Lanthanide-doped upconversion nanoparticles (UCNPs), which can convert near-infrared (NIR) excitation light into high energy anti-Stokes luminescence, have recently emerged as a new generation of promising optical nanomaterial. The anti-Stokes emission of UCNPs attributes to the successive absorption of multiple photons, based on the long life-time energy levels in the inner f-orbitals of embedded trivalent lanthanide ions [1-3]. Such an emission behavior is greatly different from downconversion materials including organic dyes and quantum dots, and endows UCNPs with a series of additional advantages [4]. Since most fluorophores cannot be excited by NIR light, UCNPs can greatly decrease the background fluorescence and improve the detection sensitivity for potential applications in biosensing and bioimaging [5-7]. In addition, as NIR range (700-1100 nm) refers to the "optical transparency window" of biological tissues, the exploitation of NIR excitation of UCNPs shows unique merits including deeper tissue penetration and decreased photodamage effects, promising UCNPs as a powerful platform in tissue imaging and therapy [8-10]. Moreover, UCNPs display other intriguing properties including easy synthesis, ready modification, and high resistance to photobleaching [11, 12]. Benefiting from these advantages and properties, UCNPs have attracted increasing attention in biosensing, bioimaging and disease therapy. To implement UCNPs as optical functional nanomaterials for practical biomedical applications, it is of critical significance to rationally optimize the interaction between UCNPs and biosystems, including biomolecules, cells and tissues [1, 4, 6]. For instance, to improve the target-recognition ability for biosensing and bioimaging, optimizing the interaction selectivity between UCNPs and targets is necessary [11, 13, 14]. As for cancer therapy, it is of great importance to improve the affinity between UCNPs and nidus to enhance the therapy efficacy [5, 15]. However, UCNPs per se cannot interact to targets and nidus with high selectivity and high affinity. In this regard, integration of functional molecules to UCNPs can provide an efficient strategy to optimize the interaction between UCNPs and biosystems.
Long recognized as the carrier of genetic information, DNA has recently attracted increasing interest as a functional biomolecule to construct elaborate materials [16]. Specifically, DNA can form double helix through Watson-Crick base-pairing principle [17]. Moreover, owing to the precise nature of base-pairing principle, such self-assembly behavior of DNA can be highly programmed at nanometer scale to optimize the interaction between DNA and sophisticated nanostructures [18-20]. Besides the properties of self-assembly, DNA molecules also reveal additional functions. Through van der Waal's force, hydrogen bonding, hydrophilic hydrophobic interaction and electrostatic interaction, DNA sequence can fold into two-dimensional and three-dimensional structure and bind to target with high affinity and selectivity, which endows DNA with the capacity of molecular recognition [21-24]. Besides, DNA also exhibits therapeutic effects such as regulation of protein activity and suppression of RNA expression, demonstrating potential applications in disease therapy [25, 26]. These properties and functions make DNA an outstanding functional element to optimize the interaction between UCNPs and biosystems [27-30]. Functionalization of UCNPs with DNA molecules allows for a variety of applications in selective biosensing, targeted bioimaging and therapy.
For DNA-functionalized upconversion nanomaterials, UCNPs generally act as efficient photoconverter and signal generator, while DNA further endows UCNPs with specific recognition ability, material assembly potential and drug loading capacity. The combination of UCNPs and DNA greatly promotes the application of upconversion material in biomedicine area. In this review, we conclude recent biomedical progresses of DNA-functionalized UCNPs. In particular, design principle, synthesis strategies and applications in biosensing, bioimaging and therapy are highlighted to provide the state of the art. Finally, more insights of future perspectives are also offered, aiming to further motivate their application in biomedicine field
2. Design and synthesis strategies of DNA-functionalized UCNPsAttaching DNA on UCNPs is the most crucial step for material application. In terms of disparate application fields, the design criteria and synthetic strategies of DNA-functionalized UCNPs are different. For biodetection and bioimaging, DNA is generally conjugated with UCNPs to endow this material with target recognition ability and material assemble capacity. For disease therapy, especially gene therapy, as nucleic acids are used as therapeutic agents and need to be released to nidus, they are usually attached with modified UCNPs through reversible encapsulation. Following we would discuss the design and synthesis strategies of DNA-functionalized UCNPs in details.
2.1. Conjugation of DNA and UCNPsThe conjugation of DNA and UCNPs is essential for their application in biosensing and bioimaging. The key of experiment design is to make use of the developed nanoparticle modification strategies and biomolecule conjugation methods, ensuring that the properties of both UCNPs and DNA would not be influenced after conjugation. Generally, three steps are required for such conjugation. Firstly, hydrophobic synthesized UCNPs are converted to hydrophilic nanoparticles through surface modification strategies, including polymer capping, surface silanization and ligand exchange. Secondly, single-stranded DNA is modified with functional groups, such as amino groups and thiol groups, at the terminus for further conjugation. Because of the left extra functional groups on both the UCNP surface and DNA terminus, biocompatible coupling methods could be subsequently employed for their attachment. For instance, in biosensing with DNAfunctionalized UCNPs [27, 31-36], oleic acid-capped UCNP was firstly coated with polyacrylic acid (PAA) polymer shell through ligand exchange. Oligonucleotides were modified with amino groups at the terminus. Then EDC/NHS coupling method was introduced to link carboxyl-modified UCNPs and amino-modified DNA via catalyzed condensation reactions. The reaction condition is gentle and the related operations are simple, which has been widely applied for biomolecule conjugation. Another similar method is Sulfo-SMCC coupling. In a typical procedure, poly ethylenimine (PEI)-capped UCNPs and thiol-modified DNA are conjugated by Sulfo-SMCC linker through two-step chemical reactions [28, 37]. Other polymer coating including polydopamine (PDA) [38, 39] and biocompatible cross-linking agents such as glutaraldehyde [40, 41] were also used for UCNP-DNA conjugation. Besides polymers, silica shells are frequently applied for UCNP surface modification and subsequent conjugation. In particular, UCNPs were firstly coated with a SiO2 layer through the wellknown StÖber process. Then the silica layer was activated by CNBr acetonitrile solution and conjugated with 50 end amino-modified DNA strands. Successful detection of target gene was achieved in this system [32].
Despite the advantages of aforementioned conjugation strategies, a series of modification and coupling procedures are still required. Therefore, these methods are usually assumed as indirect conjugating methods between DNA and UCNP. The invention of a direct conjugation method without preliminary modifications would simplify the operation procedures and further promote their biomedical application. Recently, Lu's group reported such a one-step ligand exchange method for the direct DNA-UCNP conjugation [29]. Specifically, a chloroform solution containing oleic acid-capped UCNPs and a water solution of unmodified DNA were mixed and vigorously stirred for 24 h. Because of the specific interaction between negatively charged phosphates with surface lanthanide ions in UCNPs after DNA attachment, it was observed that the UCNPs were transferred into the water layer from the chloroform layer. The resulting DNA-UCNPs were separated after the phase transfer and re-dispersed in water. Noteworthy, the characteristic upconversion emission of UCNPs and the baseparing nature of DNA were not influenced by the conjugation. This simple and direct conjugation method was a breakthrough in synthetic strategies of DNA-functionalized UCNPs and greatly contributed to their applications in biosensing [13, 42] and bioimaging [29].
2.2. Encapsulation of DNA on UCNPsFor UCNP-based gene therapy, the key of design criteria is to ensure that payload gene could reach to target sites without previous loss. Overall, three strategies are often used for this endeavor, which are silica coating-based physical absorption, polymer coating-based electrostatic absorption and photolabile material-based absorption.
Porous silica coating is an effective molecule loading method for UCNPs, due to the highly specific surface area in the silica layer that is ideal to physical encapsulation of payload molecules. For instance, Zhang's group utilized silica coating for the absorption of DNA on UCNPs for gene therapy [43]. After the coating of a silica shell on UCNPs, the solution of UCNPs was mixed with solution containing small interfering RNAs (siRNAs) and incubated for appropriate time. Successful loading was proved by gel electrophoresis and the detachment of siRNA in cell experiment was characterized by fluorescence spectroscopy. Asides from silica coating, mesoporous silica coating with even larger pore size (2- 50 nm) and higher surface area is attractive for loading efficiency improvement. Gu et al. [44] reported the high loading level of nucleic acids into mesoporous silica with a range between ~75.4 mg/g to ~111.5 mg/g, in comparison with that of dense silica ranging between ~40 to ~50 mg/g. The employment of mesoporous silica coating with UCNPs has greatly promoted the application of DNA-functionalized UCNPs in disease therapy.
