Interleukin-5 (IL-5), produced by Th2 cells, is a haematopoietic growth factor that involves in maturation, growth, activation, and survival of eosinophils and basophils in human body [1-4]. Particularly, through binding to the heterodimeric interleukin-5 receptor (IL-5R), IL-5 regulates the maturation of eosinophils in the bone marrow and their release into blood [5-8]. Recent studies suggest that abnormal accumulation of eosinophils causes pathological damage in tissue and results in eosinophilic disorders including a number of severe allergic diseases, including asthma, atopic dermatitis, and nasal rhinitis [9-11]. Considering the physiological roles of IL-5, blockage of the interaction between IL-5 and IL-5R using high affinity ligands, such as antibodies, may decrease the amount of eosinophils accumulated, thus becoming an attractive approach to treat related diseases. To date, two monoclonal antibodies of IL-5 have been approved in the clinical treatment of severe asthma, and many candidates targeting IL-5/ IL-5R interactions are in different stages of clinical trials for other eosinophilic disorders [12, 13].
While being proven to be highly effective with exceptional specificities and relatively long half-lives in comparison to conventional small molecule drugs, antibody drugs also have several disadvantages, including latent immunogenicity, difficult quality control during production due to their heterogeneity, high cost for the patients under treatment, and so on. Thus, development of therapeutic agents with improved properties and lower cost are still desired [14]. Recent years, D-peptides have drawn increasing attentions for their promising abilities to bind target proteins and intercept protein/protein interactions (PPI), as well as other advantageous properties comparing to antibodies, including smaller size, better organ or tumor penetration, reduced potential for triggering the immune system, and lower production costs, etc. [15]. Moreover, comparing to native L-peptides, the unnatural D-peptides exhibit better bioavailability as they are not recognized by natural enzymes [16]. However, identification of proper D-peptides with high binding affinity against target protein has been a challenge due to the difficulties of establishing large D-peptide libraries. Therefore, indirect approaches, such as mirrorimage phage display, become particularly useful. By screening up to billions of L-peptides displayed on phages against an unnatural protein composed of D-amino acids, high affinity ligands are selected. Based on the principles of Van't Hoff-Le Bel stereochemistry, the D-peptides that have the sequence mirroring the selected L-peptides should be the corresponding antagonist to the natural protein target [17-19]. To access proteins that consist of all D-amino acids, chemical synthesis is currently the only feasible approach whilst other approaches like bioengineering become incompetent [20-23].
Aiming to screen out peptide inhibitors to block the IL-5/IL-5R interaction, mirror-image protein of IL-5 must be prepared. Although chemical synthetic approaches have been successfully utilized in the preparation of several unnatural D-proteins [24-27], efficient synthetic route to IL-5 should be explored using natural Lamino acids, due to the relatively high price of D-amino acid derivatives. Herein we disclose our attempts toward the chemical synthesis of natural interleukin-5.
Structurally, IL-5 is a homodimeric protein linked by a pair of intermolecular disulfide bonds (Cys44, Cys86), wherein each monomer of IL-5 comprises 115 amino acid residues (Fig. 1) [28, 29]. Retrosynthetically, it was envisaged that prior to the final folding step, the reduced monomer form of IL-5 could be assembled from three segments, Ile1-Leu43, Cys44-Lys85, and Cys86-Ser115 (Fig. 2). Solid-phase peptide synthesis (SPPS) [30] and peptide hydrazide-based native chemical ligation (NCL) [31-38] would be the key technologies utilized to facilitate the synthesis.
