文章信息
- Zhang Lin, Tan Xiaofeng, Hu Jiao, Wuyun Tan, Yuan Deyi, He Xiaoyong, Long Hongxu, Li Xiugen
- 张琳, 谭晓风, 胡姣, 乌云塔娜, 袁德义, 何小勇, 龙洪旭, 李秀根
- Molecular Cloning and Characterization of PbSFBB13-gamma from Chinese White Pear (Pyrus bretschneideri)
- 中国白梨PbSFBB13-gamma基因的分子克隆与序列特征
- Scientia Silvae Sinicae, 2009, 45(11): 36-43.
- 林业科学, 2009, 45(11): 36-43.
-
文章历史
- 收稿日期:2008-02-25
-
作者相关文章
2. 浙江省丽水市科普工作指导站 丽水 323000;
3. 中国农业科学研究院郑州果树研究所 郑州 450009
2. Lishui Popular Science Station of Zhejiang Province Lishui 323000;
3. Zhengzhou Fruit Research Institute, Chinese Academy of Agricultural Sciences Zhengzhou 450009
Gametophytic self-incompatibility (GSI) is a mechanism adopted by many plant species to prevent self-fertilization and promote out-crossing (Franklin-Tong et al., 2003). This type of self-incompatibility (SI) is controlled by an apparently single multi-allelic locus, designated S-locus, which determines the specificity of the response. Early studies have shown that two separate genes encoding male (pollen) and female (pistil) determinants at the S-locus control male and female specificities of SI response, respectively. In Japanese Pear (Pyrus pyrifolia), the female specificity determinants have been identified as a class of polymorphic proteins which have ribonuclease activity, thus they were labeled S-RNases (Sassa et al., 1992). More recently, Sassa et al.(2007) have analyzed BAC contig sequences around S3- and S9- RNases in apple to find pollen-specific polymorphic F-box protein genes termed S-locus F-box brothers (SFBB)(MdSFBB3-alpha, MdSFBB3-beta, MdSFBB9-alpha and MdSFBB9-beta), and additionally conducted RACE (rapid amplification of cDNA ends) to isolate six SFBB genes (PpSFBB4-alpha, PpSFBB4-beta, PpSFBB4-gamma, PpSFBB5-alpha, PpSFBB5-beta and PpSFBB5-gamma) from Japanese Pear. Later, Kakui et al. (2007) isolated eight PpSFBB-gamma genes from different S-genotypes of Japanese Pear by genomic PCR. These genes show S-haplotype-specific polymorphism, but sequence similarities among them are very high. Based on the sequence polymorphism of the PpSFBB-gamma genes, a CAPS/dCAPS system for identifying the S-genotypes of Japanese Pear cultivars was developed.
Chinese pears originated from China are economically important fruit trees of Rosaceae family, which include about 13 species (Teng et al., 2002). Among these species, Chinese White Pear (Pyrus bretschneideri) distributing extensively in China covers a large number of important cultivars such as 'Yali', 'Jinhuali' and 'Dangshansuli' etc. As many species in Rosaceae do, Chinese White Pear also exhibits GSI. To date, a number of S-RNase genes have been isolated from this pear species, and many cultivars have been S-genotyped based on S-RNase-specific amplification (Wuyun et al., 2005; 2006; Tan et al., 2005a; 2005b). However, the molecular mechanism of GSI of Chinese White Pear is not completely clear yet. To fully understand the mechanism, it is necessary to clone pollen S-genes and investigate their functions in GSI response. Therefore, the aim of this study is to isolate, identify and characterize SFB/SLF gene, the possible pollen S-gene, in Chinese White Pear, which will provide a scientific base for clarifying the mechanism of pear SI at the molecular level.
