Protein, an important component of cells and tissues of the human body, is a natural drug delivery vehicle. While protein serum albumins serve as carriers for various hydrophobic molecules including hydrophobic drugs, it has a significant impact on the half-life of albumin-loading drugs [1]. Hence, protein-based nanomaterials are extensively investigated for drug delivery purposes [2]. Tremendous interest are paid to the self-assembly of proteins towards functional nanomaterials, which are inspired by naturally evolving self-assembly biological construction, consisting of multiple and sophisticated protein architectures in living organisms [3-7]. Proteins and protein-based assemblies represent the most diverse molecules in nature both structurally and functionally. The ability of DNA to self-assemble into a variety of nanostructures and nanomachines has been highlighted in many emerging articles [8-17]. Among these, the novel assembled structures were supported by strict base pairing of DNA [18, 19]. The assembled protein provides a template for the assembly of enzyme and immobilization substances consisting of assembled enzymes can be used to fabricate and improve efficiency and utilization of enzyme catalysis in several types of filter films, which can effectively decompose and filter bacteria or small molecules.
In this paper, fibers were formed via self-assembly of DNAconjugated bovine serum albumin (BSA) mediated by the DNA base pairing. The key strategy was to synthesize DNA–protein conjugate by using heterobifunctional crosslinker agent sulfosuccinimidyl–4 [N-maleimidomethyl]-cyclohexane-1-carboxylate (sulfo-SMCC). Although the cross-linking of proteins and DNAs with sulfo-SMCC has already been reported [20, 21], to the best of our knowledge, few studies focused on the assembly of fibers via DNA-protein conjugation. Our study presented for the first time the protein bovine serum albumin (BSA) linked to DNAs, and the resulting selfassembled system mediated by complementary base pairing of DNAs. The synthesis of the fibers was carried out by a four-step procedure as schematically illustrated in Fig. 1.
|
Download:
|
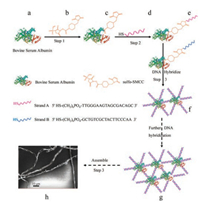
| Fig. 1. Schematic of the fiber formation process. In step 1, the crosslinker sulfoSMCC (b) reacted with the dissociative amino-group of the protein BSA (a) to produce a maleimide-activated protein (c: BSA-SMCC). In step 2, the maleimideactivated protein (c: BSA-SMCC) reacted with thiolated DNAs (d) in the appropriate molar ratio and produced BSA-SMCC-DNAs (e) through addition reaction. The H-S bond of thiolated DNA was added to the double bond in the SMCC of BSA-SMCC, while the crosslinking agent was omitted in the schematic picture of the following step in order to make the graph simple. In this step, the protein BSA was connected with thiolated DNA via sulfo-SMCC and product (e). In step 3, the protein BSA molecules were connected with each other through DNA hybridizing and produced polymer (f). Then the polymer (f) formed polymer (g) through further DNA hybridization. The produced BSA-SMCC-DNAs (g) self-assembled into fibers (h) via further complementary base paring of the DNAs. | |
In step 1, crosslinker sulfo-SMCC (b) reacted with amino-group of protein BSA (a) to produce a maleimide-activated protein BSASMCC (c). After removing the excess of crosslinker and byproducts, the maleimide-activated protein BSA-SMCC (c) reacted with thiolated DNAs (d) in an appropriate molar ratio to produce BSA-SMCC-DNAs (e) through addition reaction in Step 2. In this step, the protein BSA was conjugated with thiolated DNA through sulfo-SMCC linker and produced (e) (the H-S bond of thiolated DNA was added to the double bond of SMCC in BSA-SMCC, however, the crosslinking agent was omitted in the schematic illustration for simplification). Each protein molecule can conjugate with indefinite number of thiolated DNA molecules. In step 3, two molecules of BSA-DNA were linked through DNA hybridization to produce a dimer (f), which, in turn, further hybridized with other BSA-DNA so that a polymer (g) was formed. Finally, the polymer BSA-SMCCDNAs (g) self-assembled into fibers (h) via further complementary base paring of the DNAs (the number of DNA base was 18). The sequences of complementary DNAs (strands A and B) were shown in Table S1 in the Supporting information.