Polymer-based electrostatic absorption is another widely used method for DNA encapsulation. Positively charged polymer shells could absorb negatively charged nucleic acid and realize further delivery. For instance, using layer-by-layer engineering technique, Duan's group synthesized UCNP-PAA@PEI and attached siRNA on these nanoparticles through facile incubation for 60 min [54]. The encapsulation efficiency and loading capacity were determined to be 97.264% and 34.1%, respectively. He et al. reported that an extra PEI layer was effective in both the reduction of cytoxicity and the enhancement of gene transfection ability through the use of UCNPPEG@2×PEI for siRNA loading [46]. More efforts should be focused on related investigation.
Although successful absorption and detachment of gene were achieved with these strategies, the release of gene in a controlled manner was not realized in above systems. Disadvantages such as the irregular payload release or burst release in the initial stage were observed [47]. Light-controlled gene delivery can be realized through the caging of therapeutic gene with photolabile groups, and subsequent uncaging under photoactivated group degradation. For instance, Xing et al. constructed a gene delivery platform with the employment of catonic photoactive linker o-nitrobenzyl [48]. This linker was covalently conjugated on silica-coated UCNPs and anionic siRNA was therefore absorbed to this material through electrostatic interaction. Successful siRNA delivery in a lightcontrolled manner was proved by in vitro agarose gel electrophoresis and intracellular imaging. Other photoactive materials were also used for controlled release and would be specifically introduced in Section 5.1.
In conclusion, with the development of carbodiimide chemistry, amine chemistry, thiol chemistry and silica chemistry, various attaching methods have been used for UCNP-DNA interaction. For further improvement of the conjugation of these two moieties, more focus should be put on the invention and optimization of direct conjugation methods to simplify the synthesis procedures and promote subsequent application. For encapsulation of DNA on UCNPs, combination of photoactive compounds with gene is a new encapsulation direction, which is promising for light-controlled molecule release.
3. DNA-functionalized UCNPs in biosensingUCNP-based biosensing has two main advantages: (ⅰ) Nearly negligible background autofluorescence in carrier fluid allows for decreased detection of limit (LOD). (ⅱ) Stable and environmentinert fluorescence signals provide accurate and convincing analysis results [49]. However, there are two crucial steps in biosensing that cannot be solely satisfied by upconverison materials. Firstly, the detection platform needs to have target recognition ability. Secondly, after the biochemical recognition procedure, targetresponsive signal needs to be generated. Nevertheless, UCNPs cannot specifically recognize analytes. Their luminescence also does not directly relate to any biochemical property of a system [1]. Thus, it is necessary to combine UCNPs with suitable functional materials in biosensing.
For the target recognition procedure, introducing recognition elements to upconversion materials is necessary. Among various choices, nucleic acid has attracted increasing attention for two main reasons. On the one hand, as the analysis of DNA/RNA per se is of great importance in biomedicine, it is obvious that targetcomplementary oligonucleotide can function as probes in gene detection through base-paring [50, 51]. On the other hand, aptamers, which are single-strand functional oligonucleotides with high affinity towards various targets [52, 53], greatly expand the detection range of DNA-functionalized UCNPs. In fact, in comparison with other recognition elements (e.g., antibodies and enzymes), aptamers exhibit additional advantages such as wider range of targets, small sizes and easy modification. As for the signal transduction procedure, DNA can also act as an excellent medium to transform the luminescence of UCNPs into a target-responsive signal. Specifically DNA can function as the linkers in material assembly of UCNPs with other components for signal transduction [17]. With the employment of sequence design and enzymatic toolkit, this interaction shows more programmability than previous employed strategies, such as the biotin-avidin interaction or immunoreaction between antibodies and antigens [54]. These unique properties of DNA simplify the material assembly process and expand the interacted material scopes.
As mentioned above, DNA-functionalized UCNPs are endowed with both target-recognize ability and signal-transform capacity, which provide an outstanding platform for the detection of various biomolecules. In the following section, we classify the biosensing with DNA-functionalized UCNPs into two categories, which are homogenous assay and heterogeneous assay respectively, and would specifically discuss their design strategies, detection principles and relevant applications.
3.1. Homogenous assayA homogeneous assay is carried out in solution through the simple mix of analytes and sample [1]. In general, the majority of UCNP-related homogeneous assay utilizes luminescence resonance energy transfer (LRET) as the signal transduction mechanism. LRET is a nonradiative process containing energy transfer from an excited state donor to a ground state acceptor [55, 56]. Donor-acceptor separation distance and appropriate spectral overlap between donor emission and acceptor absorption are two important factors influencing the efficiency of LRET [40]. In LRET-based assays, UCNPs are promising donors because they are excited by NIR light, which lies discretely from the excitation spectra of most acceptors [57]. Thus, the cross-talk problem can be minimized and a relative high signal-to-background ratio can be achieved. However, to construct an efficient LRET system, UCNPs need to be connected with acceptors through linkers with suitable length, as the LRET efficiency relies heavily on the donor-acceptor distance (within 10 nm). The high programmability and developed modification methods make DNA probes promising LRET linkers.
The first LRET system employing DNA-functionalized UCNPs was developed by Zhang and his coworkers for the detection of nucleic acids [58], in which DNA functions as the linker between UCNP donor and organic dye acceptor, as well as the recognition sequence of target DNA. As mentioned in Section 2, UCNPs were conjugated with 50 end amino-modified DNA strands (DNA 1) through silica chemistry. Another DNA probe (DNA 2) was labeled with fluorophore TAMRA also at the 50 end to ensure appropriate distance between donors and acceptors. In the absence of target DNA, the distance between randomly diffused donors and acceptors were not close enough for LRET. With the targets, because both DNA 1 and DNA 2 were complementary to adjacent regions of the target DNA, UCNP and TAMAR fluorophore were brought into close proximity and the fluorescence energy was transferred from UCNP to TAMAR (Fig. 1A). Subsequently the signal of UCNP (537 nm) decreased and that of TAMAR (575 nm) increased, generating target-responsive signal. In this assay, a LOD of 1.3 nmol/L was obtained. Afterwards, this detection mode has been developed in several aspects. Detecting procedures are simplified using label-free method with the application of intercalation dye [27, 31, 32, 59]. The detection capacity was also expanded by exploiting different emission parameters of UCNPs [60, 61]. One of the most important developments is the replacement of original oligonucleotide probes with aptamers [33, 40, 61]. Because aptamers can be selected towards numerous targets through the systematic evolution of ligands by exponential enrichment (SELEX) approach, the detection range of this model is largely extended. For instance, Wang et al. used this model for the detection of antibiotic oxytetracycline (OTC). OTC aptamers were immobilized on UCNPs. Aptamer-complementary DNA probes (cDNA) were attached with quencher SYBR Green Ⅰ and then added to the solution. Without OTC, the fluorescence of UCNPs was quenched by SYBR Green Ⅰ because of their close proximity in aptamer-cDNA double helix. In the presence of OTC, considering the stronger interaction between aptamers and OTCs, the double helix unwound and the fluorescence of UCNPs recovered (Fig. 1B). A linear calibration was obtained ranging from 0.1ng/mL to 10ng/mL with a LOD of 0.054ng/mL.

|
Download:
|
| Fig. 1. Scheme representations of homogeneous assays with DNA-functionalized UCNPs. (A) Detection of target gene with fluorophore as LRET acceptors. Reproduced with permission [58]. Copyright 2006, American Chemical Society. (B) Detection of antibiotics with fluorophore as LRET acceptor. Reproduced with permission [40]. Copyright 2015, Elsevier. (C) Detection of target DNAwith AuNP as LRETacceptors. Reproduced with permission [63]. Copyright 2015, American Chemical Society. (D) Au-UCNP pyramid probe for miRNA detection. Reproduced with permission [62]. Copyright 2016, American Chemical Society. (E) Detection of ATP with graphene as LRETacceptors. Reproduced with permission [36]. Copyright 2011, Royal Society of Chemistry. | |
Although organic dyes have been generally applied as LRET acceptors, they suffer from self-quenching at high concentrations, susceptibility to photobleaching, and narrow absorption windows [55, 56], impeding the sensitivity improvement of UCNP-based LRETassay. Gold nanoparticles (AuNPs), instead, have been applied as the alternative of organic dyes. Their surface plasmon reference absorption lies in the NIR-to-IR region. They also have larger surface area and longer working distance than organic dyes. These properties make them better UCNP quenchers. Original UCNPAuNP system was developed by Li et al. for the detection of avidin [54]. UCNP and AuNP were both modified with biotin and the luminescence would be quenched with the addition of avidin. However, employing biotin-avidin interaction as the "linker" confined the further expansion of detection targets. The introduction of DNA, especiallyaptamers, intothis system greatly promotes the detection scope, which contains nucleic acids [41, 45, 62-65], metal ions [34, 66], protein [35, 67, 68], bacteria [69], etc. For instance, Zourob et al. designed a hairpin-shaped molecular beacon (MB) with UCNP-decorated PSA/SiO2 nanohybrids as donor at one end and the [(ppy)2Ir(dcbpy)(4-ABT)2]/AuNPs complex as an acceptor at the other end for DNA detection [63]. With the target, the formation of double helix broke the MB structureand extended the donor-acceptor distance, restoring the luminescence signal (Fig. 1C). Results showed a linear relationship between the fluorescence intensity and target DNA concentration down to the picomolar. This study combines the programmability of DNA probes, the high quenching efficiency of AuNPs and the assistant quenching effect of Ir(Ⅲ) complexes, which noticeably improves the detection sensitivity and deserves further investigation towards multiplexed detection. Besides unitary fluorescence assay, dual-modal detection could also be achieved with the application of gold nanoparticles as acceptors. For instance, Kuang's group designed an Au-UCNP pyramid probe for miRNA detection, utilizing the self-assemble behavior of DNA [62]. In particular, each DNA frame side was embedded with miRNA recognition sequences (part a) and noncomplementary sequences (part b). When miRNA was present, it would hybridize with the recognition sequence on each side and subsequently dissociated the DNA frame. The UCNPs and AuNPs were separated, which leaded to the decrease of plasmonic circular dichroism (CD) intensity and the increase of UCNP fluorescence intensity (Fig. 1D). Experimental results showed that the CD intensity had a LOD of 0.03fmol/10 mg RNA, while the luminescence intensity had a LOD of 0.12fmol/ 10 mg RNA. This system provides a practical and valuable platform for multimode detection using DNA-functionalized UCNPs. Afterwards, detection of thrombin and antigen with aptamer-based AuUCNP pyramids [65] and multiplexed detection of miRNA using Ag2S-Au-UCNP pyramids [45, 70] were also achieved.
