Marine-derived organisms have been recognized as a great reservoir for biologically active and structurally unusual secondary metabolites that have greatly impacted the quality of human life [1-6]. For example, bacteria derived from marine environment have yielded several promising drug candidates such as lipopeptide antibiotic taromycin [7], anticancer agentssalinosporamide A [8-10] and bryostatins [11, 12]. Recently, we have reported the characterization of a rare macrodilactone named streptoseomycin with potent anti-Helicobacter pyroli activity from Streptomyces seoulensis A01 [13], which was originally isolated from a marine prawn, and an antitumor agent chartreusin [14], as well as antibacterial alkaloids, strepchazolins and streptazolin from a marine sediment-derived S. chartreusis NA02069 [15]. Our continued chemical investigation of S. chartreusis NA02069 strain has led to the discovery of a novel alkaloid, chartrenoline (1), featuring a unique 6/6/5/5 ring system (Fig. 1). Herein, we report the isolation, structure elucidation and putative biosynthetic pathway for 1.
|
Download:
|
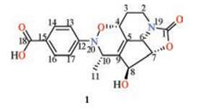
| Fig. 1. Structure of chartrenoline (1). | |
Compound 1 was obtained as pale yellow crystals. The molecular formula of 1was determined to be C18H18N2O6 based on its HR-ESI-MS data (m/z 359.1237 [M + H]+, calcd. for C18H19N2O6: 359.1238), indicating 11 degrees of unsaturation. Analysis of the 13C NMR spectrum, together with the DEPT135 and HSQC spectra, indicated ten sp2-hydribridzed carbon resonances ascribe to two carbonyl carbons, four olefinic methines and four olefinic carbonyl carbons; eight sp3-hydribridzed carbon signals including one methyl group, two methylenes, and five methine carbons. The 1H NMR spectrum data (Table 1) exhibited signals for a pair of 1, 4-disubstituted phenyl ring at δH 7.85 (d, 2H, J = 8.9 Hz, H-14, H-16) and 7.11 (d, 2H, J = 8.9 Hz, H-13, H-17). In addition, the 1H-1H COSY spectral data revealed a partial structure of -CH2-CH2-CH-, -CH-CH-CH-OH-, and CH3-CH-, respectively; indicating the connection of C-2/C-3/C-4, C-6/C-7/C-8/8-OH, and C-11/C-10, respectively.
|
|
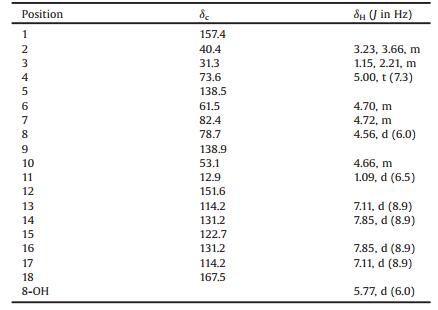
Table 1 1H and 13C NMR spectral data for 1 |
The gross structure of 1 was established by extensive analyses of 2D NMRspectral data, in particular with HMBC spectrum. The HMBC correlations of H-2 (δH 3.66) with C-4 (δC 73.6) and C-6 (δC 61.5), of H-3 (δH 2.21) with C-5 (δC 138.5), of H-7 (δH 4.72) with C-5 (δC 138.5) and C-9 (δC 138.9), of H-8 (δH 4.56) with C-6 (δC 61.5), together with the chemical shifts of δC 40.4, δC 61.5 and δC 82.4 for C-2, C-6 and C-7, respectively, suggested the presence of a 6/5 bicyclic ring system as shown in Fig. 2. The presence of a hydroxyl group substituted at C-8 was reinforced by HMBC correlations of 8-OH (δH 5.77) with C-7 (δC 82.4), C-8 (δC 78.7) and C-9 (δC 138.9). In addition, in the HMBC spectrum, H-2 (δH 3.66), H-6 (δH 4.70) and H-7 (δH 4.72) showed correlations to a downfield carbon C-1 (δC 157.4) indicated the existence of an oxazolidin-2-one ring.