|
Download:
|
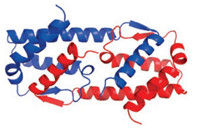
| Fig. 1. Crystal structure of hIL-5 (recombinant). | |

|
Download:
|
| Fig. 2. Primary sequence of the reduced monomeric form of human interleukin-5 (IL-5). Proposed ligation sites depicted in blue. | |
We carried out the forward synthesis first by preparing the peptide segments. While peptidyl hydrazine 3 could be generated following a reported SPPS procedure using HATU/HOBt as coupling reagents [39], it was noticed that an inseparable Thr23-deleted product 3′ was also formed (Scheme 1A). This byproduct formation could be suppressed by conducting double coupling of Fmoc-Thr(tBu)-OH using DIC/Oxyma as coupling reagents, and capping the unreacted sites with Boc-Gly-OH. The synthesis of segment 4a (Cys44-Lys85) also required optimization, as large amount of aspartimide was formed at Asp80-Gly81 under standard SPPS conditions (Scheme 1B). After a literature search, 2-hydroxy- 4-methoxybenzyl (Hmb) was used as a cleavable side chain to protect Gly81 [40, 41], which was able to inhibit the byproduct formation, but with dramatically decreased synthetic efficiency. Ultimately, the addition of 0.1 mol/L Oxyma to the Fmoc deprotection solution was found to effectively prevent aspartimide formation, providing pure peptide 4a in a decent isolated yield [42, 43]. Synthesis of the third segment turned out to be problematic, presumably because of the hydrophobic nature of the sequence, which significantly decreased the efficiency of SPPS and peptide purification. To solve this solubility issue, a depsidipeptide unit (Asn108-O-Thr109) was incorporated, which significantly increased the solubility of peptide and improved the synthetic efficiency (Scheme 1C). Such modification to the segment could be converted to the native amide bond via an O-to-N acyl transfer in the later ligation step conducted at neutral conditions [44-46]. The details of peptide synthesis can be found in Supporting information.

|
Download:
|
| Scheme 1. Syntheses of peptide segment 3 (A), segments 4a and 4b (B), and segment 5 (C). (a) Fmoc-based SPPS (standard coupling conditions: HATU, HOBt, DIEA, DMF, r.t., 20 min). (b) TFA/TIPS/H2O (95:2.5:2.5, v/v/v), r.t., 2–3 h. (c) Fmoc-Thr(tBu)-OH, DIC, Oxyma, DMF, r.t., 2 h × 2; then capped with Boc-Gly-OH, DIC, Oxyma, DMF, r.t., 2 h. (d) Fmoc-based SPPS (standard coupling conditions: HATU, HOAt, DIEA, DMF, r.t., 30 min). (e) Fmoc-Asn(Trt)-OH, DIC, NMI, DCM, r.t., 2 h × 2. | |
Next, we attempted to assemble the monomer of IL-5 in the C to N direction (Scheme 2A). The ligation of peptides 4a and 5, as well as the removal of acetamidomethyl (Acm) group on peptide 6 proceeded smoothly, affording peptide 7 in an acceptable conversion (26%). However, the followed reaction between peptide segments 3 and 7 was sluggish, presumably due to the hydrophobicity of 7 and the sterics derived from the C-terminal leucine of peptide 3. Alternatively, the assembly of segments in the N to C direction was investigated (Scheme 2B), where peptide 4b without the Acm-protection at the N-terminal cysteine was utilized accordingly. Although the ligation between segments 3 and 4b gave no conversion at room temperature, when the reaction was incubated at 60 ℃ for 1 h, the desired product was generated and isolated in a yield of 23%. In the event of ligation between segments 8 and 5, incomplete oxidation of peptidyl hydrazide 8 was observed under the condition of pH > 3.0. When the pH was adjusted to 2.9, peptide 8 could be completely converted to the corresponding hydrazide, which reacted with peptide 4 to afford IL-5 monomer 2 in 35% isolated yield after HPLC purification. The details of peptide ligation can be found in Supporting information.