The development of UCNP quenchers continued. Single-strand DNA (ssDNA) was reported being strongly absorbed on graphene oxide (GO) through π-π stacking between nucleobases of DNA and sp2 atoms of GO [71]. Considering the large absorption range of GO from 200nm to 800nm, it has been further exploited as the quencher of DNA-functionalized UCNPs [36, 42, 72-74]. The pioneered attempt was achieved by Li and his co-workers [36]. In this study, ATP aptamer and UCNPs were conjugated through EDC/NHS chemistry. In the absence of model target ATP, the fluorescence of aptamer-modified UCNP was efficiently quenched with the binding between of GO. When adding the target, ATP-aptamer complex formed and thus decreased the exposure of DNA nucleobases, detaching ssDNA from GO surface and restoring the fluorescence signal (Fig. 1E). The detection limit was calculated to be 80nmol/L. Utilizing the π-π stacking interaction between DNA and materialsfull of sp2 carbonatoms, othercarbon materials, including carbon nanoparticles [28, 75], poly-mphenylenediamine nanospheres [72] and graphene quantum dots [76, 77] were also used as UCNP acceptors for various biomolecule detection.
3.2. Heterogeneous assayIn contrast to homogeneous assays, heterogeneous assays are performed in multiple steps with analytes being added, washed, and separated during the detection process. Although the procedures of heterogeneous assay are more complex than those of homogeneous assay, the generated signal is amplified with the concentration of UCNPs in solid-state platforms [1]. Meanwhile, the collection of DNA-functionalized UCNPs surpasses the instability of solution-phase reaction and constitutes the foundation of portable detection devices [49], which better satisfies the requirement of platform instrumentation.
Generally, for signal generation, either the oligonucleotide capture probe or UCNP is immobilized on solid platforms, including on strip [78, 79], cellulose paper [80, 81], coverslip [82-84], microplate [85-87], and membrane [88] in heterogeneous assays with DNA-functionalized UCNPs. For instance, Hao et al. transformed the LRET homogeneous assay to heterogeneous assay by immobilizing UCNPs on a nanoporous alumina membrane, which successfully decreased the LOD of virus oligo from pmol/L level to amol/L level (Fig. 2A) [88]. Natural enzymatic toolkit also provides possibility for further signal amplification of molecule detection with DNA-functionalized materials. Lan et al. combined exonuclease Ⅲ (ExoⅢ)-assisted amplification and long-range selfassembly DNA concatamers for the detection of c-erbB-2 oncogene [87]. The hairpin capture probe was firstly immobilized on a microplate. When adding target solution, the hybridization between target and capture probe broke the hairpin structure of capture probe and formed double-strand DNA sequences. As Exo Ⅲ only degraded double-strand DNA, one single-strand target was able to combine with numerous capture probes and thus amplified the signal. Subsequently two auxiliary probes, one was part complementary to capture probe and the other attached with UCNPs, were added for signal generation (Fig. 2B). LOD as low as 40amol/L was achieved. This design greatly utilized the programmability and structure diversity of DNA, showing great potential for ultrasensitive detection of biomolecules.

|
Download:
|
| Fig. 2. Scheme representations of heterogenous assayswith DNA-functionalized UCNPs. (A) Nanoporous alumina membrane-based assayfor virus oligonucleotide detection. Reproduced with permission [88]. Copyright 2016, American Chemical Society. (B) ExoⅢ-assisted DNA concatamers for c-erbB-2 oncogene detection. Reproduced with permission [87]. Copyright 2016, Scientific Reports. (C) Magnetic separation for target DNA detection. Reproduced with permission [89]. Copyright 2006, Royal Society of Chemistry. | |
Magnetic separation is another frequently employed method in heterogeneous assays with DNA-functionalized UCNPs. In this detection mode, oligonucleotide probe functions as both the recognition element and the linker between UCNPs and magnetic nanoparticles (MNPs). Different than LRET-based assays, since UCNPs are assembled with MNPs, there is no need of acceptor for signal transformation. The UCNP luminescence is directly related to the amount of targets, as the UCNP-target-MNP assemblycan be separated from carrier fluid through magnetic interaction. Li et al. firstly reported this detection mode for the quantification of nucleic acids [89]. Because both the capture probe (attached on MNPs) and the signal probe (attached on UCNPs) were complementary to adjacent regions of target DNA, the UCNP-target-MNP sandwiched assembly formed in the presence of targets (Fig. 2C). After separating and redispersing the collected assembly, fluorescent analysis was conducted and showed that the signal was proportional to the quantity of target DNA with a LOD of 10 nmol/L. This detection mode is subsequently developed with the employment of UCNPs with distinctive emissions for multiple detection [90, 91]. Its target scope is also expanded with the use of aptamers, including mycotoxins, antibiotics, viruses and cells [81, 91-94].
As stated above, DNA-functionalized UCNPs have been applied to the detection of various biomolecule with three main detection modes, including LRET-based detection, DNA concatamer and magnetic separation. For further developments, in homogeneous assay, focus should be placed on the invention of efficient acceptors and the optimization of donor-acceptor distance. Because of the complex excited states and multiple emission wavelengths of UCNPs, the acceptor choices are largely baffled. Generally, the quenching efficiency of organic dye was reported as 50% and that of AuNP was around of 70% [28, 37]. Carbon nanoparticles show better quenching efficiency but they improve the detection cost instead. Therefore, more efforts should be put into producing acceptors with broad absorption band and lower cost. Furthermore, tuning the donor-acceptor distance is also crucial, as the distance is the deciding factor of LRET efficiency. The structure variety and welldeveloped modification methods of DNA could contribute to the settlement of this problem, such as exploiting the DNA hairpin structure to shorten the donor-acceptor distance [37]. In heterogeneous assay, the relative ratio of capture probe and signal probe should be in careful control. For instance, in magnetic separation, the strongest signal was created when the amount of UCNP-DNA and MNP-DNA matched [61]. The higher concentration of MNPDNA would deepen the solution color and thus block the fluorescence [91]. Also, UCNPs with smaller sizes and fewer attached probes would prevent the UCNP from aggregation, resulting in better target-to-reporter ratios and assay kinetics [1]. In addition, more concentration should be placed on the instrumentation of detection devices, such as constructing portable biochip [83, 84], satisfying the requirement of commercialization and point-of-care detection.
4. DNA-functionalized UCNPs in bioimagingUCNP is well known as outstanding bioimaging nanomaterial for three reasons. Firstly, their NIR excitation property, which stands for deeper tissue penetration, negligible background noises and minimal tissue photodamage, presents unique advantages over other downconverstion fluorescent materials [4]. Secondly, UCNPs exhibit excellent photostability with neither photoblinking on second timescales nor phtobleaching under hours of continuous excitation [9]. Besides, both in vitro and in vivo experiments showed negligible toxicity of UCNP bioimaging probes [5, 10]. Although passive imaging is meaningful to investigate the feasibility of UCNP probes, active-targeted imaging is more practical for real disease monitoring [1]. Therefore, recognition elements should be introduced to upvonversion materials for targeted in vivo imaging. DNA, specifically aptamer, is a promising recognition element for this purpose. Aptamers that are aimed at specific proteins on disease cell surface, such as NFkB, HER3, VEGF, PDGF, and PMSA have already been produced. Moreover, with the development of cell-SELEX [95], aptamers that have affinity towards whole living cells, including various cancer cells, have been selected. These aptamers retain biological function in vivo and thus can be exploited as suitable recognition component in UCNP-based bioimaging. In addition, DNA can also function as a great linker in the integration of UCNP with other imaging nanoprobes, realizing multimodal imaging. Compared with solely luminescence imaging, multimodal imaging is able to provide more information and thus is more promising for future disease diagnosis. In the following section, we would specifically discuss recent progress in bioimaging with DNA-functionalized UCNPs, containing luminescence imaging and multimodal imaging.