|
Download:
|
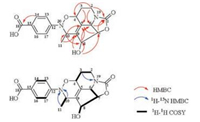
| Fig. 2. Key 2D NMR correlations of 1. | |
The HMBC correlations of H-11 (δH 1.09) with C-9 (δC 138.9), and of H-10 (δH 4.66) with C-5 (δC 138.5) and C-8 (δC 78.7) revealed that C-10 (δC 53.1) was connected to C-9 (δC 138.9). Moreover, a benzoic acid partial structure was elucidated by the HMBC correlation of H-14/16 (δH 7.85) with C-18 (δC 167.5). The HMBC correlation between H-10 (δH 4.66) and C-12 (δC 151.6), together with the chemical shift for C-10 (δC 53.1), revealed the benzoic acid group and C-10 was connected through a nitrogen atom. The only remaining oxygen atom was substituted at C-4 as revealed by chemical shifts of H-4 (δH 5.00) and C-4 (δC 73.6). Finally, owing to the shortage of any more double bonds, the last unsaturation degree could be ascribe to an oxazine ring through generating an O-N bond. The proposed structure was further supported by the 1H-15N HMBC correlations of H-10 and H-11 with N-20, and of H-3 with N-19.
Compound 1 contained five chiral centers, which were difficult to be fully solved by analysis of its NMR data. Fortunately, a high quality crystal suitable for X-ray diffraction was obtained from CH2Cl2-methanol (1/1, v/v) solvent mixture. The subsequent X-ray crystallographic data collected in Cu Kα radiation at low temperature with a flack parameter of -0.07 (14) confirmed the above proposed planar structure (Fig. 3), and further established the absolute configuration of 1 as shown in Fig. 1. Compound 1 was therefore determined as an unusual 6/6/5/5-tetracyclic alkaloid with a novel skeleton.
|
Download:
|
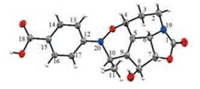
| Fig. 3. X-ray crystallographic analysis of compound 1. | |
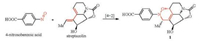
Structurally, compound 1 is actually related to streptazolin [16], which has been isolated from the same strain when fermented using different medium [15]. Therefore, a plausible biosynthetic pathway for 1 is proposed as shown in Scheme 1. Streptazolin is proposed to be biosynthesized through bacterial type Ⅰ polyketide synthases (PKSs) [17]. The C-terminal thioester reductase domain in PKS is responsible for release of the assembled polyketide chain as an aldehyde, and its subsequent reductive amination and cyclization gives streptazolin [17, 18]. Streptazolin can then undergo an intermolecular Diels-Alder reaction with 4-nitrosobenzoic acid to give 1 [19].
|
Download:
|
| Scheme 1. Proposed biosynthetic pathway for chartrenoline (1) | |
Compound 1 was evaluated for its antibacterial activities against four pathogenic bacteria including Staphylococcus aureus CMCC 26003, Bacillus subtilis CICC 10283, Escherichia coli ATCC 25922, and Pseudomonas aeruginosa CICC 10351. However, compound 1 is almost inactive to them at the concentration of 128 μmol/L.
In conclusion, a novel alkaloid, chartrenoline (1), was isolated from a marine actinobacterium S. chartreusis NA02069. The structure of 1 possessing a novel skeleton with a fused 6/6/5/5 ring system was unambiguously established by extensive analysis of its spectroscopic data including HR-ESI-MS, NMR, as well as single crystal X-ray diffraction data. Furthermore, the proposed biosynthetic pathway starting from a known alkaloid streptazolin was proposed. Our work presented here sets the stage for future biosynthetic study.
