| 棉花植株形态学性状遗传网络的关联分析 |
2. 中国农业科学院棉花研究所,棉花生物学国家重点实验室/农业部棉花生物学与遗传育种重点实验室,河南 安阳 455000;
3. 浙江大学农业与生物技术学院,浙江省作物种质资源重点实验室,杭州 310058
2. State Key Laboratory of Cotton Biology/Key Laboratory of Cotton Biology and Genetic Breeding of Ministry of Agriculture, Institute of Cotton Research, Chinese Academy of Agricultural Sciences, Anyang 455000, Henan, China;
3. Zhejiang Key Lab of Crop Germplasm, College of Agriculture and Biotechnology, Zhejiang University, Hangzhou 310058, China
Upland cotton (Gossypium hirsutum L.) is one of the most important economic crops in the world. One of the main objectives for the cotton-producing countries is to improve fiber production. Vegetative growth is the basis for life circle of the plants,which provides the essential nutrition for initiating cotton flowers,bolls,and fiber. Revealing genetic basis and estimating effects for quantitative trait loci (QTLs) related to plant morphological traits[1] by linkage mapping can provide useful information for conducting marker-assisted selection (MAS).
Association mapping is an efficient method for detecting polymorphic genotypes to phenotypic variations,and the genome-wide association study (GWAS) is a special analysis method for association mapping. It can scan the molecular markers of entire genome for particular traits of interest[2, 3]. Mapping genetic loci can be performed by using RAPD (random amplification polymorphism DNA),SCAR (sequence characterized amplified region),SSR (simple sequence repeat),and SNP (single nucleotide polymorphism) markers,etc. In this study,we utilized PCR-based molecular markers SSRs,which came mostly from the cotton marker database (CMD).
Linkage mapping has been widely used in cotton among the previous studies. But most of them concentrated on the fiber traits and yield traits,such as micronaire value,lint yield,and boll mass[4, 5, 6, 7, 8]. These traits were direct factors related to cotton industry,so the breeders wanted to understand their genetic basis and find ways to improve them. Few works had been reported on plant morphological traits[9, 10, 11]. Indirect factors including plant height,stem diameter,and stem node,also influenced the growth of cotton and hence the yield. Therefore,we aimed to dissect the genetic basis,find the QTSs (quantitative trait SSRs) associated to the traits and provide more information for cotton breeding.
In this study,we performed association mapping for 872 marker loci of 39 cultivars and their 178 F1 hybrids in three environments. A newly developed software QTXNetwork (http://ibi.zju.edu.cn/software/QTXNetwork/) based on mixed linear model approaches[12] was applied in this study. A full genetic model with genetic effects of additive,dominance,epistasis and their environment interactions was built. The association mapping results revealed the importance of genetic architecture on plant morphological traits in cotton.
1 Materials and methods 1.1 Parent selection and experimental designIn this study,the mapping population was provided by the Institute of Cotton Research,Chinese Academy of Agricultural Sciences and Tarimu University. These cultivars were representatives of cotton-growing regions in China. Thirty-nine cotton cultivars were sampled and their 178 F1 hybrids were analyzed for four morphological traits at three environments (e1: Anyang of Henan Province in 2012; e2: Anyang of Henan Province in 2013; e3: Alar of Xinjiang Autonomous Region in 2012). Anyang of Henan Province located in the Yellow River valley cotton-growing region,and Alar of Xinjiang Autonomous Region located in the northwestward cotton-growing region. For field experiments,a completely randomized block design with three replications was applied. Each block was settled with two rows,and each row was kept in a plot of 5.00 m in length and 0.70 m in width. The distance between plants was 0.25 m in width in Anyang,and 0.10 m in width in Alar. Plant height (PLH),stem diameter (STD),stem first (STF) and stem node (STN) were evaluated to dissect the genetic network of these morphological traits.