|
Download:
|
| Scheme 2. Synthesis toward human interleukin-5 (IL-5). Reagent and conditions: (ⅰ) 4a (2 mmol/L), 500 μL of Buffer A (6mol/L Gn·HCl, 200 mmol/L NaH2PO4, pH 3–4), 6.5 equiv. NaNO2 (0.2 mol/L), -15 ℃, 20min, 340 μL of buffer (6 mol/L Gn·HCl, 200 mmol/L Na2HPO4, 200 mmol/L MPAA, pH 7.0), 5 (0.9 equiv.), adjust pH to 7.0, r.t., 12h, then 880 μL of Buffer B (6 mol/L Gn·HCl, 200 mmol/L Na2HPO4, 200 mmol/LTCEP·HCl, pH 6.5–6.9), 34%; (ⅱ) 6 (1.2mmol/L), 900 μL of buffer (6mol/L Gn·HCl, 200 mmol/L NaH2PO4, pH 7), 15 equiv. PdCl2, 37 ℃, 30 min, then 1mL of buffer (6 mol/L Gn·HCl, 1mol/L DTT, pH 7), 76%; (ⅲ) 3 (1mmol/L), 2.4μL of Buffer A, 6.5 equiv. NaNO2 (0.2mol/L), -15 ℃, 20 min, 50mg of MPAA (200 mmol/L), adjust pH to 7.0, 4b (0.9 equiv.), 60 ℃, 1h, then 1 mL of Buffer B, 23%; (ⅳ) 8 (1 mmol/L), 2.4 mL of Buffer A (pH 2.9), 15 equiv. NaNO2 (0.2mol/L), -15 ℃, 30 min, 100 equiv. MPAA, adjust pH to 7.0, 5 (0.9 equiv.), 37 ℃, 2 h, then 0.2μL of Buffer B, 35%. | |
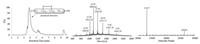
With the full sequence of interleukin-5 monomer 2 in hand, we attempted the refolding following a literature report studying E. Coli expressed IL-5 [47]. Unfortunately, only the oxidized monomer containing one intramolecular disulfide bond was observed as the major product based on mass spectropic analysis (Fig. 3). Other folding conditions, including the redox system mediated byglutathione (GSH) [48], were also evaluated. However, none of them afforded the desired product, where poor solubility of the linear peptide in buffers might be the major cause for the unsuccessful folding.

|
Download:
|
| Fig. 3. UPLC-MS analysis of the mixture obtained from folding of 2. Left: Mass signal as a function of time. Middle and Right: Q-TOF data of the major product (tR =4 min). Calcd. for C588H956N160O174S3 (oxidized form of IL-5 monomer containing one intramolecular disulfide): 13147.24 Da (average isotopes), (m/z) [M+8H]8+: 1644.01, [M+9H]9+: 1461.45, [M+10H]10+: 1315.41, [M+11H]11+: 1195.92, [M+12H]12+: 1096.34, [M+13H]13+: 1012.09, [M+14H]14+: 939.87; found [M+8H]8+: 1644.39, [M+9H]9+: 1461.79, [M+10H]10+: 1315.71, [M+11H]11+: 1196.20, [M+12H]12+: 1096.60, [M+13H]13+: 1012.25, [M+14H]14+: 940.09. | |
In summary, we have successfully synthesized the native sequence of reduced L-IL-5 using a convergent synthetic route featuring two peptidyl hydrazide-based NCL. The preliminary folding experiments suggest that the hydrophobicity of the IL-5 sequence brings many challenges to the formation of correct dimeric structure. This issue may be overcome by increasing the concentration of unfolded monomer, which requires large scale production of the folding substrate [47]. Alternatively, by introducing glycosylation to the protein, the solubility may be increased for both unfolded and folded forms. Further investigation of refolding conditions, as well as synthesizing glycosylated IL-5, is currently undergoing, and will be disclosed in due course.
AcknowledgmentsThe authors are grateful for financial support from the National Recruitment Program of Global Youth Experts (1000 Plan), Peking University Health Science Center (No. BMU2017QQ008), and the State Key Laboratory of Natural and Biomimetic Drugs. We thank Hongxing Li, Yuankun Dao, and Lingqi Qiu (Peking University) for helpful discussions, and Dr. Yuan Wang and Weiqing Zhang (Peking University) for spectroscopic assistance.
Appendix A. Supplementary dataSupplementary material related to this article can be found, inthe online version, at doi: https://doi.org/10.1016/j.cclet.2018.05.014.