4.1. Luminescence imagingAs discussed in Section 2, Lu's group developed a direct conjugation method of DNA-functionalized UCNPs and further applied them to cellular luminescence imaging [29]. The produced DNA-UCNPs were able to cross cell membranes without the aid of transfection agents, making them excellent probes for bioimaging. Specifically, a DNA aptamer AS1411, which is able to target nucleolin, was attached to UCNPs for targeted imaging of the nucleolin-overexpressed breast cancer cell line MCF-7 (Fig. 3A). Overlays of bright-field images and confocal fluorescence proved the successful accumulation of aptamer-UCNPs in MCF-7 cells. In control experiments, these aptamer-UCNPs showed far less affinity to normal cells, which demonstrated their excellent recognition ability. Besides fluorescence imaging, aptamer-functionalized UCNPs have also been employed for in vivo detection through LRET mechanism. For instance, Wang et al. built an UCNP@PDAbased aptasensor for intracellular sensing of cytochrome c (cyt c) [38]. Aptamer-complementary DNA probes (cDNA) were firstly conjugated on UCNP@PDA. Then Cy3 fluorophore-labeled aptamer would combine with cDNA-UCNP through base-pairing. The PDA shell was introduced to quench the fluorescence of fluorophore Cy3 and protect DNA absorbed on its surface from nuclease digestion. When uptaken by targeted cells, aptamer would combine with cyt c and dissociate from UCNP surface, restoring the Cy3 fluorescence (Fig. 3B). Successful imaging and quantification of cyt c was achieved in HepG-2 cells with Cy3 fluorescence alteration, and the consistent UCNP fluorescence was used as the reference signal for self-calibrating. This imaging mode provides a general platform for intracellular target quantification with aptamer-modified UCNPs.

|
Download:
|
| Fig. 3. Scheme representations of bioimaging with DNA-functionalized UCNPs. (A) Aptamer-modified UCNPs for targeted luminescence imaging. Reproduced with permission [29]. Copyright 2013, American Chemical Society. (B) UCNP@PDA (polydopamine) aptasensor for intracellular sensing of cytochrome c. Reproduced with permission [38]. Copyright 2017, Elsevier. (C) DNA-AuNP/UCNP assembly for multimodal imaging. Reproduced with permission [97]. Copyright 2015, American Chemical Society. | |
4.2. Multimodal imaging
Asides from aforementioned recognition element, DNA also functions as a linker between UCNPs and other imaging probes, which provides great potential for multimodal imaging [96]. For instance, Lu's group designed an aptamer-functionalized Au-UCNP assembly for intracellular dual-imaging [97]. This assembly was built through the electrostatic interaction between negatively charged DNA-AuNP and hexagonal phospholipids-capped UCNPs (Lipo-UCNP). Interestingly, with the coating of Lipo shell on UCNPs, minimal AuNPs-induced luminescence quenching was observed in these DNA-AuNP/UCNP structures, showing its potential as dualmode imaging agent. When uptaken by breast cancer cells, this hetero-assembly demonstrated both strong upconversion fluorescence and light scattering signals (Fig. 3C). These two kinds of signals were distinctive and can be used for analysis of different properties.
Noteworthy, UCNPs per se are also good candidates for multimodal imaging. Because lanthanide elements have higher atomic numbers and K-edge values within the X-ray spectrum, UCNPs can create strong X-ray attenuation for computed tomography (CT). Moreover, Gd3+ doped UCNPs are also excellent agents for T1-weighted magnetic resonance imaging (MRI) [9, 39]. Therefore, UCNP-involved imaging categories can be further enlarged through the material assembly with DNA linkers. For instance, by combing UCNP with photothermal materials [98, 99], high-resolution photoacoustic tomography (PA) could be obtained with this contrast agent, providing both depth and spatial information of tissues. Xu et al. [100] further assembled Gd3+-doped UCNPs with gold nanorods through DNA hybridization, obtaining successful multimodal imaging in tumorbearing mice experiments, including luminescence imaging, MRI, CT and PA. With the provided various information, an exhaustive and accurate analysis towards the tumor morphology could be created.
In conclusion, DNA endows upconversion material with targeted-imaging ability and multimodal imaging capacity. Despite these aforementioned progresses, bioimaging with DNAfunctionalized UCNPs is still in its infancy. Two problems might impede the real application of these materials. The first one is the nuclease digestion problem of DNA for in vivo analysis. This problem can be further solved by introducing protective shells on UCNPs [30, 38] or building rigid DNA-UCNP nanostructures with DNA nanotechnology [62]. The second problem is the seemingly redundancy of material assembly between UCNP and other probes for multimode imaging, as the direct seed-mediate growth [101] or layer-by-layer synthesis [102] of multifunctional UCNPs imaging probes (such as UCNP@Fe3O4@Au) is apparently more facile in material synthesis. However, the spatial separation of imaging modalities can reduce signal interference [1], offering extra significance to the linking between disparate imaging probes. Moreover, the modality separation would provide enough conjugation space for further functionalization of each probe, which may bring extra therapeutic effect or detection function to the imaging system. Therefore, DNA-mediated UCNP material assembly is promising in bioimaging and more efforts should be made for feasibility test.
5. DNA-functionalized UCNPs in disease therapyUCNP-involved disease therapies, including molecule delivery and phototherapy, demonstrate obvious advantages in real application due to the unique optical properties of UCNPs. On the one hand, the stable and high-sensitivity luminescence signal can be used to track the molecule propagating channels, which is meaningful in medicine research [10]. On the other hand, NIR light-controlled molecule release and phototherapy activation can be achieved with UCNPs [15]. Although most of lightresponsive materials only respond to UV or visible light, UCNPs that possess upconversion property can function as excellent nanotranducers in this system, allowing for NIR-activated therapy with deep tissue penetration and low specimen damage [10, 15, 103]. However, inert UCNPs do not have molecule-loading capacity. They also cannot automatically transfer their emission to heat or singlet oxygen for phototherapy. Therefore, it is important to impart UCNP with therapeutic effect through structure regulation and material assembly.
DNA can be good candidate for these endeavors for three reasons. Firstly, specific nucleic acids, such as plasmid DNA, siRNA and antisense oligonucleotides, are therapeutic agents themselves, as they have transfection ability and can be applied to gene-related diseases [25]. Secondly, DNA with double-helix framework is capable of encapsulating small drug molecules, such as anticancer drug doxorubicin (DOX), through sequenceselective binding, which makes it a great drug carrier [26, 104]. As for phototherapy, DNA could be linkers in material assembly to endow UCNP-involved system with both photothermal and photodynamic effect. Therefore, it is beneficial to introduce DNA in upconversion material for building multifunctional therapeutic platforms. The following section reviews a set of NIR-activated smart delivery systems and advanced therapeutic applications with the use of DNA-functionalized UCNPs, including gene therapy, drug delivery and phototherapy.
5.1. Gene therapyGene therapy, the treatment of targeted cells with therapeutic nucleic acids to regulate abnormal gene expression, has been widely applied to gene-related diseases and cancer [47]. As aforementioned, UCNPs are excellent cargoes of nucleic acids for both imaging tracking and gene release control. The pioneering investigation was completed by Zhang's group [105]. In this study, amino-modified UCNPs were firstly conjugated with antiHer2 antibody for targeted delivery, and then attached with siRNAs through electrostatic interaction. Material uptake by Her2-overexpressed SK-BR-3 cells was proved successful under fluorescence monitoring, and 45% down-regulation of luciferase gene expression was achieved. However, the tracking of gene release from nanoplatform was not realized in this system [47]. To address the problem, the same group employed LRET mechanism in material design [43]. By introducing intercalating dye BOBO-3 in siRNA as acceptors, gene release can be visualized through fluorescence alteration and quantified by LRET efficiency calculation. Afterwards, UCNP surface modifications were further applied to this delivery system on geneloading efficiency improvement and targeted delivery enhancement [45 46, 106].