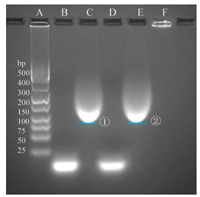
As shown in Fig. 2, Lanes A, B, and C represented DNA ladder, thiolated DNAs (strand A), and BSA-SMCC-DNAs (strand A), respectively. Comparing the thiolated DNAs in lane B with that of BSA-SMCC-DNAs in lane C, a new band ① was observed. This indicated that BSA was successfully combined with thiolated the DNAs (strand A) and formed BSA-SMCC-DNAs complex. In addition, lanes D and E were the thiolated DNAs (strand B) and BSA-SMCC-DNAs (strand B), respectively. Comparing lanes D and E, a new band ② of BSA-SMCC-DNAs (Strand B) appeared in lane E, indicating that BSA was successfully combined with the thiolated DNAs (Strand B). Both bands ① and ② verified that a BSA molecule could conjugate with indefinite number of DNA, as a result, compounds of different molecular weights were produced.
|
Download:
|
| Fig. 2. Agarose gel electrophoresis of thiolated DNAs, BSA-SMCC-DNAs and fibers at the initial stage. Lane A is the DNA ladder. Lane B is the thiolated DNAs (Strand A). Lane C is the BSA-SMCC-DNAs (Strand A) solution, comparing with thiolated DNAs in lane B, the BSA-SMCC-DNAs formed new band ①. Lane D is the thiolated DNAs (Strand B). Lane E is the BSA-SMCC-DNAs (Strand B) solution, comparing with thiolated DNAs in lane D, the BSA-SMCC-DNAs (Strand B) formed new band ②. Lane F was the self-assembled BSA via complementary base pairing of thiolated DNAs at the initial stage of fibers genesis, comparing with the new band ① and ② in the lane C and lane E, the fibers formed a new band in the well. | |
Lane F represented the self-assembled BSA via complementary base pairing of thiolated DNAs. When compare with the two new bands ① and ② in lanes C and E, respectively, the fibers formed was stuck in the well. These results demonstrated that BSA was successfully conjugated with thiolated DNAs, and further selfassembled into fibers via complementary DNA base pairing hybrid.
The fibers were observed by three microscopic techniques including optical microscopy (OM), scanning electron microscopy (SEM), and atomic force microscopy (AFM). The images were shown in Figs. 3 and 4. As shown in Fig. 3a and Fig. S1 in Supporting information, dendritic features were observed. Interestingly, some fibers grew on top of the silicon wafer and had waterweed-like shape (Fig. 3b). Moreover, the corkscrew spin phenomenon was observed around the fibers and the details were highlighted in Fig. S1b. The resulting fibers exhibited dendritic characteristic, yielding a semiregular linear array over micrometers [22]. In addition, little limb that grew on the trunk was also found.

|
Download:
|
| Fig. 3. Optical microscopy (a, c), SEM (b), fluorescence microscopy images and (d) of assembled fibers. (f) The sketch map of the assembling mechanism of fibers. | |
The experiment was repeated under the same condition but by using FITC-labelled BSA. Fig. 3c shows the fiber in bright field channel. It could be seen from Fig. 3d that the fiber emitted green fluorescence under blue light excitation due to the FITC, the fluorescent group that was labelled on BSA. These green spots thus indicated BSA molecules. Thus, it could be inferred that BSA molecules were first conjugated with DNA molecules (Fig. 3f) and formed linear chains. These chains were further combined in a parallel manner through DNA hybridization so that the fibers were formed.