AcknowledgmentsThis work was financially supported by the National Natural Science Foundation of China (Nos. 81522042, 21572100, 81773591, 81421091, 81500059, 81673333, 21672101, 21761142001, 21661140001, J1210026, and J1103512), Natural Science Foundation of Jiangsu Province (No. BK20151397), and Fundamental Research Funds for the Central Universities (No. 020814380092).
Appendix A. Supplementary dataSupplementary material related to this article can be found, in the online version, at doi:https://doi.org/10.1016/j.cclet.2018.10.030.
| [1] |
J.W. Blunt, A.R. Carroll, B.R. Copp, et al., Nat. Prod. Rep. 35 (2018) 8-53. DOI:10.1039/C7NP00052A |
| [2] |
M. Kita, H. Kigoshi, Nat. Prod. Rep. 32 (2015) 534-542. DOI:10.1039/C4NP00129J |
| [3] |
L.A. Salvador-Reyes, H. Luesch, Nat. Prod. Rep. 32 (2015) 478-503. DOI:10.1039/C4NP00104D |
| [4] |
D.Q. Xue, H.L. Liu, S.H. Chen, et al., Chin. Chem. Lett. 28 (2017) 1190-1193. DOI:10.1016/j.cclet.2017.03.040 |
| [5] |
R. Montaser, H. Luesch, Future Med. Chem. 3 (2011) 1475-1489. DOI:10.4155/fmc.11.118 |
| [6] |
Y. Gao, W. Xian, H.C. Liu, et al., Chin. Chem. Lett. 28 (2017) 905-908. DOI:10.1016/j.cclet.2017.01.004 |
| [7] |
K.A. Reynolds, H. Luhavaya, J. Li, et al., J. Antibiot. 71 (2018) 333-338. DOI:10.1038/ja.2017.146 |
| [8] |
D. Zhou, C. Wang, M.L. Li, et al., Chin. Chem. Lett. 29 (2018) 191-193. DOI:10.1016/j.cclet.2017.06.007 |
| [9] |
R.H. Feling, G.O. Buchanan, T.J. Mincer, et al., Angew. Chem. Int. Ed. 42 (2003) 355-357. DOI:10.1002/anie.200390115 |
| [10] |
T.A.M. Gulder, B.S. Moore, Angew. Chem. Int. Ed. 49 (2010) 9346-9367. DOI:10.1002/anie.201000728 |
| [11] |
S.K. Davidson, S.W. Allen, G.E. Lim, et al., Appl. Environ. Microb. 67 (2001) 4531-4537. DOI:10.1128/AEM.67.10.4531-4537.2001 |
| [12] |
B. Zhang, K.B. Wang, W. Wang, et al., Org. Lett. 20 (2018) 2967-2971. DOI:10.1021/acs.orglett.8b01006 |
| [13] |
A.E. Trindade-Silva, G.E. Lim-Fong, K.H. Sharp, et al., Curr. Opin. Biotech. 21 (2010) 834-842. DOI:10.1016/j.copbio.2010.09.018 |
| [14] |
Y.S. Wang, B. Zhang, J.P. Zhu, et al., J. Am. Chem. Soc. 140 (2018) 10909-10914. DOI:10.1021/jacs.8b06623 |
| [15] |
C.L. Yang, Y.S. Wang, C.L. Liu, et al., Mar. Drugs 15 (2017) 244. DOI:10.3390/md15080244 |
| [16] |
S.F. Bi, F. Li, Y.C. Song, et al., Nat. Prod. Commun. 6 (2011) 353-355. |
| [17] |
U.R. Awodi, J.L. Ronan, J. Masschelein, et al., Chem. Sci. 8 (2017) 411-415. DOI:10.1039/C6SC02803A |
| [18] |
W. Huang, S.J. Kim, J. Liu, et al., Org. Lett. 17 (2015) 5344-5347. DOI:10.1021/acs.orglett.5b02707 |
| [19] |
S. Garbley, H. Kluge, H.U. Hope, et al., Angew. Chem. Int. Ed. 26 (1987) 664-665. |
 2019, Vol. 30
2019, Vol. 30