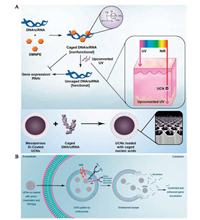
Despite these developments on molecule imaging tracking, gene delivery in a controlled release manner had not been accomplished. To achieve this function, introducing photoactive material to "cage" the transfection genes is a practical solution, as the photoactiver would be cleaved when absorbing UV light. Zhang's group further designed a photocaged gene loading system using NIR-to-UV UCNPs to realize NIR-activated releasing [30]. Plasmid DNA or siRNA was firstly bond with photoactive compound 4, 5-dimethoxy-2-nitroacetophenone (DMNPE) through chemical modification, and then loaded onto mesoporous silica-coated UCNPs by physical absorption. Under NIR excitation, DMNPE would absorb the UV emission from UCNP and thus be cleaved, initiating the cellular release of delivered genes and realizing light-activated release control (Fig. 4A). Tissue phantom experiments proved successful activation of genes under 0.4 cm tissue thickness, demonstrating the great therapy potential of this gene-delivering material.
|
Download:
|
| Fig. 4. Scheme representations of gene therapy with DNA-functionalized UCNPs. (A) Photo-controlled gene delivery with NIR-to-UV UCNPs. Reproduced with permission [30]. Copyright 2012, National Academy of Sciences. (B) Gene knockdown with UCNP-based photoactive delivery platform. Reproduced with permission [108]. Copyright 2014, American Chemical Society. | |
When microinjected into biological system, one important obstacle of nanocarrier-based gene delivery is the sequestration of endosomal vesicles, which may lead to material breaking down or exocytosing out from cells [107]. To further promote the endosomal escape of delivered genes and thus improve the therapeutic effect, Zhang's group employed photochemical internalization (PCI) method in gene release control. UCNPs were co-loaded with TPPS2a (a PCI photosentizer) and photomorpholino (anti-STAT3, nucleic analogous for gene knockdown) [108]. Under NIR excitation, visible light emission (413 nm) from UCNPs was absorbed by TPPS2a and lead to the production of ROS, which locally disrupted the endosomal vesicle walls and thus released the endosome contents into the cytoplasm. Meanwhile, UV light emission from UCNPs cleaved the sense photomorpholino and freed the antisense morpholino, resulting in gene knockdown of STAT3 (Fig. 4B). A 30% improvement of gene knockdown was achieved compared with control group without endosomal escape facilitation, showing its excellent clinical potential.
5.2. Drug deliveryBesides gene therapy, UCNPs also have great potential in drug delivery for both imaging-guided investigation and light-activated release control. However, natural UCNPs do not have molecule loading capacity. Although introducing chemical modifications to UCNPs, such as coating with amphiphilic polymer [109] or mesoporous silica shells [110], are useful in drug loading, NIRrelease control and targeted delivery are rather difficult to be both achieved in these systems. Therefore, it is of great value to combine UCNPs with functional molecule that simultaneously possesses drug loading capacity, drug release control potential and selectivity. DNA can be excellent candidate for these endeavors. Hairpinshaped DNA is able to encapsulate small drug molecules through selective-sequence binding. It can also undergo conformational change in high temperature or in the presence of complementary strands, leading to the control of drug delivery and release. These properties make DNA promising functional molecules in UCNPbased drug delivery.
Recently, Liu et al. designed a hairpin DNA (hpDNA) functionalized UCNP system for DOX delivery [111]. Silica coated UCNPs were attached with small-sized gold nanoparticles (2 nm), which functioned as the photothermal converter for DNA unwinding. Because AuNPs were able to selectively absorb the UV-vis emission from UCNPs, the surface temperature of Au gradually increased with NIR excitation. Meanwhile, NIR emission of UCNP (800 nm) was not influenced by the introduction of AuNPs and thus could be used as imaging signal, owning additional advantage in terms of tissue penetration. Next, hpDNA strands were immobilized on AuNPs by Au-S chemistry and loaded with DOX molecules. When irradiated by NIR light, the temperature increase brought by AuNP's localized photothermal effect led to the unwinding of hpDNAs, realizing light-activated DOX release control (Fig. 5A). Successful drug unleashing under 980 nm excitation was achieved in both cell viability experiments and tumor-bearing mice experiments. This system provides a new modality for both deep-tissue imaging and drug controlling release. Besides temperature-induced unwinding, hpDNA undergoes conformation change when hybridizing with complementary strands and thus realize targeted delivery. Utilizing this property, Xing et al. proposed a novel upconversion nanobeacon for tumor therapy [112]. In this molecule beacon, UCNP was applied as donor and BHQ-1, the quencher labeled on hairpin DNA, was used as acceptor. Then drug DOX was loaded on hpDNAs for therapy tests. Normally the green emission from UCNP was quenched by BHQ-1 through LRET mechanism. Because the sequence of hpDNA was designed complementary to target thymidine kinase 1 (TK1) mRNA, hpDNA would hybridize with target upon target binding, leading to the drug detachment and fluorescence restoration (Fig. 5B). Hence, both the quantification of target mRNA and the release of drugs can be monitored by UCNP emission. Strong selectivity and sensitivity of this system was achieved in living cell (TK1 mRNA overexpressed MCF-7 cells) experiments. By combing the drug loading capacity and selectivity of DNA, this novel agent realized mRNA detection, targeted drug delivery and drug release control, making it a promising theranostic platform for tumor treatment.

|
Download:
|
| Fig. 5. Scheme representations of drug delivery with DNA-functionalized UCNPs. (A) Temperature-induced drug release with hairpin DNA-Au/UCNP nanoconjugates. Reproduced with permission [111]. Copyright 2017, John Wiley and Sons. (B) Targeted drug delivery and target mRNA quantification with UCNP-based molecule beacons. Reproduced with permission [112]. Copyright 2016, John Wiley and Sons. | |
In addition with chemical modifications, another strategy for drug loading is to use hollow UCNPs with mesoporous shell [1]. Usually the formation of hollow structure was fabricated with sacrificial templates such as colloidal spheres [113] and magnetic nanoparticles [114], which is complex and sometimes environment-unfriendly. Recently, Qu et al. reported a facile and green synthesis method of hollow UCNP using DNA as the structure mediator [115]. They found that DNA could function as morphology transformer to promote the hollow structure formation of UCNPs employing hydrothermal synthesis method. With the increase of DNA concentration, UCNPs were transformed from flower-like nanoaggregates to small-sized spheres. Interestingly, when adjusting the DNA concentration to a suitable level, sphere particles with a hollow interior would occur, attributed to the influence of DNA on particle nucleation and growth. These DNA-mediated hollow UCNPs have great potential for drug loading due to its large interior space. More efforts should be devoted to this area for further applications.
5.3. PhototherapyPhototherapy, including photothermal therapy (PTT) and photodynamic therapy (PDT), is well-known for its minimal invasiveness, remote controllability and high therapeutic effect. In PTT, photoabsorbers are usually employed to transfer absorbed light to heat, resulting in thermal ablation of cells [104, 116]. In PDT, photosentisizers are exploited to generate singlet oxygen (1O2), leading to the death of targeted cells [117, 118]. UCNPs have been widely applied in phototherapy due to their upconversion property, which allows for NIR-triggered phototherapy with high tissue penetration and low photodamage [119]. However, to achieve expected therapy effect, high density irradiation is always required in solely PTTor PDT, which might cause damage to healthy cells if over the skin maximum permissible exposure (MPE). Therefore, the combination of PTT and PDT has become a trend to increase the therapeutic efficacy and minimize the side effects. DNA, as a linker with high programmability, as well as a recognition sequence with high selectivity, is thus applied in the construction of UCNP-involved multifunctional nanoplatforms to achieve combination therapy.
Recently, Xu et al. designed a plasmonic NR dimer-UCNP-Ce6 (Chlorin e6) system for PTT and PDT synergistic remedy [94]. Specifically, gold nanorod (NR) was chosen as PTT photoabsorber and Ce6 was chosen as PDT photosentisizer. NR dimers were built through DNA hybridization, which exhibited better photothermal effect than NR monomers. PEG coated UCNPs (NaGdF4:Yb, Er) were firstly conjugated with Ce6 and then assembled with NR dimers through DNA hybridization (Fig. 6A). When tail-injected to tumorbearing mice, this assembly showed both PDT (980 nm, 2 mW/cm2, 30 min, lower than MPE) and PTT (808 nm, 0.2 W/cm2, 5 min, lower than MPE) therapeutic effects, and completely eliminated the tumor after 15-day irradiation. In contrast, mice treated with solely PDT or PTT had limited tissue ablation during this period. Multimodal imaging was also achieved with the use of Gd3+- doped UCNPs and NPs. This system exhibited both multimodal imaging ability and successful elimination with tumor under safe power dose, which is promising for future clinical cancer treatment.