Theoretically, protein BSA can self-aggregate through sulfoSMCC because it contains amino and thiol groups. In order to verify that the fibers were not formed as the result of BSA selfaggregation, a control experiment was performed in the absence of thiolated DNA under the same condition and the optical microscopic images were shown in Fig. S2 in Supporting information. The result showed that while several irregularly shaped substances were found, fiber was not observed. This indicated that fibers were not formed through self-aggregation of BSA. Additional control experiments in the absence of BSA or sulfoSMCC were also performed at the same condition and fibers were not observed in the optical microscopic images. These results demonstrated that all components including protein BSA, thiolated DNA, and the linker sulfo-SMCC were indispensable for the formation of fiber.
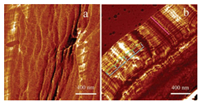
Microstructure of the assembled fiber was investigated by tapping-mode AFM. The selected AFM phase images were shown in Fig. S3 in Supporting information. The images presented in Figs. 4a and b were corresponded to the selected red area in Figs. S3a and S3b, respectively. Figs. 4a and b show the helical and straight regions of the fibers, respectively. In contrast to Fig. 4a, which lamellar structures were observed on the surface of the fibers, many parallel lamellae were observed in Fig. 4b. Moreover, as shown in Fig. 4b, the lamellae were composed of repeated lamella of about 220 nm with an average width (thickness) of about 20 nm (a number averaged from 26 lamellae with the length of 523 nm, as indicated by the pink line in Fig. 4b). This can be inferred that one assembled repeat contains three parallel DNAs (18 base-pair of about 6 nm in length) [23] and one BSA molecule. BSA assembled into fibers through multiple parallel layers of lamellae. Similar characteristics of the assembled fibers were observed in all microscopic observation including optical microscopy, SEM, confocal laser scanning microscopy, and AFM.

|
Download:
|
| Fig. 4. AFM phase images of the assembled fiber. The scan bar is 400 nm. | |
Protein is a biological macromolecule with complex structure that can provide randomly distributed sites for hydrophobic and electrostatic interactions on its surface. Thus, it is challenging to effectively control the process of protein self-assembly and to obtain accurate and orderly assembled structures. In this study, the amino groups of BSA were conjugated with thiol DNAs through heterobifunctional crosslinker agent (sulfo-SMCC), resulting in BSA molecules capable of self-assembly. The mechanism, in which the assembly took placed, could be inferred that BSA were first linked to the complementary base paring of DNA and formed linear chains. These chains were then further assembled through DNA hybridization and formed the fibers. Enzyme is a type of protein that possesses catalytic ability. Filter membranes that are made from enzymes not only can filter but also decompose some small molecules and bacteria. Thus, assembled enzymes could increase efficiency of the filter due to increasing numbers of enzymes it contains. Such assembly can therefore lead to immobilization of enzymes, which may overcome possible drawbacks when enzymes play the key role.
In conclusion, the dendritic threaded fibers were obtained though assembly of DNA-conjugated BSA molecules via complementary base pairing of DNA. The method was based on the use of heterobifunctional cross-linker sulfo-SMCC, which covalently linked protein and thiolated DNAs. By the use of fluorencence labelling method, microscopic studies, and control experiments, the mechanism of which the fibers were assembled, was analysed and inferred. The assembly strategy adopted here would pave the route for future potential in biomedical applications, such as supramolecular assembly, biosensor, delivery, and release of biological molecules and tissue engineering.
AcknowledgmentsThis work was supported by the National Natural Science Foundation of China (NSFC, No. 21174029), the Industry Academia Cooperation Innovation Fund of Jiangsu Province (No. BY2014127-07), the Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD) and the Fundamental Research Funds for the Central Universities (No. 2242016K41020).
Appendix A. Supplementary dataSupplementary data associated with this article can be found, in the online version, at https://doi.org/10.1016/j.cclet.2017.09.055.