1 Materials and methods 1.1 Plant materialsFive Chinese White Pear cultivars with known S-genotypes 'Jinhua' (S13S18) (Tan et al., 2005b), 'Jinhuasihao' (S13S18) (Tan et al., 2005b), 'liuleng' (S16S19) (Zhang et al., 2008), 'Eli' (S13S34) (Zeng, 2007), and 'Mili' (S19S29) (Zhang et al., 2008) were used to clone SFBB genes in this study. Young leaves were collected in spring and stored at -80℃ until used. Flowers of 'Jinhua' pear were collected at balloon stage. Subsequently, pistils and pollen grains were detached, immediately frozen in liquid nitrogen and stored at -80℃ until used. All plant materials were provided by Zhengzhou Fruit Research Institute, Chinese Academy of Agricultural Sciences (CAAS), Zhengzhou City, Henan Province.
1.2 Preparation of total DNA and confirmation of the S-genotypesTotal genomic DNA was isolated from young leaves of the five cultivars as described by Tan et al. (2007). The S-genotypes of the five cultivars were confirmed by S-RNase-specific amplification with primers PF1 and PR1 (Tan et al., 2007). PCR reactions were performed on PE-9600 (Perkin-Elmer, USA). The PCR cycling parameters were consisted of an initial step of 94℃ for 3 min for pre-denaturation, 35 cycles of 94℃ for 45 s for denaturing, 50℃ for 45 s for primer annealing, and 72℃ for 2 min for extension, followed by final 1 cycle of 5 min at 72℃ for extension. PCR products were analyzed by electrophoresis in 1.8% TAE agarose gel.
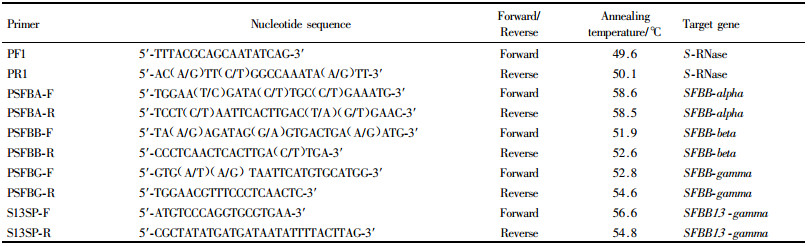
1.3 Isolation of SFBB gene in Pyrus bretschneideriThree pairs of primers (Tab. 1) were designed using SFB sequence information (Sassa et al., 2007) to amplify complete coding region of pear SFBBs. PCR was carried out in a total volume of 50 μL containing 5 μL 10×PCR Buffer, 4 μL dNTP Mix (2.5 mmol·L-1 each), 3 μL MgCl2 (25 mmol·L-1), 0.3 μL (around 1.5 U) ExTaq (TaKaRa, Japan), 2 μL of each primer (10 μmol·L-1), 1 μL total DNA (about 50 ng) and 32.7 ddH2O. The reactions were programmed as described above with a slightly-modified annealing temperature of 58.5, 52.0 and 53.0℃, respectively. PCR products were extracted from 2% agarose gel using a Gel Extraction Kit (Ambiogen, China), TA-ligated into pMD18-T Vector (Takara, Japan) and transformed into Escherichia coli strain DH5α. The positive clones were screened by the white/blue colony, and identified by plasmid PCR with BcaBESTTM Sequencing Primer M13-47 (5′-CGCCAGGGTTTTCCCAGTCACGAC-3′) and BcaBESTTM Sequencing Primer RV-M (5′-GAGCGGATAACAATTTCA CACAGG-3′). Two independent positive clones derived from each cultivar were sequenced.
|
|
Total RNAs were isolated from leaves, pistils and pollen grains of 'Jinhua' pear using Micro-to-Midi Total RNA Purification System (Invitrogen) and following the manufacture's protocols. Quality of the extracted RNA was checked by agarose gel electrophoresis, and concentrations of RNA were determined spectrophotometrically. To examine the organ expression specificity of the isolated PbSFBB13-gamma, reverse transcription (RT)-PCR was conducted using a cDNA amplification Kit (Takara, Japan) according to the manufacturer's directions from total RNAs from leaves, pistils and pollen grains, respectively. The first-step RT reaction was programmed for 1 cycle of 30℃ 10 min, 50 ℃ 25 min, and 95℃ 5 min. The second-step PCR amplification was carried out with primer pair S13SP-F/S13SP-R (Tab. 1) for 38 cycles of denaturation for 30 s at 94℃, annealing for 30 s at 55℃, and extension for 5 min at 72℃. S13SP-F and S13SP-R, a set of SFBB13-gamma-specific primers, were designed for distinguishing SFBB13-gamma from other possible pear SFBB genes on the basis of the sequence of SFBB13-gamma and nucleotide sequence comparison.