1.2 DNA extraction and SSR markers screeningTwo cotyledons from two plants were collected randomly for each line in the plots in March 2012. DNA from 39 parent samples was extracted according to the method of PATERSON et al.[13]. The sequences of SSR primers,which came mostly from the cotton marker database (CMD,http://www.cottonmarker.org/cgi-bin/panel.cgi),were downloaded to obtain specific information about the markers. This database is an integrated web-enabled database,providing a centralized access to all publicly available cotton microsatellites and single nucleotide polymorphisms,which provides more efficient applications of molecular markers and facilitates basic and applied research in molecular breeding and genetic mapping in cotton. A standard screening panel contained the polymorphic information of upland cotton cultivars and other tetraploid species. Among the 39 lines,there were 5 052 SSR primer pairs in total to be examined for screening polymorphisms. Those markers included DPL,GH,HAU,JESPR,NAU,BNL,CIR,CGR,DC,MUSS,MGHES,C,CER and CM series. The marker nomenclature consisted of a letter that indicated the origin of the marker,followed by the primer number. SSR analysis was carried out according to the procedure proposed by YAO et al.[14]. To sum up,872 marker loci were obtained to conduct association analysis.
1.3 Statistical analysisTo analyze different effect values,a full genetic model was used for the phenotypic value of the k-th individual in the h-th environment (yhk),which can be presented by the following mixed linear model,
| $\begin{gathered} {y_{hk}} = \mu + \sum\limits_i {{a_i}x{A_{ik}} + } \sum\limits_i {{d_i}x{D_{ik}} + } \sum\limits_{i{\text{ < }}j} {a{a_{ij}}xA{A_{ijk}} + } \hfill \\ \sum\limits_{i{\text{ < }}j} {a{d_{ij}}xA{D_{ijk}} + } \sum\limits_{i{\text{ < }}j} {d{a_{ij}}xD{A_{ijk}} + } \hfill \\ \sum\limits_{i{\text{ < }}j} {d{d_{ij}}xD{D_{ijk}} + } {e_h} + \sum\limits_i {a{e_{ih}}uA{E_{ihk}} + } \hfill \\ \sum\limits_i {d{e_{ih}}uD{E_{ihk}} + } \sum\limits_{i{\text{ < }}j} {aa{e_{ijh}}xAA{E_{ijhk}} + } \hfill \\ \sum\limits_{i{\text{ < }}j} {ad{e_{ijh}}uAD{E_{ijhk}} + } \sum\limits_{i{\text{ < }}j} {da{e_{ijh}}uDA{E_{ijhk}} + } \hfill \\ \sum\limits_{i{\text{ < }}j} {ad{e_{ijh}}uDD{E_{ijhk}} + } {\varepsilon _{hk}} \hfill \\ \end{gathered} $ | (Equation1) |
The mixed linear model can be expressed in matrix notation as follows,
| $\begin{gathered} y = 1\mu + {X_A}{e_A} + {X_D}{e_D} + {X_{AA}}{e_{AA}} + {X_{AD}}{e_{AD}} + \hfill \\ {X_{DA}}{e_{DA}} + {X_{DD}}{e_{DD}} + {U_E}{e_E} + {U_{AE}}{e_{AE}} + \hfill \\ {U_{DE}}{e_{DE}} + {U_{AAE}}{e_{AAE}} + {U_{ADE}}{e_{ADE}} + {U_{DAE}}{e_{DAE}} + \hfill \\ {U_{DDE}}{e_{DDE}} + {e_\varepsilon } \hfill \\ = Xb + \sum\limits_{v = 1}^7 {{U_v}{e_v} + {e_\varepsilon }~{\text{MVN}}} \left( {Xb,\sum\limits_{\nu = 1}^7 {\sigma _v^2{U_v}U_v^T + I\sigma _\varepsilon ^2} } \right) \hfill \\ \end{gathered}$ | (Equation2) |
