAzyme that exhibited enzyme-like activity is very appealing for sensitive detection of metal ions. As a result, a number of metal ion specific DNAzymes have been isolated and used for sensitive detection of various metal ions including Pb2+ [13], Cu2+ [14], Ce3+ [15], Hg2+ [16], Ca2+/Mg2+ [17] as well as UO22+ [18, 19].
The typical DNAzyme-based metal ion sensing system is fabricated via fluorescence resonance energy transfer (FRET) method which requires labeling a fluorophore on one end of the substrate strand and a fluorescence quencher on the corresponding end of the enzyme strand [20]. The labeling of DNA leads to complicated synthetic routes, high cost and even interference with DNAzyme cleavage. Accordingly, much effort was also paid to label-free metal ion sensors. One common way is via directly using DNA intercalating dyes such as SYBR green Ⅰ (SG) [21], ethidium bromide (EB) [22], and picogreen (PG) [23] for signaling. These dyes exhibited strong fluorescence in the presence of DNAzymes and low fluorescence after cleavage. Although these kind of metal sensors are quite simple and also can be readily used for detection of UO22+, they are always not as sensitive as the FRET-based sensors, making it difficult to detect trace UO22+ in water samples.
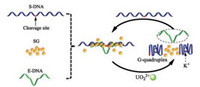
An effective approach for enhancing the sensitivity of DNAzyme-based metal sensor is to improve catalytic turnovers [20]. Herein, we developed a G-quadruplex-assisted enzyme strand recycling strategy for increasing the catalytic turnovers and further used it for detection of nanomolar UO22+. As shown in Fig. 1, the substrate strand was designed with a G-rich sequence at the both ends; after UO22+-induced cleavage, the two cleaved fragments were folded into stable G-quadruplex structure, helping the separation of the complex between enzyme strand and fragment, and also improving the recycle use of enzyme strand. Hence, sensitive and label-free detection of UO22+ can be readily achieved by this G-quadruplex-assisted enzyme strand recycling strategy.
|
Download:
|
| Fig. 1. The scheme of G-quadruplex-assisted enzyme strand recycling strategy for label-free detection of UO22+. | |
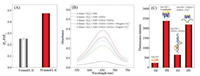
Initially, the conventional substrate sequence that was unable to form G-quadruplex after cleavage was used for construction of label-free UO22+ sensor. For the design with short hybridization length (only 4 bps), the complex of enzyme strand and substrate strand is quite unstable and easily dissociated automatically, thus resulting only a small SG fluorescent signal and also a tiny decrease after UO22+-induced cleavage (Fig. 2A). The longer hybridization length (8 bps) leads to much stronger SG fluorescent signal, but the fluorescence decrease of SG decreased only about 20% (Fig. 2B). The low fluorescent drop of SG here is probably attributed to incomplete separation between enzyme strand and the cleaved fragments. When the dsDNA interacting dye, namely SG, was incubated with the enzyme strand and split conventional substrate fragments, we found a remarkably increased fluorescent signal, indicating the easy formation of dsDNA between the enzyme strand and split conventional substrate fragment (Fig. S1 in Supporting information). This result also means the difficult separation between enzyme strand and the cleaved fragments. Therefore, G-quadruplex was subsequently used for assisting the separation of enzyme strand with the cleaved fragments and further improving its recycle use.

|
Download:
|
| Fig. 2. The UO22+-triggered fluorescence decrease using different substrate strand: (A) hybridizing 4 bps with enzyme strand, (B) hybridizing 8 bps with enzyme strand, and (C) containing G-quadruplex sequence and hybridizing 8 bps with E-DNA. Experimental conditions: substrate strand and enzyme strand concentration, 100 nmol/L; UO22+ concentration, 100 ng/mL; SG concentration, 2×; DNAzyme hybridization time, 30 min; UO22+ cleavage time, 1 h. | |
It has been demonstrated that the G-quadruplex structure can even lead to the separation of the dsDNA formed by G-quadruplex sequence and its complementary strand [24]. In the G-quadruplexassisted enzyme strand recycling strategy, the substrate strand contained G-quadruplex sequence at the both ends; after cleavage, the two fragments were folded into G-quadruplex structure in the presence of K+, leading to the complete release of enzyme strand. The result in Fig. S1 also proves that the formation of G-quadruplex will contribute to the separation of enzyme strand and cleaved substrate fragments. As expected, the strategy lead to an obvious SG fluorescence drop (>58%) in the presence of 100 ng/mL UO22+ (Fig. 2C). Therefore, sensitive and label-free quantification of UO22+ can be achieved.