Extraction of RT-PCR products, TA ligation, transformation into E. coli and screening of positive clones were performed as described in section 1.3.
1.5 DNA sequencing and sequence analysisTarget clones were sequenced in both directions by Shanghai Biosune Biotechnology Ltd. Sequence data were analyzed with DNAMAN (version 6.0.40). Prediction of isoeletric point (pI), molecular weight (Wm) and Prosite was performed by using online ExPASy proteomics tools (http://expasy.org/tools/pi_tool.html). The BLAST algorithm was used to search the NCBI GenBank (http://www.ncbi.nlm.nih.gov/) databases for homologous sequences. Motif analysis of the target gene was conducted by Motif Scan (http://www.myhits.isb-sib.ch/) and sequence comparisons. Multiple amino acid sequences alignment was performed using the CLUSTAL X (version 1.83). Based on this alignment, a phylogenetic tree was constructed by the neighbor-joining method with MEGA (version 3.1). The following SFB/SLF sequences together with the Chinese White Pear SFBB13-gamma (PbSFBB13-gamma, EU081892) sequence were used for the phylogenetic analysis: 14 Japanese Pear (P. pyrifolia) SFBB sequences (PpSFBB1-gamma, AB297933; PpSFBB2-gamma, AB297934; PpSFBB3-gamma, AB297935; PpSFBB4-alpha, AB270797; PpSFBB4-beta, AB270798; PpSFBB4-gamma, AB270799; PpSFBB5-alpha, AB270800; PpSFBB5-beta, AB270801; PpSFBB5-gamma, AB270802; PpSFBB6-gamma, AB297936; PpSFBB7-gamma, AB297937;PpSFBB8-gamma, AB297938; PpSFBB9-gamma, AB297939; PpSFBBk-gamma, AB297940), six apple (Malus domestica) SFB/SLF sequences (MdSFBB3-alpha, AB270795; MdSFBB3-beta, AB270796; MdSFBB9-alpha, AB270793; MdSFBB9- beta, AB270794; MdSLF1, DQ422810; MdSLF2, DQ422811), four almond (Prunus dulcis) SFB sequences (PdSFBa, AB092966; PdSFBb, AB092967; PdSFBc, AB079776; PdSFBd, AB081648), two apricot (Prunus armeniaca) SFB sequences (ParSFB1, AY587563; ParSFB2, AY587562), three Japanese Apricot (Prunus mume) SFB sequences (PmSFB1, AB101440; PmSFB7, AB101441; PmSFB9, AB092645), and four Sweet Cherry (Prunus avium) SFB sequences (PavSFB1, AY805048; PavSFB2, AY805049; PavSFB3, AY805057; PavSFB4, AY649872).