The total heritability can be estimated by the following equation,
| $\begin{gathered} h_T^2 = h_G^2 + h_{GE}^2 \hfill \\ = \left( {\sum\limits_i {h_a^2} + \sum\limits_j {h_d^2} + \sum\limits_{i{\text{ < }}j} {h_{aa}^2} + \sum\limits_{i{\text{ < }}j} {h_{ad}^2} + \sum\limits_{i{\text{ < }}j} {h_{da}^2} + \sum\limits_{i{\text{ < }}j} {h_{dd}^2} } \right) + \hfill \\ \left( {\sum\limits_i {h_{ae}^2} + \sum\limits_j {h_{de}^2} + \sum\limits_{i{\text{ < }}j} {h_{aae}^2} + \sum\limits_{i{\text{ < }}j} {h_{ade}^2} + \sum\limits_{i{\text{ < }}j} {h_{dae}^2} + \sum\limits_{i{\text{ < }}j} {h_{dde}^2} } \right) \hfill \\ = \left( {h_A^2 + h_D^2 + h_{AA}^2 + h_{AD}^2 + h_{DA}^2 + h_{DD}^2} \right) + \hfill \\ \left( {h_{AE}^2 + h_{DE}^2 + h_{AAE}^2 + h_{ADE}^2 + h_{DAE}^2 + h_{DDE}^2} \right) = \hfill \\ \left( {h_A^2 + h_D^2 + h_I^2} \right) + \left( {h_{AE}^2 + h_{DE}^2 + h_{IE}^2} \right) \hfill \\ \end{gathered}$ | (Equation3) |
The marker-trait association analyses were carried out by the above mixed linear model. All the detected marker-trait association loci were fitted by a full genetic model to predict the additive effects,dominance effects,epistasis effects,and their environment interaction effects. The mixed linear model framework with Henderson method Ⅲ[15] was used to construct the F-statistic test to implement the association analysis. Permutation test was set for a total of 2 000 times to calculate the critical F-value to control the experiment-wise type Ⅰ error (αEW < 0.05). The QTS effects were predicted by using the MCMC (Markov Chain Monte Carlo) algorithm with 20 000 Gibbs sample iterations[16]. Based on the information of genetic effects of detected-QTSs for the four plant morphological traits,the breeding values ($\hat u + \hat G + \hat GE$) for the superior lines and superior hybrids of the mapping population were predicted[17].
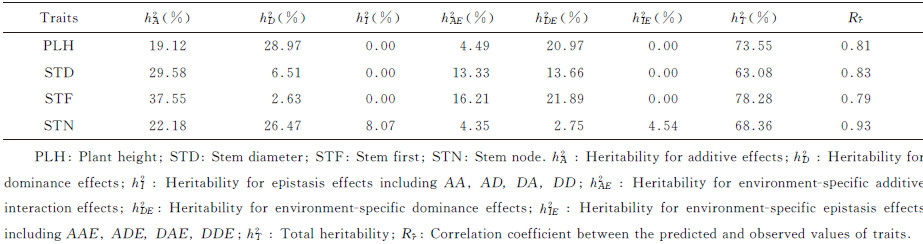
2 Results 2.1 Estimated heritability and genetic effectsA total of 25 experiment-wise significant QTSs (PEW-value < 0.05) were detected for the four plant morphological traits (5-7 QTSs for each trait) in cotton (Fig. 1,Table S1 available at http://www.journals.zju.edu.cn/agr/EN/article/showSupportInfo.do?id=10359). Estimated heritability was quite high (63.08%-78.28%),and mainly contributed by genetic main effects (hG2= 48.09% for PLH,36.09% for STD,40.18% for STF,and 56.72% for STN) (Table 1). It was suggested that plant morphological traits were quite stable across environments. The additive effects were the primary genetic component for STD and STF,while dominance effects were dominated for PLH and STN.