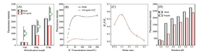
G-quadruplex was reported to be more stable in the presence of K+ [25]. The experiments showed small fluorescent decrease in the absence of K+ (Fig. 3A); upon addition of KCl solution, the fluorescent intensity has a noticeable decrease. The result implied the possible formation of G-quadruplex after cleavage. Subsequently, hemin that was known for its catalytic activity after embedding into the G-quadruplex was used for confirmation of G-quadruplex formation [26]. Fig. 3B displayed that the absorbance of 3, 3', 5, 5'-tetramethylbenzidine (TMB) signal enhanced significantly upon addition of UO22+, implying that formation of G-quadruplex which catalyze the oxidation of TMB by H2O2. To further identify the formation of G-quadruplex, we used thioflavin T (ThT) as a probe, since ThT has been confirmed as highly responsive for G-quadruplexes as compared with duplex DNA [27, 28]. As shown in Fig. 3C, after cleavage by UO22+, the fluorescent signal of ThT greatly enhanced, revealing the formation of G-quadruplex. It was noteworthy that one G-quadruplex interacts with only one ThT molecule, whereas several SG molecules could be embedded into one dsDNA strand. So, the dissociation of the complex between enzyme strand and cleaved substrate strand can lead to release several SG molecules, resulting in higher sensitivity.

|
Download:
|
| Fig. 3. Verifying the formation of G-quadruplex via the effect of K+ (A), hemin (B) and ThT (C). The experimental conditions are the same as Fig. 2. | |
Sensing conditions have profound influence on the performance of the sensor. Thus several important conditions were inspected. The effect of hybridization length between enzyme strand and substrate strand was examined firstly by fixing the enzyme strand sequence. When the hybridization length is short (4bps), the dsDNA between enzyme strand and substrate strand is not stable and easily dissociated, so the signal decline of SG is tiny in the presence of UO22+ (Fig. 4A and Fig. S2 in Supporting information); however, the long hybridization length (12bps) leads to difficult separation between the cleaved substrate strand and enzyme strand. The optimum hybridization length between enzyme strand and substrate strand is 8 bps.

|
Download:
|
| Fig. 4. The effect of hybridization length between substrate strand and enzyme strand concentration (A), K+ concentration (B), the molar ratio between S-DNA and E-DNA (C) and solution pH (D) on the detection of UO22+. | |
The effect of K+ concentration was showed in Fig. 4B and Fig. S3 in Supporting information. With the increasing amount of K+ solution, the (F0-F)/F0 values went up. When 50mmol/L of K+ solution was added, the (F0-F)/F0 reached its plateau. Afterwards, little change was observed upon addition of 100–200mmol/LK+. Therefore, 50mmol/LK+ was chosen to ensure a high (F0-F)/F0.
The cleavage reaction was also performed with different molar ratio between substrate strand and enzyme strand. As shown in Fig. 4C and Fig. S4 in Supporting information, the (F0-F)/F0 was greatly dependent on the ration of substrate strand and enzyme strand, and a maximum (F0-F)/F0 value was attained at molar ratio of 1:1. A higher concentration of substrate strand induced more cycles of DNAzyme cleavage [29], providing higher cleavage efficiency. Inadequate substrate strand will lead to the incomplete hybridization of enzyme strand as well as decreased the (F0-F)/F0. Thereby, a molar ratio of 1:1 for substrate strand and enzyme strand was considered to be optimum.