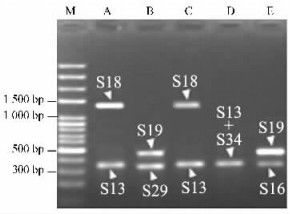
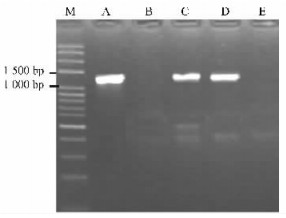
2 Results 2.1 Isolation, cloning and sequencing of SFB gene in Pyrus bretschneideriFollowing PCR amplification on genomic DNAs of 'Jinhua', 'Jinhuasihao', 'Eli', 'Yingzhiqing' and 'Liuleng', S-RNase-specific primers PF1/PR1 successfully amplified two fragments in each of the five cultivars apart from 'Eli' (Fig. 1). In the case of 'Eli', PCR products of around 380 bp were generated as blend band. The sizes of the total of nine fragments ranging from 380 bp to 1 400 bp well corresponded to S13, S16, S18, S19, S29 and S34, respectively. The S-genotypes of the five cultivars detected herein agreed with those reported previously. Thus, they were used for the next step of SFBB gene amplification with three different primer pairs PSFBA-F/PSFBA-R, PSFBB-F/PSFBB-R and PSFBG-F/PSFBG-R. The primer pair of PSFBA-F/PSFBA-R and PSFBB-F/PSFBB-R failed to amplify expected products in all of the five cultivars by testing of agarose gel electrophoresis analysis (data not shown). Using PSFBG-F/PSFBG-R, however, one fragment of approximately 1 300 bp was generated in each of 'Jinhua', 'Jinhuasihao' and 'Eli' (Fig. 2). In contrast, no PCR products were yielded in 'Liuleng' and 'Mili', indicating that this pair of primers might not be suitable for amplifying SFBB16-gamma, SFBB19-gamma, SFBB29-gamma and SFBB34-gamma genes. Two clones were randomly selected for sequencing from each of the three cultivars. Interestingly, six identical nucleotide sequences were obtained. A preliminary analysis by Blastn revealed that the sequence represented S-locus-linked F-box gene. Because the same sequence was isolated from three P. bretschneideri cultivars possessing an S-allele (S13) in common, this sequence was denominated as PbSFBB13-gamma (GenBank accession No. EU081892).
|
Fig.1 S-genotype detection of five Chinese White Pear cultivars by S-RNase-specific amplification M: 100 bp DNA ladder (Oumay, China); A: 'Jinhua'; B: 'Mili'; C: 'Jinhuasiha o'; D: 'Eli'; E: 'Liuleng'. Arrowheads represent the corresponding S- RNase alleles amplified by PF1/PR1. |
|
Fig.2 Amplification of SFBB-gamma genes in five Chinese White Pear cultivars with primer pair PSFBG-F/PSFBG-R M: 100 bp DNA ladder (Oumay, China); A: 'Jinhua'; B: 'Mili'; C: 'Jinhuasihao'; D: 'Eli'; E: 'Liuleng'. |
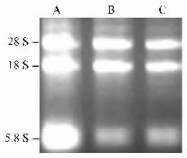
Using Total RNA Purification System (Invitrogen), the total RNAs were successfully isolated from leaves, pistils and pollen grains of 'Jinhua' pear. All of the three RNA samples had an average ratio OD 260/280 in the range 1.8-2.0, suggesting the RNAs were not contaminated by DNA. The RNA quality was verified by visualization on the agarose gel (Fig. 3), which showed no degradation and was of high quality enabling us to use it in the next RT-PCR.
|
Fig.3 Electrophoresis detection of total RNA A: Leaves; B: Pistils; C: Pollen grains. |
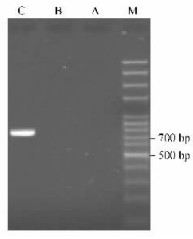
To examine organ expression of the PbSFBB13-gamma, RT-PCR was carried out from the total RNAs from leaves, pistils and pollen grains, respectively. As a result, only one cDNA fragment of approximately 750 bp was generated from the RNA of pollen grains, but not from leaves and pistils as shown by agarose gel electrophoresis (Fig. 4). Subsequently, the 750 bp-sized fragment was cloned and sequenced, and only one sequence was obtained. Sequence comparison revealed that the obtained cDNA sequence corresponded to SFBB13-gamma. These results indicated that the SFBB13-gamma was expressed specifically in pollen grains.