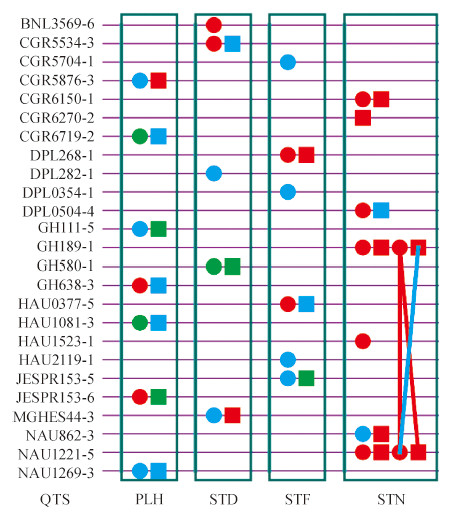 |
| Fig. 1 G×G plot of detected significant QTSs (PEW-value < 0.05) for four plant morphological traits |
| Table 1 Estimates of heritability of QTSs detected for four plant morphological traits |
 |
| 点击放大 |
Plant height (PLH) is one of the most common traits among the plant morphological traits,which plays an important role in formation of plant morphology type as a key trait. Cotton plant height is also considered to be related with yield and fiber quality traits. In this study,it was discovered that dominance and its environment interaction effects were the main components of total heritability (hD2≈28.97%,hDE2≈20.97%). No epitasis or their environment interaction was detected.
A total of seven QTSs were detected for PLH (Fig. 1,Table S1 available at http://www.journals.zju.edu.cn/agr/EN/article/showSupportInfo.do?id=10359). There were seven individual QTSs highly significant (PEW-value < 1×10-5) (Table 2),among which three QTSs (GH111-5,GH638-3 and JESPR153-6) were found with additive effects. And the other four QTSs were detected with dominance effects. The GH638-3 was detected with environment-specific dominance effects in e1,while NAU1269-3 was discovered with environment-specific additive effects in e1. No pair epistasis interaction effect was identified. Selecting genotypes qq of GH111-5 and QQ of GH638-3,JESPR153-6 for different environments could improve PLH. Due to large environment-specific dominance effect,heterozygote Qq of GH638-3 in e1 could increase PLH. Besides,selecting genotype qq of NAU1269-3 in e1 could also result in higher PLH.
| Table 2 Estimated genetic effects, standard error (SE), significance and heritability for highly significant QTSs (PEW-value < 1×10-5) of four plant morphological traits |
 |
| 点击放大 |
For STD,a total of five QTSs were detected (Fig. 1,Table S1 available at http://www.journals.zju.edu.cn/agr/EN/article/showSupportInfo.do?id=10359). Four highly significant QTSs (PEW-value < 1×10-5) were listed in Table 2. All the four highly significant QTSs were detected for additive effects. Environment-specific effects were identified for CGR5534-3 by de1 and DPL282-1 by ae1 and ae3. Increasing thick diameter of stem could be expected by selecting genotypes QQ of BNL3569-6 and qq of MGHES44-3. Selecting genotypes qq of DPL282-1 in e1 and Qq of CGR5534-3 in e1 could also improve this trait due to environment-specific additive/dominance effects.
For STF,there were six QTSs (CGR5704-1,DPL268-1,DPL0354-1,HAU0377-5,HAU2119-1 and JESPR153-5) highly significant with positive additive effects (Fig. 1,Table 2). For this trait,additive effects contributed nearly half heritability. In addition,CGR5704-1,HAU2119-1 and JESPR153-5 had environment-specific additive effects in e1 and e3,and the same was DPL0354-1 in all three environments. Besides,DPL268-1 was detected with positive effects. Among these six QTSs,all the effects were negative in e1 but positive in e3; therefore STF could be reduced via selecting genotypes QQ of CGR5704-1,DPL0354-1,HAU2119-1 and JESPR153-5 in e1 or qq of CGR5704-1 in e3.