|
Download:
|
| Fig. 6. Scheme representations of phototherapy with DNA-functionalized UCNPs. (A) Gold nanorod-UCNP-Ce6 assembly for phototherapy and multimodal imaging. Reproduced with permission [100]. Copyright 2015, John Wiley and Sons. (B) UCNP-methylene blue-hairpin DNA nanoconstructs for imaging-guided phototherapy and gene detection. Reproduced with permission [39]. Copyright 2017, American Chemical Society. | |
Despite aforementioned advantages, the material delivery selectivity of the above system relied on enhanced permeability and retention effect, which might not be ideal in elaborate medical treatments. Chu et al. developed a new platform exploiting the target recognition ability of DNA to discriminate cancer cells from normal cells [91]. Specifically, UCNPs were firstly coated with silica shell loading with photosentisizer methylene blue (MB). Then the PDA shell was further modified on UCNPs, which could function as both PTT photoabsorber and the ssDNA linker due to its rich catechol and amine groups. Two different hairpin DNA probes, one targeted c-myc mRNA with FAM fluorophore labelling and the other aimed at TK1 mRNA with Cy3 fluorophore labelling, were then conjugated to PDA shell. Both of the signals from these two fluorophores were quenched by PDA through energy/electron transfer. When uptaken by MCF-7 breast cancer cells, the overexpression of target genes leaded to the detachment of hairpin DNAs from PDA surface, restoring the fluorophore fluorescence (Fig. 6B). Successful imaging of both two channels was achieved. As for phototherapy, under 980 nm excitation, the 605 nm emission of UCNPs was absorbed by MB and thus caused the generation of 1O2; the 800 nm emission of UCNPs was absorbed by PDA, producing PTT effect. The PTT-PDT synergistic therapy induced efficient cancer cell apoptosis in cell viability test. One remarkable feature of this system is that in real application, the fluorescence restoration could be a signal for therapy initiation as it stands for the successful locating of cancer cells, making it an imaging-guided multifunctional platform.
As stated above, DNA-functionalized UCNPs platforms provide great advances for spatially and temporally controlled molecule release (e.g., nucleic acids, drugs) and phototherapy activation. With the employment of different properties of DNA, the platform function has been gradually extended, started from imaging tracking to multi-therapy combination. For further development, more efficient NIR-to-UV UCNPs should be developed to improve the release control efficiency and therapeutic effect [120]. Meanwhile, to circumvent the heat effect caused by the 980 nm absorption of water, it is desirable to develop novel UCNPs with an excitation wavelength between 700 nm and 900 nm (i.e., where water's absorption is negligible). Moreover, most of the present works are limited to in vitro experiments. The in vivo therapeutic effect and toxicity investigation of DNA-functionalized UCNPs is important for their clinical translation. More systematic and rigorous evaluations of their biosafety and degradability are still required before their application in clinics.
6. ConclusionIn this review, we concluded recent developments in material design, nanochemistry, and therapeutic applications of DNAfunctionalized UCNPs. In particular, they could be used to construct effective luminescent biosensors for biomolecule detection, sensitive nanoprobes for optical and multimodal imaging, and multifunctional platforms for disease therapy. To further promote their biomedical applications, we conclude existed issues from three aspects, aiming to offer insights for their further developments.
Firstly, in regard to UCNPs, fluorescent intensity enhancement is crucial for efficiency improvement. Despite the discussed advantages of UCNPs, the quantum yield of these materials is usually less than 1% [6]. Size control, host matrix optimization and core-shell structure construction should be further developed to address this issue. For instance, Wang et al. reported an effective core-shell UCNP system with CaF2 as shell material, which generated around 300 times emission amplification [121]. This study provides a direction for further emission enhancement. In addition, the invention of UCNPs with high emission intensity and small size below 10 nm is promising for clinical devices, as nanoparticles in this size can be efficiently cleared from the body [122]. However, smaller sized UCNPs generally possess lower emission intensity. To overcome this limitation, Hao et al. synthesized KGdF4:Tm3+, Yb3+ UCNPs with average sizes of 7.4 nm and comparable emission intensity with traditional UCNPs, attributing to the doping of Gd3+. More efforts should be devoted into this field to promote their clinical applications.
Secondly, the selection of functional DNA towards more effective disease targets is also important for selectivity improvement of DNA-functionalized nanomaterials. This should be accompanied with the finding of reliable molecular and cellular markers of specific diseases [2] and the development of SELEX approaches. Moreover, more attention should be paid on the introduction of DNA nanotechnology to UCNP-based nanostructures. Currently, most nanomaterials are built through linear assembly of two complementary DNA strands [17]. Twodimensional and three-dimensional nanostructures, such as the introduction of branched, dendritic and networked DNA to UCNPbased nanostructures, are promising for multiple function integration and deserves further exploration.
Thirdly, for future developments, constructing an all-purpose theranostic platform, which integrates modalities of intracellular biomolecule detection, multimodal imaging, controlled drug release and phototherapy, is a promising research direction. With the improvement of UCNP emission properties, the development of DNA technologies and the advancement of surface modification methods, this could be achieved in near future.
AcknowledgmentsThis work was supported by the National Key R & D Program of China (No. 2017YFA0208000), the National Natural Science Foundation of China (Nos. 21422105, 21675120), the Natural Science Foundation of Hubei Province (No. 2015CFA032), and Ten Thousand Talents Program for Young Talents.
| [1] |
G. Chen, H. Qiu, P.N. Prasad, et al., Chem. Rev. 114 (2014) 5161-5214. DOI:10.1021/cr400425h |