|
Fig.4 RT-PCR for detection of organ expression of SFBB13-gamma M: 100 bp DNA ladder (Oumay, China); A: Leaves; B: Pistils; C: Pollen grains. |
The PbSFBB13-gamma clone was 1 245 bp in length. It contained a putative initiation codon ATG and a putative termination codon TGA, which were identified by aligning the PbSFBB13-gamma with other reported PpSFBB-gamma genes. Comparison between the PbSFBB13-gamma and other 10 PpSFBB-gamma alleles (S1-9 and Sk) revealed extremely sequence similarities varying from 98.0% (with S6) to 98.9% (with S3) for the coding region at the nucleotide level. In comparison with other types of pear SFBB alleles, however, the PbSFBB13-gamma showed 80.3% similarity to PpSFBB4-alpha, 77.2% to PpSFBB4-beta, 80.0% to PpSFBB5-alpha, and 77.1% to PpSFBB5-beta. The similarity values supported that pear SFB genes were grouped into three types of alpha, beta and gamma as proposed by Sassa et al. (2007).
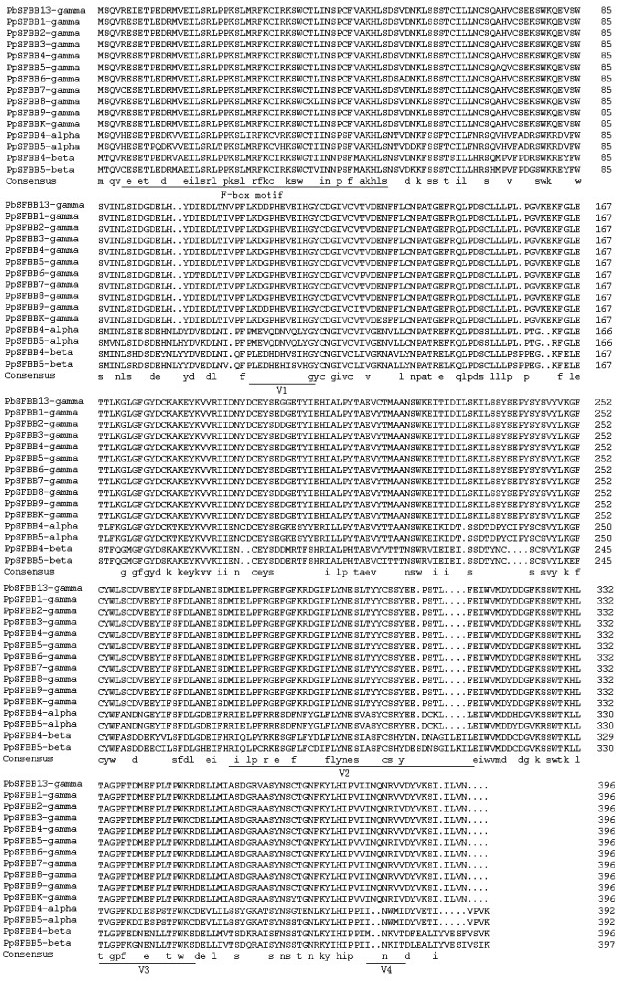
The Complete coding sequence of 1 191 bp for the PbSFBB13-gamma encoded a putative polypeptide of 396 amino acids. The molecular weight and isoeletric point (pI) of the PbSFBB13-gamma were predicted to be 45.4 ku and 4.63, respectively, by online proteomics tools. Motif Scan predicted an F-box motif of around 45 amino acids in the N-terminal region of the PbSFBB13-gamma (Fig. 5) which was common to reported SFB genes. Four variable regions (V1, V2, V3 and V4) were also defined as described by Sassa et al. (2007). Numbers of amino acid substitutions between the PbSFBB13-gamma and PpSFBB1 to PpSFBB9 and PpSFBBk were calculated to be only 9, 11, 9, 9, 11, 14, 10, 11, 10, and 13 of the total of 396 amino acids, respectively. At the deduced amino acid sequence level, the PbSFBB13-gamma also exhibited extremely high similarities ranging from 96.5% to 97.7% with PpSFBB-gamma genes which were remarkably higher than those with SFBB-alpha, SFBB-beta and SLF genes of pear and apple ranging from 62.8% (with MdSLF2) to 80.3% (with PpSFBB4-alpha). When compared with Prunus SFB genes, furthermore, the PbSFBB13-gamma showed the lowest similarities varying from 17.8% (with PdSFBa) to 25.1% (with PmSFB7).