The number of STN was determined by genetic and environmental factors,and had an essential impact on the final plant height. A total of eight QTSs were significantly dissected for STN (Fig. 1,Table S1 available at http://www.journals.zju.edu.cn/agr/EN/article/showSupportInfo.do?id=10359). Most genetic effects were contributed by additive and dominance effects,ranging from 22.18% to 26.47% (Table 1). For the highly significant QTSs (Table 2),CGR6270-2 had negative dominance effect,while other six QTSs (CGR6150-1,DPL0504-4,GH189-1,HAU1523-1,NAU862-3 and NAU1221-5) were additive effects. NAU862-3 was discovered with environment-specific additive effect in e1. The only one epistasis was between GH189-1 and NAU1221-5,and environment-specific dominance-additive effect in e1 was discovered (Table S1 available at http://www.journals.zju.edu.cn/agr/EN/article/showSupportInfo.do?id=10359),which/ had the highest heritability in STN. Selecting genotypes qq of DPL0504-4,NAU862-3 or QQ of CGR6150-1,GH189-1,HAU1523-1 and NAU1221-5 could increase STN. Genotype QQ of NAU862-3 in e1 could also be chosen to further increase STN.
2.4 Predicted genetic effects for different genotypesBased on the estimated genetic effects of each QTS for three environments,the maximum and minimum genotypic effects of the superior lines and superior hybrids were predicted (Table 3). Moreover,the genotypic effects of homozygotes (QQ,qq) and heterozygote (Qq) were also predicted for the four morphological traits in the three environments. For the traits of PLH,STD,STF and STN,the predicted genotypic effects were positive for major-allele homozygote QQ,but negative for minor-allele homozygote qq. It was suggested that the selection had the potential to increase plant height,stem diameter,first stem,and stem node. The predicted genotypic effects of heterozygote Qq were positive for PLH and STN in all environments,but various for STF and STD across the environments,suggesting that the PLH and STN could be improved through hybrid vigor.
| Table 3 Predicted total genotypic effects across environments (G) and in three environments for genotypes of QQ, qq, Qq, superior line and superior hybrid for four plant morphological traits |
 |
| 点击放大 |
These four morphological traits are moderate,and need to be balanced through breeding and cultivation techniques. Except for the superior lines (±) of STF,most of the absolute breeding values of superior lines (±) in the mapping population were larger than those of homozygote genotypes QQ and qq. It was indicated that the breeders could increase or decrease three morphological traits (PLH,STN and STD),through selecting superior lines with a mixture of homozygote QQ and qq genotypes of detected QTSs. And the breeding value of the superior hybrid could also be reinforced for each trait by selecting a hybrid with a mixture of heterozygote (Qq) and homozygotes (QQ,qq).
3 DiscussionAssociation mapping is a powerful tool to dissect quantitative trait loci underlying complex traits,and has been widely used in different studies. But due to the lack of suitable statistical methodology and heavy computational demand,epistasis and environment interaction effects were often missed in current GWAS[18, 19, 20]. Ignoring epistasis and environment interaction might lead to biased estimation of effects on loci and lower detecting power[21]. Furthermore,the heritability of target traits would be missed if reduced models were used[22, 23]. To solve the computational burden problems,LOU et al.[24] developed a new approach,which combined GPU (graphics processing unit) with GMDR (generalized multifactor dimensionality reduction). GPU could accelerate the operating speed in order to deal with millions of markers,and GMDR was able to detect high dimension interactions.
In this study,the software QTXNetwork was utilized to perform association mapping[12]. QTXNetwork is a software for GPU computation of linkage and association analyses on full genetic model,including epistasis and environment interaction of complex traits based on omics genotypes. Compared with the reduced models used in the previous research,a full mixed linear model (MLM) could dissect total heritability into different parts: additive,dominance,epistasis and their environment interactions. Although diverse softwares such as PLINK[25],TASSEL[26],R[27] were used,and different models including MLM,GLM were applied in previous studies,only the narrow or broad-sense heritability could be calculated[28, 29, 30, 31].