| [1] |
S. Greenfeder, S.P. Umland, F.M. Cuss, R.W. Chapman, R.W. Egan, Respir. Res. 2 (2001) 71-79. DOI:10.1186/rr41 |
| [2] |
R.R. Dickason, D.P. Huston, Nature 379 (1996) 652-655. DOI:10.1038/379652a0 |
| [3] |
K. Hirai, M. Yamaguchi, Y. Misaki, et al., J. Exp. Med. 172 (1990) 1525-1528. DOI:10.1084/jem.172.5.1525 |
| [4] |
R.L. Coffman, B.W.P. Seymour, S. Hudak, J. Jackson, D. Rennick, Science 245 (1989) 308-310. DOI:10.1126/science.2787531 |
| [5] |
K.D. Stone, C. Prussin, D.D. Metcalfe, J. Allergy Clin. Immunol. 125 (2010) S73-S80. |
| [6] |
E.J. Clutterbuck, E.M.A. Hirst, C.J. Sanderson, Blood 73 (1989) 1504-1512. |
| [7] |
S.E. Broughton, T.L. Nero, U. Dhagat, et al., Cytokine 74 (2015) 247-258. DOI:10.1016/j.cyto.2015.02.005 |
| [8] |
M.E. Rothenberg, S.P. Hogan, Annu. Rev. Immunol. 24 (2006) 147-174. DOI:10.1146/annurev.immunol.24.021605.090720 |
| [9] |
D. Simon, H.U. Simon, J. Allergy Clin. Immunol. 119 (2007) 1291-1300. DOI:10.1016/j.jaci.2007.02.010 |
| [10] |
E. Patino, A. Kotzsch, S. Saremba, et al., Structure 19 (2011) 1864-1875. DOI:10.1016/j.str.2011.08.015 |
| [11] |
M.F. Patterson, L. Borish, J.L. Kennedy, J. Asthma Allergy 8 (2015) 125-134. |
| [12] |
G. Varricchi, D. Bagnasco, F. Borriello, E. Heffler, G.W. Canonica, Curr. Opin. Allergy Clin. Immunol. 16 (2016) 186-200. DOI:10.1097/ACI.0000000000000251 |
| [13] |
J. Nixon, P. Newbold, T. Mustelin, G.P. Anderson, R. Kolbeck, Pharmacol. Ther. 169 (2017) 57-77. DOI:10.1016/j.pharmthera.2016.10.016 |
| [14] |
T.T. Hansel, H. Kropshofer, T. Singer, J.A. Mitchell, A.J.T. George, Nat. Rev. Drug Discov. 9 (2010) 325-338. DOI:10.1038/nrd3003 |
| [15] |
P. Vlieghe, V. Lisowski, J. Martinez, M. Khrestchatisky, Drug Discov. Today 15 (2010) 40-56. DOI:10.1016/j.drudis.2009.10.009 |
| [16] |
S.A. Funke, D. Willbold, Mol. Biosyst. 5 (2009) 783-786. DOI:10.1039/b904138a |
| [17] |
T.N.W. Schumacher, L.M. Mayr, D.L. Minor Jr., et al., Science 271 (1996) 1854-1857. DOI:10.1126/science.271.5257.1854 |
| [18] |
K. Wiesehan, D. Willbold, ChemBioChem 4 (2003) 811-815. DOI:10.1002/cbic.v4:9 |
| [19] |
N. Sun, S.A. Funke, D.J. Willbold, Biotechnology 161 (2012) 121-125. |
| [20] |
C.L. Lee, X.C. Li, Sci. China Chem. 59 (2016) 1061-1064. |
| [21] |
H.X. Li, S.W. Dong, Sci. China Chem. 60 (2017) 201-213. |
| [22] |
J.H. Yang, J.F. Zhao, Sci. China Chem. 61 (2018) 97-112. DOI:10.1007/s11426-017-9056-5 |
| [23] |
C.H. Kim, J.Y. Axup, P.G. Schultz, Curr. Opin. Chem. Biol. 17 (2013) 412-419. DOI:10.1016/j.cbpa.2013.04.017 |
| [24] |
M. Liu, C. Li, M. Pazgier, et al., Proc. Natl. Acad. Sci. U. S. A. 107 (2010) 14321-14326. DOI:10.1073/pnas.1008930107 |
| [25] |