The effect of pH was investigated in the range of pH 3.0–5.5. Fig. 4D showed that the blank fluorescence signal increased as pH increased, probably because the dsDNA is more stable at low acidity. However, the complex between cleaved fragment and enzyme strand also becomes more difficult to be dissociated under low acidic conditions. Therefore, a maximum (F0-F)/F0 reached at pH 3.5 as the pH increased from 3.0 to 5.5 (shown in Fig. S5 in Supporting information). Therefore, the pH 3.5 was chosen as the optimum condition for the detection of UO22+ in this work.
It was reported that ethanol could influence the DNA hybridization [30]. Therefore, the effect of anhydrous ethanol was investigated from 0 to 20%. As shown in Fig. S6 (Supporting information), the (F0-F)/F0 value increased with the concentration of anhydrous ethanol ranging from 0 to 10%, and then decreased at the concentration higher than 10%. The complex between enzyme strand and substrate strand fragment is stable at low concentrations of ethanol and become labile at high concentration of ethanol, benefiting to the release of SG molecule. However, too high concentration of ethanol leads to low background fluorescence (Fig. S7 in Supporting information). Hence, 10% of ethanol was chosen for further work.
To test the performance of the sensor, the fluorescence decrease of SG in the presence of varied UO22+ concentrations was monitored (Fig. 5A). As shown in Fig. 5A, upon the addition of UO22+, the fluorescence intensity gradually decreased. The decreased fluorescence showed a good linear correlation with the concentration ranging from 0.2 ng/mL to 200 ng/mL. The detection of limit (LOD, 3δ) for UO22+ was calculated to be 0.06 ng/mL (about 0.2 nmol/L), which is over 25-fold lower than that of the reported DNAzyme catalytic system without G-quadruplex structure and even comparable to those obtained by labeled DNAzymes and ICP-MS (Table 1). The sensitivity of our method meets the requirement of fast detection of UO22+ in drinking waters, whose maximum allowable level is 20 ng/mL defined by the WHO. Additionally, the sensing system is quite simple and easy-to-operate because it avoids the labeling process, the signal amplification procedure as well as the use of nanomaterials.

|
Download:
|
| Fig. 5. (A) The fluorescence spectraof SG in the presence of UO22+ from0.2ng/mL to 200ng/mL (inset: the calibration curve obtained by the proposedsensing platform), (B) The fluorescence decline of the sensing system in the presence of 100ng/mL UO22+ and 5 μg/mL other metal ions. Experimental conditions: substrate strand and enzyme strand concentration, 100 nmol/L; UO22+ concentration, 100ng/mL; SG concentration, 2×; hybridization time of the two strands, 30min; UO22+ cleavage time, 1h. | |
|
|
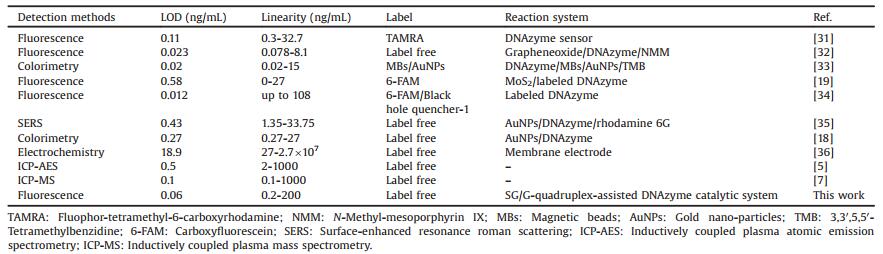
Table 1 Comparison of different methods for UO22+ determination. |
In order to investigate the specificity of the system, twelve potentially co-existing metal ions including Na+, Cd2+, Co2+, Mg2+, Mn2+, Ni2+, Zn2+, Ca2+, Hg2+, Fe3+, Pb2+ and Al3+ were also tested with our sensing system. As shown in Fig. 5B, obvious fluorescence drop was found for 100 ng/mL UO22+; other metallic ions which are 50-fold higher in concentration induced negligible fluorescence changes. This indicated that only UO22+ can trigger the cleavage of the substrate strand and the release of SG. Therefore, the proposed sensing system had a high selectivity to UO22+ over other metal ions.