| [1] |
A.O. Elzoghby, W.M. Samy, N.A. Elgindy, J. Control Release 157 (2012) 168-182. DOI:10.1016/j.jconrel.2011.07.031 |
| [2] |
M.J. Hawkins, P. Soon-Shiong, N. Desai, Adv. Drug Deliver. Rev. 60 (2008) 876-885. DOI:10.1016/j.addr.2007.08.044 |
| [3] |
H.C. Sun, L. Miao, J.X. Li, et al., ACS Nano 9 (2015) 5461-5469. DOI:10.1021/acsnano.5b01311 |
| [4] |
S.I. Lim, C.I. Lukianov, J.A. Champion, J. Control Release 249 (2017) 1-10. DOI:10.1016/j.jconrel.2017.01.007 |
| [5] |
U. Shimanovich, I. Efimov, T.O. Mason, et al., ACS Nano 9 (2015) 43-51. DOI:10.1021/nn504869d |
| [6] |
B. Rad, T.K. Haxton, A. Shon, et al., ACS Nano 9 (2015) 180-190. DOI:10.1021/nn502992x |
| [7] |
C.B. Winiger, S.M. Langenegger, G. Calzaferri, R. Haner, Angew. Chem. Int. Ed. 54 (2015) 3643-3647. DOI:10.1002/anie.201410041 |
| [8] |
M.A. Kostiainen, P. Hiekkataipale, A. Laiho, et al., Nat. Nanotechnol. 8 (2013) 52-56. DOI:10.1038/nnano.2012.220 |
| [9] |
V. Liljestrom, J. Mikkila, M.A. Kostiainen, Nat. Commun. 5 (2014) 1-9. |
| [10] |
Y.S. Bai, Q. Luo, W. Zhang, et al., J. Am. Chem. Soc. 135 (2013) 10966-10969. DOI:10.1021/ja405519s |
| [11] |
L. Miao, J.S. Han, H. Zhang, et al., ACS Nano 8 (2014) 3743-3751. DOI:10.1021/nn500414u |
| [12] |
H.C. Sun, L. Miao, J.X. Li, et al., ACS Nano 9 (2015) 5461-5469. DOI:10.1021/acsnano.5b01311 |
| [13] |
E. Auyeung, W. Morris, J.E. Mondloch, et al., J. Am. Chem. Soc. 137 (2015) 1658-1662. DOI:10.1021/ja512116p |
| [14] |
M. Bartke, B. Eickenberg, F. Wittbracht, A. Hutten, Part. Part. Syst. Charact. 32 (2015) 583-587. DOI:10.1002/ppsc.v32.5 |
| [15] |
I.M. Storm, M. Kornreich, A. Hernandez-Garcia, et al., J. Phys. Chem. B 119 (2015) 4084-4092. DOI:10.1021/jp511412t |
| [16] |
Z.M. Zhang, P.C. Gao, Z.F. Wang, B.W. Sun, Y. Jiang, Chem. Commun. 51 (2015) 12996-12999. DOI:10.1039/C5CC05164A |
| [17] |
J. Wei, Y.L. Li, P.C. Gao, et al., Chem. Commun. 53 (2017) 4581-4584. DOI:10.1039/C6CC09858D |
| [18] |
C.M. Niemeyer, J. Koehler, C. Wuerdemann, Chembiochem 3 (2002) 242-245. DOI:10.1002/1439-7633(20020301)3:2/3<242::AID-CBIC242>3.0.CO;2-F |
| [19] |
C.M. Niemeyer, T. Sano, C.L. Smith, C.R. Cantor, Nucleic Acids Res. 22 (1994) 5530-5539. DOI:10.1093/nar/22.25.5530 |
| [20] |
F. Kukolka, C.M. Niemeyer, Org. Biomol. Chem. 2 (2004) 2203-2206. DOI:10.1039/B406492E |
| [21] |
L.K. Zhao, J.Y. Zhang, H.M. Xu, H. Geng, Y.Q. Cheng, ACS Appl. Mater. Inter. 8 (2016) 22923-22929. DOI:10.1021/acsami.6b07803 |
| [22] |
M. Bruning, L. Kreplak, S. Leopoldseder, et al., Nano Lett. 11 (2010) 4533-4537. |
| [23] |
S.A. Contera, H. Iwasaki, S. Suzuki, Ultramicroscopy 97 (2003) 65-72. DOI:10.1016/S0304-3991(03)00030-5 |
 2018, Vol. 29
2018, Vol. 29 