B.D. Welch, J.N. Francis, J.S. Redman, et al., J. Virol. 84 (2010) 11235-11244. DOI:10.1128/JVI.01339-10 |
| [26] |
K. Mandal, M. Uppalapati, D. Ault-Riché, et al., Proc. Natl. Acad. Sci. U. S. A. 109 (2012) 14779-14784. DOI:10.1073/pnas.1210483109 |
| [27] |
H.N. Chang, B.Y. Liu, Y.K. Qi, et al., Angew. Chem. Int. Ed. 54 (2015) 11760-11764. DOI:10.1002/anie.201506225 |
| [28] |
M.V. Milburn, A.M. Hassell, M.H. Lambert, et al., Nature 363 (1993) 172-176. DOI:10.1038/363172a0 |
| [29] |
Y. Minamitake, S. Kodama, T. Katayama, et al., J. Biochem. 107 (1990) 292-297. DOI:10.1093/oxfordjournals.jbchem.a123041 |
| [30] |
R.B. Merrifield, J. Am. Chem. Soc. 85 (1963) 2149-2154. DOI:10.1021/ja00897a025 |
| [31] |
P.E. Dawson, T.W. Muir, I. Clark-Lewis, S.B.H. Kent, Science 266 (1994) 776-779. DOI:10.1126/science.7973629 |
| [32] |
E.C.B. Johnson, S.B.H. Kent, J. Am. Chem. Soc. 128 (2006) 6640-6646. DOI:10.1021/ja058344i |
| [33] |
G.M. Fang, Y.M. Li, F. Shen, et al., Angew. Chem. Int. Ed. 50 (2011) 7645-7649. DOI:10.1002/anie.201100996 |
| [34] |
G.M. Fang, J.X. Wang, L. Liu, Angew. Chem. Int. Ed. 51 (2012) 10347-10350. DOI:10.1002/anie.201203843 |
| [35] |
J.S. Zheng, S. Tang, Y.K. Qi, Z.P. Wang, L. Liu, Nat. Protoc. 8 (2013) 2483-2495. DOI:10.1038/nprot.2013.152 |
| [36] |
J.S. Zheng, M. Yu, Y.K. Qi, et al., J. Am. Chem. Soc. 136 (2014) 3695-3704. DOI:10.1021/ja500222u |
| [37] |
S. Tang, Y.Y. Si, Z.P. Wang, et al., Angew. Chem. Int. Ed. 54 (2015) 5713-5717. DOI:10.1002/anie.201500051 |
| [38] |
M. Pan, S. Gao, Y. Zheng, et al., J. Am. Chem. Soc. 138 (2016) 7429-7435. DOI:10.1021/jacs.6b04031 |
| [39] |
K.J. Doores, Y. Mimura, R.A. Dwek, et al., Chem. Commun. 13 (2006) 1401-1403. |
| [40] |
T. Johnson, M. Quibell, D. Owen, R.C. Sheppard, J. Chem. Soc. Chem. Commun. (1993), 369-372. |
| [41] |
T. Johnson, M. Quibell, R.C. Sheppard, J. Pept. Sci. 1 (1995) 11-25. DOI:10.1002/(ISSN)1099-1387 |
| [42] |
M.E. Petersen, M.T. Jacobsen, M.S. Kay, Org. Biomol. Chem. 14 (2016) 5298-5303. DOI:10.1039/C6OB00824K |
| [43] |
R.S. Funosas, A.E. Faham, F. Albericio, J. Pept. Sci. 98 (2012) 89-97. DOI:10.1002/bip.v98.2 |
| [44] |
I. Coin, J. Pept. Sci. 16 (2010) 223-230. DOI:10.1002/psc.v16:5 |
| [45] |
F. Liu, E.Y. Luo, D.B. Flora, A.R. Mezo, Angew. Chem. Int. Ed. 53 (2014) 3983-3987. DOI:10.1002/anie.201310735 |
| [46] |
Z.M. Wang, W.L. Xu, L. Liu, T.F. Zhu, Nat. Chem. 8 (2016) 698-704. DOI:10.1038/nchem.2517 |
| [47] |
A.E.I. Proudfoot, D. Fattah, E.H. Kawashima, A. Bernard, P.T. Wingfield, Biochem. J. 270 (1990) 357-361. DOI:10.1042/bj2700357 |
| [48] |
K. Mandal, S.B.H. Kent, Angew. Chem. Int. Ed. 50 (2011) 8029-8033. DOI:10.1002/anie.v50.35 |
 2018, Vol. 29
2018, Vol. 29 