|
Fig.5 Alignment of the deduced amino acid sequences of 15 pear SFBB genes The F-box motif and four variable regions (V1, V2, V3 and V4) are underlined. |
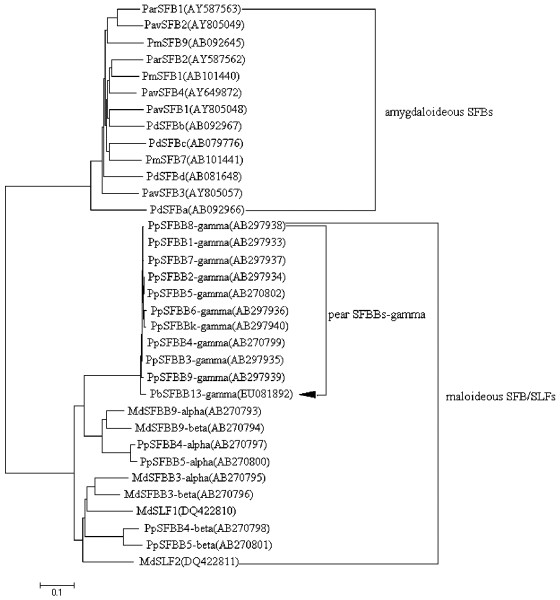
To better understand the evolutionary relationship of rosaceous SFB/SLF genes, the multiple amino acid sequences alignment was conducted using clustal X. A total of 34 complete amino acid sequences for SFB/SLF genes of seven species including Chinese White Pear (P. bretschneideri), Japanese Pear (P. pyrifolia), Apple (M. domestica), Almond (Prunus dulcis), Apricot (Prunus armeniaca), Sweet Cherry (Prunus avium), and Japanese Apricot (Prunus mume) were used in the alignment. Based on the alignment, a neighbor-joining phylogenetic tree was generated (Fig. 6). These 34 sequences mainly fell into two groups: maloideous SFB/SLF s and amygdaloideous SFBs. In amygdaloideous SFBs, four P. dulcis SFBs PdSFBa, PdSFBb, PdSFBc and PdSFBd did not pairwise cluster in the same branch, but with other Prunus SFBs. In maloideous group, likewise, PpSFBB-gamma genes were clustered with PbSFBB13-gamma forming a pear SFBB-gamma clade, but not with PpSFBB-alpha and PpSFBB-beta. This well mirrors the highest degree of similarities between PbSFBB13-gamma and PpSFBB-gamma genes. MdSFBB/SLFs, in addition, did not form a species-specific group, but clustered with PpSFBBs. The phylogenetic tree illustrated the distinct evolutionary path of SFB/SLFs between Maloideae and Amygdaloideae plants, and indicated that SFB/SLFs occurred after the divergence of subfamilies Maloideae and Amygdaloideae, but before the divergence of species such as P. pyrifolia, P. bretschneideri, M. domestica, P. dulcis, P. armeniaca, P. avium, and P. mume.