| [2] |
S.F. Torabi, Y. Lu, Curr. Opin. Biotechnol. 28 (2014) 88-95. DOI:10.1016/j.copbio.2013.12.011 |
| [3] |
A. Sedlmeier, H.H. Gorris, Chem. Soc. Rev. 44 (2015) 1526-1560. DOI:10.1039/C4CS00186A |
| [4] |
M. Wang, G. Abbineni, A. Clevenger, et al., Nanomedicine 7 (2011) 710-729. DOI:10.1016/j.nano.2011.02.013 |
| [5] |
L. Cheng, C. Wang, Z. Liu, Nanoscale 5 (2013) 23-37. DOI:10.1039/C2NR32311G |
| [6] |
Z. Gu, L. Yan, G. Tian, et al., Adv. Mater. 25 (2013) 3758-3779. DOI:10.1002/adma.v25.28 |
| [7] |
G. Chen, H. Agren, T.Y. Ohulchanskyy, et al., Chem. Soc. Rev. 44 (2015) 1680-1713. DOI:10.1039/C4CS00170B |
| [8] |
D. Yang, P. Ma, Z. Hou, et al., Chem. Soc. Rev. 44 (2015) 1416-1448. DOI:10.1039/C4CS00155A |
| [9] |
Y.I. Park, K.T. Lee, Y.D. Suh, et al., Chem. Soc. Rev. 44 (2015) 1302-1317. DOI:10.1039/C4CS00173G |
| [10] |
J. Shen, L. Zhao, G. Han, Adv. Drug Deliv. Rev. 65 (2013) 744-755. DOI:10.1016/j.addr.2012.05.007 |
| [11] |
M.V. DaCosta, S. Doughan, Y. Han, et al., Anal. Chim. Acta 832 (2014) 1-33. DOI:10.1016/j.aca.2014.04.030 |
| [12] |
L.D. Sun, Y.F. Wang, C.H. Yan, Acc. Chem. Res. 47 (2014) 1001-1009. DOI:10.1021/ar400218t |
| [13] |
L.J. Huang, R.Q. Yu, X. Chu, Analyst 140 (2015) 4987-4990. DOI:10.1039/C5AN00635J |
| [14] |
C. Wang, X. Li, F. Zhang, Analyst 141 (2016) 3601-3620. DOI:10.1039/C6AN00150E |
| [15] |
C. Wang, L. Cheng, Z. Liu, Theranostics 3 (2013) 317-330. DOI:10.7150/thno.5284 |
| [16] |
U. Feldkamp, C.M. Niemeyer, Angew. Chem. Int. Ed. 45 (2006) 1856-1876. DOI:10.1002/chin.200624280 |
| [17] |
D. Yang, M.R. Hartman, T.L. Derrien, et al., Acc. Chem. Res. 47 (2014) 1902-1911. DOI:10.1021/ar5001082 |
| [18] |
Y.H. Roh, R.C. Ruiz, S. Peng, et al., Chem. Soc. Rev. 40 (2011) 5730-5744. DOI:10.1039/c1cs15162b |
| [19] |
K.V. Gothelf, T.H. LaBean, Org. Biomol. Chem. 3 (2005) 4023-4037. DOI:10.1039/b510551j |
| [20] |
L.H. Tan, H. Xing, Y. Lu, Acc. Chem. Res. 47 (2014) 1881-1890. DOI:10.1021/ar500081k |
| [21] |
A.D. Keefe, S. Pai, A. Ellington, Nat. Rev. Drug. Discov. 9 (2010) 537-550. DOI:10.1038/nrd3141 |
| [22] |
J. Liu, Z. Cao, Y. Lu, Chem. Rev. 109 (2009) 1948-1998. DOI:10.1021/cr030183i |
| [23] |
T. Hermann, D.J. Patel, Science 287 (2000) 820-825. DOI:10.1126/science.287.5454.820 |
| [24] |
S.D. Jayasena, Clin. Chem. 45 (1999) 1628-1650. |
| [25] |
L. Naldini, Nature 526 (2015) 351-360. DOI:10.1038/nature15818 |
| [26] |
A. Khvorova, J.K. Watts, Nat. Biotechnol. 35 (2017) 238-248. DOI:10.1038/nbt.3765 |
| [27] |
M. Kumar, P. Zhang, Biosens. Bioelectron. 25 (2010) 2431-2435. DOI:10.1016/j.bios.2010.03.038 |
| [28] |
Y. Wang, L. Bao, Z. Liu, et al., Anal. Chem. 83 (2011) 8130-8137. DOI:10.1021/ac201631b |
| [29] |
L.L. Li, P. Wu, K. Hwang, et al., J. Am. Chem. Soc. 135 (2013) 2411-2414. DOI:10.1021/ja310432u |
| [30] |
M.K. Jayakumar, N.M. Idris, Y. Zhang, Proc. Nat. Acad. Sci. U. S. A. 109 (2012) 8483-8488. DOI:10.1073/pnas.1114551109 |
| [31] |
M. Kumar, P. Zhang, Langmuir 25 (2009) 6024-6027. DOI:10.1021/la900936p |
| [32] |
P. Wang, P. Zhang, RSC Adv. 4 (2014) 56235-56240. DOI:10.1039/C4RA10181B |
| [33] |
S. Wu, N. Duan, X. Ma, et al., Anal. Chim. Acta 782 (2013) 59-66. DOI:10.1016/j.aca.2013.04.025 |
| [34] |
H.Q. Chen, F. Yuan, S.Z. Wang, et al., Analyst 138 (2013) 2392-2397. DOI:10.1039/c3an36921h |
| [35] |
H. Chen, F. Yuan, S. Wang, et al., Biosens. Bioelectron. 48 (2013) 19-25. DOI:10.1016/j.bios.2013.03.083 |
| [36] |
C. Liu, Z. Wang, H. Jia, et al., Chem. Commun. 47 (2011) 4661-4663. DOI:10.1039/c1cc10597c |
| [37] |
Y. Yuan, Z. Liu, Chem. Commun. 48 (2012) 7510-7512. DOI:10.1039/c2cc33289b |
| [38] |
L. Ma, F. Liu, Z. Lei, et al., Biosens. Bioelectron. 87 (2017) 638-645. DOI:10.1016/j.bios.2016.09.017 |
| [39] |
Y. Cen, W.J. Deng, Y. Yang, et al., Anal. Chem. 89 (2017) 10321-10328. DOI:10.1021/acs.analchem.7b02081 |
| [40] |
H. Zhang, C. Fang, S. Wu, et al., Anal. Biochem. 489 (2015) 44-49. DOI:10.1016/j.ab.2015.08.011 |
| [41] |
W.W. Ye, M.K. Tsang, X. Liu, et al., Small 10 (2014) 2390-2397. DOI:10.1002/smll.v10.12 |
| [42] |
L. Huang, X. Tian, J. Yi, et al., Anal. Methods 7 (2015) 7474-7479. DOI:10.1039/C5AY01169H |
| [43] |
S. Jiang, Y. Zhang, Langmuir 26 (2010) 6689-6694. DOI:10.1021/la904011q |
| [44] |
X. Li, J. Zhang, H. Gu, Langmuir 28 (2012) 2827-2834. DOI:10.1021/la204443j |
| [45] |
M. Lin, Y. Gao, T.J. Diefenbach, et al., ACS Appl. Mater. Interfaces 9 (2017) 7941-7949. DOI:10.1021/acsami.6b15321 |
| [46] |
L. He, L. Feng, L. Cheng, et al., ACSAppl.Mater.Interfaces 5 (2013) 10381-10388. DOI:10.1021/am403554x |
| [47] |
M. Lin, Y. Gao, F. Hornicek, et al., Adv. Colloid Interface Sci. 226 (2015) 123-137. DOI:10.1016/j.cis.2015.10.003 |
| [48] |
Y. Yang, F. Liu, X. Liu, et al., Nanoscale 5 (2013) 231-238. DOI:10.1039/C2NR32835F |
| [49] |
D. Mendez-Gonzalez, E. Lopez-Cabarcos, J. Rubio-Retama, et al., Adv. Colloid Interface Sci. 249 (2017) 66-87. DOI:10.1016/j.cis.2017.06.003 |
| [50] |
H. Zhang, F. Li, B. Dever, et al., Chem. Rev. 113 (2013) 2812-2841. DOI:10.1021/cr300340p |
| [51] |
A. Sassolas, B.D. Leca-Bouvier, L.J. Blum, Chem. Rev. 108 (2008) 109-139. DOI:10.1021/cr0684467 |
| [52] |
C. Feng, S. Dai, L. Wang, Biosens. Bioelectron. 59 (2014) 64-74. DOI:10.1016/j.bios.2014.03.014 |
| [53] |
X.M. Fu, Z.J. Liu, S.X. Cai, et al., Chin. Chem. Lett. 27 (2016) 920-926. DOI:10.1016/j.cclet.2016.04.014 |
| [54] |
L. Wang, R. Yan, Z. Huo, et al., Angew. Chem. Int. Ed. 44 (2005) 6054-6057. |
| [55] |
K.E. Sapsford, L. Berti, I.L. Medintz, Angew. Chem. Int. Ed. 45 (2006) 4562-4589. DOI:10.1002/anie.200503873 |
| [56] |
A.R. Clapp, I.L. Medintz, J.M. Mauro, J. Am. Chem. Soc. 126 (2004) 301-310. DOI:10.1021/ja037088b |
| [57] |
Y.Y. Chen, T.T. Zhang, X.N. Gao, et al., Chin. Chem. Lett. 28 (2017) 1983-1986. DOI:10.1016/j.cclet.2017.08.024 |
| [58] |
P. Zhang, S. Rogelj, K. Nguyen, J. Am. Chem. Soc. 128 (2006) 12410-12411. DOI:10.1021/ja0644024 |
| [59] |
S. Kim, S.H. Hwang, S.G. Im, et al., Sensors 16 (2016) 1259. DOI:10.3390/s16081259 |
| [60] |
T. Rantanen, M.L. Jarvenpaa, J. Vuojola, et al., Analyst 134 (2009) 1713-1716. DOI:10.1039/b901299k |
| [61] |
S. Wu, N. Duan, H. Zhang, Anal. Bioanal. Chem. 407 (2015) 1303-1312. DOI:10.1007/s00216-014-8378-3 |
| [62] |
S. Li, L. Xu, W. Ma, et al., J. Am. Chem. Soc. 138 (2016) 306-312. DOI:10.1021/jacs.5b10309 |
| [63] |