To demonstrate the potential application in environmental analysis, four water samples were analyzed, i.e., one lake water sample, one river water sample and two mining water samples. UO22+ was not found in these water samples using our sensing system as well as ICP-MS. Therefore, 20 ng/mL of UO22+ were spiked to the water samples, and obvious fluorescent drops were detected. The recoveries of the four water samples ranged from 96 to 103% (Table 2), suggesting that the system can be applied for fast sensing of UO22+ in environmental water samples.
|
|
Table 2 Analytical results of UO22+ in spiked water samples. |
In conclusion, a G-quadruplex-assisted enzyme strand recycling strategy was successfully developed and further applied for label-free detection of UO22+ via SG fluorescence signalling. The cleavage of the substrate by the DNAzyme in the presence of UO22+ caused its conformation change from dsDNA to the energetically favored G-quadruplex, leading to release SG and decreased fluorescent signal. Such a simple strategy provides a rapid sensing platform for detection of UO22+ with a LOD as low as 0.06 ng/mL. Therefore, this sensing system exhibited the advantages of simplicity, high sensitivity as well as low cost. Using different metal ion specific DNAzymes, this strategy will become a promising alternative tool for the rapid and on-site detection for metal ions in environmental waters.
AcknowledgmentsThe authors gratefully acknowledge the financial support from the National Natural Science Foundation of China (Nos. 21475013 and 21305009), and China Postdoctoral Science Foundation (Nos. 2015M572453 and 2016T90839).
Appendix A. Supplementary dataSupplementary data associated with this article can be found, in the online version, at https://doi.org/10.1016/j.cclet.2018.02.003.
| [1] |
A.C. Miller, M. Stewart, K. Brooks, et al., J. Inorg. Biochem. 91 (2002) 246-252. DOI:10.1016/S0162-0134(02)00391-4 |
| [2] |
B. Furlow, Lancet Resp. Med. 2 (2014) 178-179. DOI:10.1016/S2213-2600(14)70005-0 |
| [3] |
S.K. Annamalai, K.D. Arunachalam, R. Selvaraj, Environ. Sci. Pollut. Res. 24 (2017) 15427-15443. DOI:10.1007/s11356-017-9111-5 |
| [4] |
Health Criteria and Other Supporting Information: Addendum Guidelines for Drinking-water Quality, vol. 2, World Health Organization, 1998.
|
| [5] |
M.R. Jamali, Y. Assadi, F. Shemirani, et al., Anal. Chim. Acta 579 (2006) 68-73. DOI:10.1016/j.aca.2006.07.006 |
| [6] |
A. Quentmeier, M. Bolshov, K. Niemax, Spectrochim. Acta B 56 (2001) 45-55. DOI:10.1016/S0584-8547(00)00289-5 |
| [7] |
D.W. Boomer, M.J. Powell, Anal. Chem. 59 (1987) 2810-2813. DOI:10.1021/ac00150a019 |
| [8] |
F. Claverie, A. Hubert, S. Berail, et al., Anal. Chem 88 (2016) 4375-4382. DOI:10.1021/acs.analchem.5b04802 |
| [9] |
X.B. Zhang, R.M. Kong, Y. Lu, Annu. Rev. Anal. Chem. 4 (2011) 105-128. DOI:10.1146/annurev.anchem.111808.073617 |
| [10] |