Several highly significant QTSs detected by this analysis for morphological traits were also found by other mapping analyses in cotton,which were DPL0504[32],CGR5534[33],NAU1269[34],HAU2119[35],JESPR153[36]. This could validate the efficiency of the mixed linear model approach for association mapping to identify relevant loci for complex traits. Furthermore,we identified different alleles of one SSR that had effects on STF (JESPR153-5) and PLH (JESPR153-6). This provided evidence that some QTSs could have pleiotropic or physiological associations for accelerating the process of MAS programs in cotton breeding. We had also calculated the R-values (correlation coefficient) of the predicted genotypic effects in Table 1. For these four traits,the R-values ranged from 0.79 to 0.93,which were close to the total heritability. It was suggested that the genotypic variation could be predicted very well by the full genetic model.
| [1] | WILLCOX M C, KHAIRALLAH M M, BERGVINSON D, et al. Selection for resistance to southwestern corn borer using marker-assisted and conventional backcrossing. Crop Science, 2002, 42(5):1516-1528. |
| [2] | NUSSBAUM R, MCINNES R R, WILLARD H F. Thompson & Thompson genetics in medicine. The Journal of the American Medical Association, 1992, 267(15):2115. |
| [3] | GIBSON G, MUSE S. A Primer of Genomic Science. 2nd ed. Sunderland, England: Sinauer Associates, 2005:408-409. |
| [4] | JIANG C X, WRIGHT R J, EL-ZIK K M, et al. Polyploid formation created unique avenues for response to selection in Gossypium (cotton). Proceedings of the National Academy of Sciences of the USA, 1998, 95(8):4419-4424. |
| [5] | ULLOA M, MEREDITH JR W R. Genetic linkage map and QTL analysis of agronomic and fiber quality traits in an intraspecific population. Journal of Cotton Science, 2000, 4(3):161-170. |
| [6] | PATERSON A H, SARANGA Y, MENZ M, et al. QTL analysis of genotype×environment interactions affecting cotton fiber quality. Theoretical and Applied Genetics, 2003, 106(3):384-396. |
| [7] | SUN F D, ZHANG J H, WANG S F, et al. QTL mapping for fiber quality traits across multiple generations and environments in upland cotton. Molecular Breeding, 2012, 30(1):569-582. |
| [8] | JIA Y H, SUN X W, SUN J L, et al. Association mapping for epistasis and environmental interaction of yield traits in 323 cotton cultivars under 9 different environments. PLoS ONE, 2014, 9(5):e95882. |
| [9] | WANG M H, JIANG N, JIA T Y, et al. Genome-wide association mapping of agronomic and morphologic traits in highly structured populations of barley cultivars. Theoretical and Applied Genetics, 2012, 124(2):233-246. |
| [10] | GUO Y F, MCCARTY J C, JENKINS J N, et al. QTLs for node of first fruiting branch in a cross of an upland cotton, Gossypium hirsutum L., cultivar with primitive accession Texas 701. Euphytica, 2008, 163(1):113-122. |
| [11] | LI C Q, SONG L, ZHAO H H, et al. Quantitative trait loci mapping for plant architecture traits across two upland cotton populations using SSR markers. The Journal of Agricultural Science, 2014, 152(2):275-287. |
| [12] | ZHANG F T, ZHU Z H, TONG X R, et al. Mixed linear model approaches of association mapping for complex traits based on omics variants. Scientific Reports, 2015(5):10298. |
| [13] | PATERSON A H, BRUBAKER C L, WENDEL J F. A rapid method for extraction of cotton (Gossypium spp.) genomic DNA suitable for RFLP or PCR analysis. Plant Molecular Biology Reporter, 1993, 11(2):122-127. |
| [14] | YAO J, WANG L X, LIU L H, et al. Association mapping of agronomic traits on chromosome 2A of wheat. Genetica, 2009, 137(1):67-75. |
| [15] | SEARLE S R, CASELLA G, MCCULLOCH C E. Variance Components. Hoboken, USA: John Wiley & Sons, 2009:208-218. |