J.G. Jesu Raj, M. Quintanilla, K.A. Mahmoud, et al., ACS Appl. Mater. Interfaces 7 (2015) 18257-18265. DOI:10.1021/acsami.5b02986 |
| [64] |
M. Wu, X. Wang, K. Wang, et al., Chem. Commun. 52 (2016) 8377-8380. DOI:10.1039/C6CC02674E |
| [65] |
T. Hao, X. Wu, L. Xu, et al., Small 13 (2017) 1603944. DOI:10.1002/smll.v13.19 |
| [66] |
S. Wu, N. Duan, Z. Shi, et al., Talanta 128 (2014) 327-336. DOI:10.1016/j.talanta.2014.04.056 |
| [67] |
A. Qu, X. Wu, L. Xu, et al., Nanoscale 9 (2017) 3865-3872. DOI:10.1039/C6NR09114H |
| [68] |
F. Lin, B. Yin, C. li, et al., Anal. Methods 5 (2013) 699-704. DOI:10.1039/C2AY25519G |
| [69] |
B. Jin, S. Wang, M. Lin, et al., Biosens. Bioelectron. 90 (2017) 525-533. DOI:10.1016/j.bios.2016.10.029 |
| [70] |
S. Li, L. Xu, M. Sun, et al., Adv. Mater. 29 (2017) 1606086. DOI:10.1002/adma.v29.19 |
| [71] |
C. Lu, H. Yang, C. Zhu, Angew. Chem. 121 (2009) 4879-4881. DOI:10.1002/ange.v121:26 |
| [72] |
Y. Wang, Z. Wu, Z. Liu, Anal. Chem. 85 (2013) 258-264. DOI:10.1021/ac302659b |
| [73] |
G.M. Han, H. Li, X.X. Huang, Talanta 147 (2016) 207-212. DOI:10.1016/j.talanta.2015.09.059 |
| [74] |
Y.M. Wu, Y. Cen, L.J. Huang, et al., Chem. Commun. 50 (2014) 4759-4762. DOI:10.1039/C4CC00569D |
| [75] |
Z. Wu, H. Li, Z. Liu, Sens. Actuators B:Chem. 206 (2015) 531-537. DOI:10.1016/j.snb.2014.09.084 |
| [76] |
M. Laurenti, M. Paez-Perez, M. Algarra, et al., ACS Appl. Mater. Interfaces 8 (2016) 12644-12651. DOI:10.1021/acsami.6b02361 |
| [77] |
L. He, L. Yang, H. Zhu, Methods Appl. Fluoresc. 5 (2017) 024010. DOI:10.1088/2050-6120/aa6e0d |
| [78] |
P.L.A.M. Corstjens, M. Zuiderwijk, M. Nilsson, et al., Anal. Biochem. 312 (2003) 191-200. DOI:10.1016/S0003-2697(02)00505-5 |
| [79] |
M. Zuiderwijk, H.J. Tanke, R. Sam Niedbala, et al., Clin. Biochem. 36 (2003) 401-403. DOI:10.1016/S0009-9120(03)00057-2 |
| [80] |
F. Zhou, M.O. Noor, U.J. Krull, Anal. Chem. 86 (2014) 2719-2726. DOI:10.1021/ac404129t |
| [81] |
S. Wu, H. Zhang, Z. Shi, et al., Food Control 50 (2015) 597-604. DOI:10.1016/j.foodcont.2014.10.003 |
| [82] |
S. Doughan, Y. Han, U. Uddayasankar, et al., ACS Appl. Mater. Interfaces 6 (2014) 14061-14068. DOI:10.1021/am503391m |
| [83] |
X. Hu, Y. Wang, H. Liu, et al., Chem. Sci. 8 (2017) 466-472. DOI:10.1039/C6SC03401B |
| [84] |
Y. Tan, X. Hu, M. Liu, et al., Chem. Eur. J. 23 (2017) 10683-10689. DOI:10.1002/chem.v23.44 |
| [85] |
J. Lan, F. Wen, F. Fu, et al., RSC Adv. 5 (2015) 18008-18012. DOI:10.1039/C5RA01288K |
| [86] |
J. Lan, L. Li, Y. Liu, et al., Microchimica Acta 183 (2016) 3201-3208. DOI:10.1007/s00604-016-1965-6 |
| [87] |
J. Lan, Y. Liu, L. Li, et al., Sci. Rep. 6 (2016) 24813. DOI:10.1038/srep24813 |
| [88] |
M.K. Tsang, W. Ye, G. Wang, et al., ACS Nano 10 (2016) 598-605. DOI:10.1021/acsnano.5b05622 |
| [89] |
L. Wang, Y. Li, Chem. Commun. (2006) 2557-2559. |
| [90] |
S. Wu, N. Duan, X. Ma, et al., Chem. Commun. 48 (2012) 4866-4868. DOI:10.1039/c2cc00092j |
| [91] |
N. Duan, S. Wu, C. Zhu, et al., Anal. Chim. Acta 723 (2012) 1-6. DOI:10.1016/j.aca.2012.02.011 |
| [92] |
S. Wu, N. Duan, Z. Shi, et al., Anal. Chem. 86 (2014) 3100-3107. DOI:10.1021/ac404205c |
| [93] |
S. Wu, N. Duan, Z. Wang, et al., Analyst 136 (2011) 2306-2314. DOI:10.1039/c0an00735h |
| [94] |
S. Fang, C. Wang, J. Xiang, et al., Nano Res. 7 (2014) 1327-1336. DOI:10.1007/s12274-014-0497-9 |
| [95] |
Z.X. Huang, Q. Xie, Q.P. Guo, et al., Chin. Chem. Lett. 28 (2017) 1252-1257. DOI:10.1016/j.cclet.2017.01.002 |
| [96] |
Y. Chen, C.L. Tan, H. Zhang, et al., Chem. Soc. Rev. 44 (2015) 2681-2701. DOI:10.1039/C4CS00300D |
| [97] |
L.L. Li, Y. Lu, J. Am. Chem. Soc. 137 (2015) 5272-5275. DOI:10.1021/jacs.5b01092 |
| [98] |
S.K. Maji, S. Sreejith, J. Joseph, et al., Adv. Mater. 26 (2014) 5633-5638. DOI:10.1002/adma.201400831 |
| [99] |
Y.S. Chen, W. Frey, S. Kim, et al., Nano Lett. 11 (2011) 348-354. DOI:10.1021/nl1042006 |
| [100] |
M. Sun, L. Xu, W. Ma, et al., Adv. Mater. 28 (2016) 898-904. DOI:10.1002/adma.v28.5 |
| [101] |
A. Xia, Y. Gao, J. Zhou, et al., Biomaterials 32 (2011) 7200-7208. DOI:10.1016/j.biomaterials.2011.05.094 |
| [102] |
L. Cheng, K. Yang, Y. Li, et al., Angew. Chem. Int. Ed. 50 (2011) 7385-7390. DOI:10.1002/anie.v50.32 |
| [103] |
J.N. Liu, W.B. Bu, L.M. Pan, et al., Angew. Chem. Int. Ed. 52 (2013) 4375-4379. DOI:10.1002/anie.201300183 |
| [104] |
J. Kim, J. Kim, C. Jeong, et al., Adv. Drug Deliv. Rev. 98 (2016) 99-112. DOI:10.1016/j.addr.2015.12.018 |
| [105] |
S. Jiang, Y. Zhang, K.M. Lim, et al., Nanotechnology 20 (2009) 155101. DOI:10.1088/0957-4484/20/15/155101 |
| [106] |
X. Bai, S. Xu, J. Liu, et al., Talanta 150 (2016) 118-124. DOI:10.1016/j.talanta.2015.08.038 |
| [107] |
S.J. Tan, P. Kiatwuthinon, Y.H. Roh, et al., Small 7 (2011) 841-856. DOI:10.1002/smll.201001389 |
| [108] |
M.K.G. Jayakumar, A. Bansal, K. Huang, et al., ACS Nano 8 (2014) 4848-4858. DOI:10.1021/nn500777n |
| [109] |
C. Wang, L. Cheng, Z. Liu, Biomaterials 32 (2011) 1110-1120. DOI:10.1016/j.biomaterials.2010.09.069 |
| [110] |
X. Zhang, P. Yang, Y. Dai, et al., Adv. Funct. Mater. 23 (2013) 4067-4078. DOI:10.1002/adfm.v23.33 |
| [111] |
S. Han, A. Samanta, X. Xie, et al., Adv. Mater. 29 (2017). |
| [112] |
Q. Ding, Q. Zhan, X. Zhou, et al., Small 12 (2016) 5944-5953. DOI:10.1002/smll.201601724 |
| [113] |
Z. Xu, P. Ma, C. Li, et al., Biomaterials 32 (2011) 4161-4173. DOI:10.1016/j.biomaterials.2011.02.026 |
| [114] |
F. Zhang, G.B. Braun, A. Pallaoro, et al., Nano Lett. 12 (2012) 61-67. DOI:10.1021/nl202949y |
| [115] |
L. Zhou, Z. Chen, K. Dong, et al., Adv. Mater. 26 (2014) 2424-2430. DOI:10.1002/adma.v26.15 |
| [116] |
H. Lin, S.S. Gao, C. Dai, et al., J. Am. Chem. Soc. 139 (2017) 16235-16247. DOI:10.1021/jacs.7b07818 |
| [117] |
D.K. Chatterjee, L.S. Fong, Y. Zhang, Adv. Drug. Deliver. Rev. 60 (2008) 1627-1637. DOI:10.1016/j.addr.2008.08.003 |
| [118] |
F. Chen, S.J. Zhang, W.B. Bu, et al., Chem. Eur. J. 18 (2012) 7082-7090. DOI:10.1002/chem.201103611 |
| [119] |
W.P. Fan, W.B. Bu, B. Shen, et al., Adv. Mater. 27 (2015) 4155-4161. DOI:10.1002/adma.v27.28 |
| [120] |
J. Wang, R. Deng, M.A. MacDonald, et al., Nat. Mater. 13 (2014) 157-162. DOI:10.1038/nmat3804 |
| [121] |
Y.F. Wang, L.D. Sun, J.W. Xiao, et al., Chem. Eur. J. 18 (2012) 5558-5564. DOI:10.1002/chem.v18.18 |
| [122] |
H.T. Wong, F. Vetrone, R. Naccache, et al., J. Mater. Chem. 21 (2011) 16589-16596. DOI:10.1039/c1jm12796a |
 2018, Vol. 29
2018, Vol. 29 