S. DasGupta, S.A. Shelke, N.S. Li, et al., Chem. Commun. 51 (2015) 9034-9037. DOI:10.1039/C5CC01526J |
| [11] |
M. Hoang, P.J.J. Huang, J.W. Liu, ACS Sens. 1 (2016) 137-143. DOI:10.1021/acssensors.5b00147 |
| [12] |
G.Q. Wang, L.X. Chen, Chin. Chem. Lett. 20 (2009) 1475-1477. DOI:10.1016/j.cclet.2009.06.029 |
| [13] |
J. Li, Y. Lu, J. Am. Chem. Soc. 122 (2000) 10466-10467. DOI:10.1021/ja0021316 |
| [14] |
J. Liu, Y. Lu, J. Am. Chem. Soc. 129 (2007) 9838-9839. DOI:10.1021/ja0717358 |
| [15] |
P.J.J. Huang, J. Lin, J. Cao, et al., Anal. Chem. 86 (2014) 1816-1821. DOI:10.1021/ac403762s |
| [16] |
M. Hollenstein, C. Hipolito, C. Lam, et al., Angew. Chem. Int. Ed. 47 (2008) 4346-4350. DOI:10.1002/(ISSN)1521-3773 |
| [17] |
W.H. Zhou, Y.P. Zhang, J.S. Ding, et al., ACS Sens. 1 (2016) 600-606. DOI:10.1021/acssensors.5b00306 |
| [18] |
J.H. Lee, Z.D. Wang, J.W. Liu, et al., J. Am. Chem. Soc. 130 (2008) 14217-14226. DOI:10.1021/ja803607z |
| [19] |
H.Y. Zhang, Y.J. Ruan, L. Lin, et al., Spectrochim. Acta A 146 (2015) 1-6. DOI:10.1016/j.saa.2015.02.113 |
| [20] |
W.H. Zhou, R. Saran, J.W. Liu, Chem. Rev. 117 (2017) 8272-8325. DOI:10.1021/acs.chemrev.7b00063 |
| [21] |
L.L. Zhang, Y.Y. Zhang, M.J. Wei, et al., New J. Chem. 37 (2013) 1252-1257. DOI:10.1039/c3nj41103f |
| [22] |
D. Ferrari, A. Peracchi, Nucleic Acids Res. 30 (2002) e112. DOI:10.1093/nar/gnf111 |
| [23] |
L.B. Zhang, B.Y. Han, T. Li, et al., Chem. Commun. 47 (2011) 3099-3101. DOI:10.1039/c0cc04523c |
| [24] |
B.L. Li, H. Wei, S.J. Dong, Chem. Commun. 1 (2007) 73-75. |
| [25] |
Z.G. Wang, J.P. Liu, J. Mol. Graphics Model 72 (2017) 168-177. DOI:10.1016/j.jmgm.2017.01.006 |
| [26] |
D.M. Kong, J. Wu, N. Wang, et al., Talanta 80 (2009) 459-465. DOI:10.1016/j.talanta.2009.07.010 |
| [27] |
H. Cai, C. Zhou, Q. Yang, et al., Chin. Chem. Lett. 29 (2018) 531-534. DOI:10.1016/j.cclet.2017.09.010 |
| [28] |
T.X. Chen, F. Ning, H.S. Liu, et al., Chin. Chem. Lett. 28 (2017) 1380-1384. DOI:10.1016/j.cclet.2017.01.006 |
| [29] |
W. Cui, L. Wang, W. Jiang, Biosens. Bioelectron. 77 (2016) 650-655. DOI:10.1016/j.bios.2015.10.040 |
| [30] |
J. Kypr, I. Kejnovska, D. Renciuk, et al., Nucleic Acids Res. 37 (2009) 1713-1725. DOI:10.1093/nar/gkp026 |
| [31] |
S.J. Xiao, J. Zuo, Z.Q. Zhu, et al., Sensor. Actuat. B-Chem. 210 (2015) 656-660. |
| [32] |
M.H. Li, Y.S. Wang, J.X. Cao, et al., Biosens. Bioelectron. 72 (2015) 294-299. DOI:10.1016/j.bios.2015.05.032 |
| [33] |
H.Y. Zhang, L. Lin, X.X. Zeng, et al., Biosens. Bioelectron. 78 (2016) 73-79. DOI:10.1016/j.bios.2015.11.024 |
| [34] |
J.W. Liu, A.K. Brown, X.L. Meng, et al., Proc. Natl. Acad. Sci. U. S. A. 104 (2007) 2056-2061. DOI:10.1073/pnas.0607875104 |
| [35] |
Z.L. Jiang, D.M. Yao, G.Q. Wen, et al., Plasmonics 8 (2013) 803-810. DOI:10.1007/s11468-012-9476-8 |
| [36] |
M. Shamsipur, M. Saeidi, A. Yari, et al., Bull. Korean Chem. Soc. 25 (2004) 629-633. DOI:10.5012/bkcs.2004.25.5.629 |
 2019, Vol. 30
2019, Vol. 30