| [16] | YANG J, ZHU J, WILLIAMS R W. Mapping the genetic architecture of complex traits in experimental populations. Bioinformatics, 2007, 23(12):1527-1536. |
| [17] | YANG J, ZHU J. Methods for predicting superior genotypes under multiple environments based on QTL effects. Theoretical and Applied Genetics, 2005, 110(7):1268-1274. |
| [18] | CARLBORG O, HALEY C S. Epistasis: Too often neglected in complex trait studies? Nature Reviews Genetics, 2004, 5(8):618-625. |
| [19] | JIM O A, RUTTEN B P F. Gene-environment-wide interaction studies in psychiatry. American Journal of Psychiatry, 2009, 166(9):964-966. |
| [20] | MANOLIO T A, COLLINS F S, COX N J, et al. Finding the missing heritability of complex disease. Nature, 2009, 461:747-753. |
| [21] | CULVERHOUSE R, SUAREZ B K, LIN J, et al. A perspective on epistasis: Limits of models displaying no main effect. American Journal of Human Genetics, 2002, 70(2):461-471. |
| [22] | BRACHI B, MORRIS G P, BOREVITZ J O. Genome-wide association studies in plants: The missing heritability is in the field. Genome Biology, 2011, 12(10):5-7. |
| [23] | ZUK O, HECHTER E, SUNYAEV S R, et al. The mystery of missing heritability: Genetic interactions create phantom heritability. Proceedings of the National Academy of Sciences of the USA, 2012, 109(4):1193-1198. |
| [24] | LOU X Y, CHEN G B, YAN L, et al. A Generalized combinatorial approach for detecting gene-by-gene and gene-by-environment interactions with application to nicotine dependence. American Journal of Human Genetics, 2007, 80(6):1125-1137. |
| [25] | PURCELL S, NEALE B, TODD-BROWN K, et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. The American Journal of Human Genetics, 2007, 81(3):559-575. |
| [26] | BRADBURY P J, ZHANG Z, KROON D E, et al. TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics, 2007, 23(19):2633-2635. |
| [27] | R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing: Vienna, Austria, 2012. http://www.R-project.org. |
| [28] | MEI H X, ZHU X F, ZHANG T Z. Favorable QTL alleles for yield and its components identified by association mapping in Chinese upland cotton cultivars. PLoS ONE, 2013, 8(12):e82193. |
| [29] | WANG B H, GUO W Z, ZHU X F, et al. QTL mapping of fiber quality in an elite hybrid derived-RIL population of upland cotton. Euphytica, 2006, 152(3):367-378. |
| [30] | KANTARTZI S K, STEWART J M D. Association analysis of fibre traits in Gossypium arboreum accessions. Plant Breeding, 2008, 127(2):173-179. |
| [31] | RAKSHIT A, RAKSHIT S, SINGH J, et al. Association of AFLP and SSR markers with agronomic and fibre quality traits in Gossypium hirsutum L. Journal of Genetics, 2010, 89(2):155-162. |
| [32] | 梁冰, 范术丽, 宋美珍, 等.陆地棉农艺性状与SSR标记的关联分析.棉花学报, 2014, 26(5):387-395. LIANG B, FAN S L, SONG M Z, et al. Association analysis of agronomic traits in upland cotton using SSR markers. Cotton Science, 2014, 26(5):387-395. (in Chinese with English abstract) |
| [33] | LI C Q, SONG L, ZHAO H H, et al. Identification of quantitative trait loci with main and epistatic effects for plant architecture traits in upland cotton (Gossypium hirsutum L.). Plant Breeding, 2014, 133(3):390-400. |
| [34] | LIU D X, LIU F, SHAN X R, et al. Construction of a high-density genetic map and lint percentage and cottonseed nutrient trait QTL identification in upland cotton (Gossypium hirsutum L.). Molecular Genetics and Genomics, 2015, 290(5):1683-1700. |
| [35] | ALI I, KAUSAR A, UR-REHMAN M, et al. Development of genetic linkage map of leaf hairiness in Gossypium hirsutum (cotton) using molecular markers. Pakistan Journal of Botany, 2009, 41(4):1627-1635. |
| [36] | LI C Q, LIU G S, ZHAO H H, et al. Marker-assisted selection of Verticillium wilt resistance in progeny populations of upland cotton derived from mass selection-mass crossing. Euphytica, 2013, 191(3):469-480. |
 2016, Vol. 42
2016, Vol. 42