|
Fig.6 Phylogenetic analysis of the members of SFB/SLF genes in rosaceous plants The GenBank accession No. is shown in parentheses and the PbSFBB13-gamma is marked by an arrowhead. Branch lengths are proportional to genetic distance. |
Recently, researchers have focused on the search and cloning of pollen S-genes aiming to fully understand the mechanism of GSI, and much progress have been made. A series of SFB/SLFs were isolated and identified as the pollen S-genes or good candidates for pollen S-genes in the Solanaceae (Sijacic et al., 2004), Rosaceae (Entani et al., 2003), and Scrophulariaceae (Lai et al., 2002). In Prunus of Rosaceae, only one type of SFB/SLFs were discovered, whereas in apple and pear SFB/SLFs were classified into three types of alpha, beta and gamma and labeled SFB brothers (SFBBs). In this study, we designed three pairs of primers attempting to amplify the corresponding three types of SFBBs from Chinese White Pear. Consequently, PSFBG-F/PSFBG-R successfully amplified the product of 1 245 bp that was named PbSFBB13-gamma in this study, whereas the other two pairs of primers failed to amplify alpha and beta genes of PbSFBB. This result indicated that the latter two primers might not match well to PbSFBB alpha and beta genes in Chinese White Pear. The PbSFBB13-gamma showed extremely high similarities with other PpSFBB-gamma genes as described by Kakui et al. (2007). To our knowledge, this is the first report of SFB gene of Chinese pear. Although its function in GSI response is not clear, it should be a good pollen S candidate as it shows pollen-specific expression and typical sequence features, i.e. an F-box motif, and four variable regions which seem to be involved in the allelic specificity of the GSI response (Ushijima et al., 2003). Previously, Tan et al. (2007) investigated evolutionary relationship of rosaceous S-RNases by phylogenetic tree construction using 35 amino acid sequences. They found that rosaceous S-RNases formed two subfamily-specific groups, namely Maloideaous S-RNases and Amygdaloideous S-RNases. And they inferred that the S-RNases of Rosaceae predated the divergence of species. This result concurs with our phylogenetic analysis of rosaceous SFBs in the present study, although we used a different type of S-genes. As is known, pollen S-gene must be linked to the S-RNase gene at the S-locus, and the two genes are passed on the later generation as one unit to avoid breakdown of GSI (Kao et al., 1996). Our finding, therefore, that the evolutionary pattern of SFB/SLFs is consistent with that of S-RNases further confirmed the co-evolution between SFB/SLFs and S-RNases, which also strongly supported SFB genes including SFBB13-gamma as good candidates for pollen S-genes.
Although the molecular mechanism of GSI is not fully clear yet, it is suggested that pollen S-gene products should physically interacts with S-RNases to accomplish incompatible or compatible reactions (Golz et al., 1999). Hence, studies in further work on the interaction between the PbSFBB13-gamma and S-RNases will facilitate clarifying its role in incompatible reactions and the mechanism of GSI at the molecular level in Chinese pear. SFBB-gamma genes show S-haplotype-specific polymorphism although sequence similarities among them are very high. Based on the sequence polymorphism of the PpSFBB-gamma genes, Japanese researchers developed a CAPS/dCAPS system for identifying S-genotypes of Japanese Pear cultivars. As the range of Chinese pear cultivars analyzed is widened, more SFBB homologies will be isolated which may provide a novel approach to identifying the S-genotypes of Chinese pear cultivars.
Entani T, Iwano M, Shiba H, et al. 2003. Comparative analysis of the self-incompatibility (S) locus region of Prunus mume: identification of a pollen expressed F-box gene with allelic diversity[J]. Genes Cells, 8: 203-213. DOI:10.1046/j.1365-2443.2003.00626.x |
Franklin-Tong N, Franklin F C H. 2003. Gametophytic self-incompatibility inhibits pollen tube growth using different mechanisms[J]. Trends Plant Sci, 8: 598-605. DOI:10.1016/j.tplants.2003.10.008 |
Golz J F, Su V, Clarke A E, et al. 1999. A molecular description of mutations affecting the pollen component of Nicotiana alata S locus[J]. Genetics, 152: 1123-1135. |
Kakui H, Tsuzuki T, Koba T, et al. 2007. Polymorphism of SFBB-gamma and its use for S genotyping in Japanese pear (Pyrus pyrifolia)[J]. Plant Cell Rep, 26(9): 1619-1625. DOI:10.1007/s00299-007-0386-8 |
Kao T H, McCubbin A G. 1996. How flowering plants discriminate between self and non-self pollen to prevent inbreeding[J]. P Natl Acad Sci, 93: 12059-12065. DOI:10.1073/pnas.93.22.12059 |
Lai Zhao, Ma Wenshi, Han Bin, et al. 2002. An F-box gene linked to the self-incompatibility (s) locus of Antirrhinum is expressed specifically in pollen and tapetum[J]. Plant Mol Biol, 50: 29-42. DOI:10.1023/A:1016050018779 |
Sassa H, Hirano H, Ikehashi H. 1992. Self-incompatibility related RNases in styles of Japanese pear (Pyrus serotina Rehd.)[J]. Plant Cell Physiol, 33: 811-814. |
Sassa H, Kakui H, Miyamoto M, et al. 2007. S locus F-box brothers: multiple and pollen-specific F-box genes with S haplotype specific polymorphisms in apple and Japanese pear[J]. Genetics, 175: 1869-1881. DOI:10.1534/genetics.106.068858 |
Sijacic P, Wang X, Skirpan A L, et al. 2004. Identification of the pollen determinant of S-RNase-mediated self-incompatibility[J]. Nature, 429: 302-305. DOI:10.1038/nature02523 |
Tan Xiaofeng (谭晓风), Bi Fangcheng (毕方铖), Wuyun Tana (乌云塔娜), et al. 2005a. Studies of Chinese pear S-gene: Determination of 9 cultivars' gen otypes and isolation of S29-RNase gene in Pyrus bretschneideri. J Cent South Forest Univ (中南林学院学报), 25(4): 1-4.
|
Tan Xiaofeng (谭晓风), Wuyun Tana (乌云塔娜), Nakanishi T, et al. 2005b. Isolation and identification of S-RNase gene of the Pyrus bretschneideri (Chinese pear). J Cent South Forest Univ (中南林学院学报), 25(1): 1-3. http://en.cnki.com.cn/Article_en/CJFDTOTAL-ZNLB200501001.htm
|
Tan Xiaofeng, Zhang Lin, Wuyun Tana, et al. 2007. Molecular identification of two new self-incompatible alleles (S-alleles) in Chinese pear (Pyrus bretschneideri)[J]. J Plant Physiol Mol Biol, 33(1): 61-70. |
Teng Yuanwen, Kenji T, Fumio T, et al. 2002. Genetic relationships of Pyrus species and cultivars native to east Asia revealed by randomly amplified polymorphic DNA markers[J]. J Am Soc Hortic Sci, 127(2): 262-270. |
Ushijima K, Sassa H, Dandekar A M, et al. 2003. Structural and transcriptional analysis of the self-incompatibility locus of almond: identification of a pollen-expressed F-box gene with haplotype-specific polymorphism[J]. Plant Cell, 15: 771-781. DOI:10.1105/tpc.009290 |
Wuyun Tana (乌云塔娜), Tan Xiaofeng (谭晓风), Bi Fangcheng (毕方铖), et al. 2005. Studies of Chinese pear S-gene: Determination of 7 cultivars of the S-genotype and identification of two new allele of self-incompatibility of Pyrus bretschneideri (Chinese pear). J Cent South Forest Univ (中南林学院学报), 25(4): 7-12. http://en.cnki.com.cn/Article_en/CJFDTotal-ZNLB200504001.htm
|
Wuyun Tana (乌云塔娜), Tan Xiaofeng (谭晓风), Li Xiugen (李秀根), et al. 2006. Isolation and identification of S12-RNase gene of Pyrus bretschneideri. Scientia Silvae Sinicae (林业科学), 42(4): 117-121. http://www.linyekexue.net/EN/10.11707/j.1001-7488.20060421
|
Zeng Yanling (曾艳玲). 2007. Identification of S-genotype of Eli and the S-alleles cDNA cloning. Hunan: Central South University of Forestry and Technology.
|
Zhang Lin(张琳), Tan Xiaofeng(谭晓风), Yuan Deyi(袁德义), et al. 2008. Determination of S-genotype of 13 Chinese pear cultivars based on sequence analysis. J Zhej Univ: Agric & Life Sci (浙江大学学报: 农业与生命科学版), 34(1): 19-28. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=zjdxxb-nyysm200801005
|
 2009, Vol. 45
2009, Vol. 45